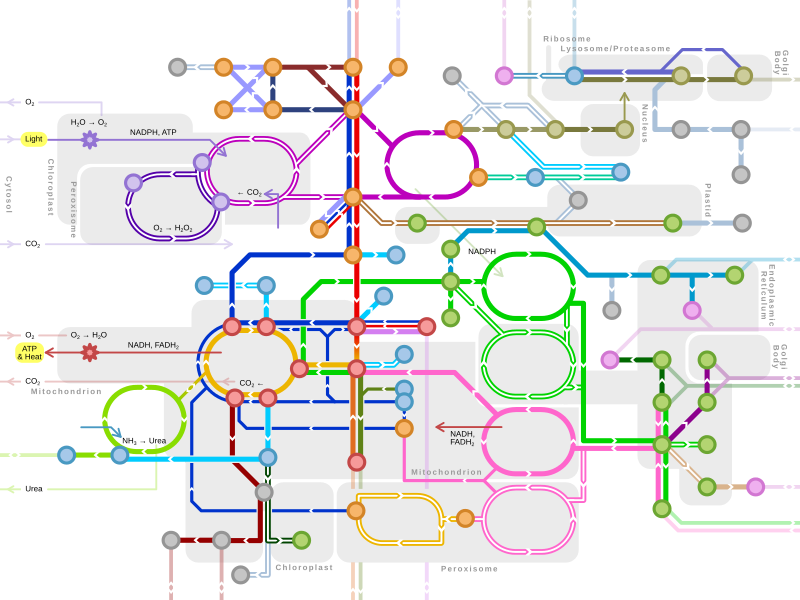
Metabolism

| Part of a series on |
| Biochemistry |
|---|
 |
Metabolism (
Metabolic reactions may be categorized as
The chemical reactions of metabolism are organized into metabolic pathways, in which one chemical is transformed through a series of steps into another chemical, each step being facilitated by a specific enzyme. Enzymes are crucial to metabolism because they allow organisms to drive desirable reactions that require energy and will not occur by themselves, by coupling them to spontaneous reactions that release energy. Enzymes act as catalysts – they allow a reaction to proceed more rapidly – and they also allow the regulation of the rate of a metabolic reaction, for example in response to changes in the cell's environment or to signals from other cells.
The metabolic system of a particular organism determines which substances it will find nutritious and which poisonous. For example, some prokaryotes use hydrogen sulfide as a nutrient, yet this gas is poisonous to animals.[1] The basal metabolic rate of an organism is the measure of the amount of energy consumed by all of these chemical reactions.
A striking feature of metabolism is the similarity of the basic metabolic pathways among vastly different species.
Key biochemicals
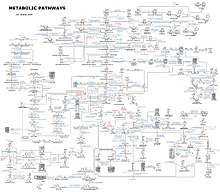
Most of the structures that make up animals, plants and microbes are made from four basic classes of
| Type of molecule | Name of monomer forms | Name of polymer forms | Examples of polymer forms |
|---|---|---|---|
| Amino acids | Amino acids | Proteins (made of polypeptides) | Fibrous proteins and globular proteins |
| Carbohydrates | Monosaccharides | Polysaccharides | Starch, glycogen and cellulose |
| Nucleic acids | Nucleotides | Polynucleotides | DNA and RNA |
Amino acids and proteins
Proteins are made of
Lipids
Lipids are the most diverse group of biochemicals. Their main structural uses are as part of
Carbohydrates
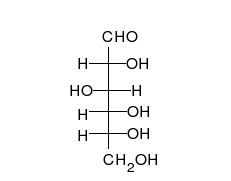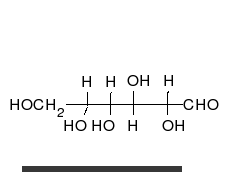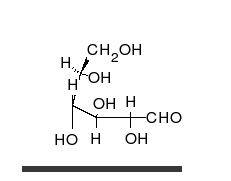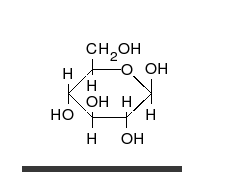
Carbohydrates are
Nucleotides
The two nucleic acids, DNA and
Coenzymes
Metabolism involves a vast array of chemical reactions, but most fall under a few basic types of reactions that involve the transfer of
One central coenzyme is adenosine triphosphate (ATP), the energy currency of cells. This nucleotide is used to transfer chemical energy between different chemical reactions. There is only a small amount of ATP in cells, but as it is continuously regenerated, the human body can use about its own weight in ATP per day.[20] ATP acts as a bridge between catabolism and anabolism. Catabolism breaks down molecules, and anabolism puts them together. Catabolic reactions generate ATP, and anabolic reactions consume it. It also serves as a carrier of phosphate groups in phosphorylation reactions.[21]
A

Mineral and cofactors
Inorganic elements play critical roles in metabolism; some are abundant (e.g. sodium and potassium) while others function at minute concentrations. About 99% of a human's body weight is made up of the elements carbon, nitrogen, calcium, sodium, chlorine, potassium, hydrogen, phosphorus, oxygen and sulfur. Organic compounds (proteins, lipids and carbohydrates) contain the majority of the carbon and nitrogen; most of the oxygen and hydrogen is present as water.[25]
The abundant inorganic elements act as
Transition metals are usually present as trace elements in organisms, with zinc and iron being most abundant of those.[29] Metal cofactors are bound tightly to specific sites in proteins; although enzyme cofactors can be modified during catalysis, they always return to their original state by the end of the reaction catalyzed. Metal micronutrients are taken up into organisms by specific transporters and bind to storage proteins such as ferritin or metallothionein when not in use.[30][31]
Catabolism
Catabolism is the set of metabolic processes that break down large molecules. These include breaking down and oxidizing food molecules. The purpose of the catabolic reactions is to provide the energy and components needed by anabolic reactions which build molecules.
| Energy source | sunlight | photo- | -troph | ||
| molecules | chemo- | ||||
| Hydrogen or electron donor | organic compound | organo- | |||
| inorganic compound | litho- | ||||
| Carbon source | organic compound | hetero- | |||
| inorganic compound | auto- | ||||
The most common set of catabolic reactions in animals can be separated into three main stages. In the first stage, large organic molecules, such as proteins, polysaccharides or lipids, are digested into their smaller components outside cells. Next, these smaller molecules are taken up by cells and converted to smaller molecules, usually acetyl coenzyme A (acetyl-CoA), which releases some energy. Finally, the acetyl group on acetyl-CoA is oxidized to water and carbon dioxide in the citric acid cycle and electron transport chain, releasing more energy while reducing the coenzyme nicotinamide adenine dinucleotide (NAD+) into NADH.[32]
Digestion
Macromolecules cannot be directly processed by cells. Macromolecules must be broken into smaller units before they can be used in cell metabolism. Different classes of enzymes are used to digest these polymers. These
Microbes simply secrete digestive enzymes into their surroundings,[37][38] while animals only secrete these enzymes from specialized cells in their guts, including the stomach and pancreas, and in salivary glands.[39] The amino acids or sugars released by these extracellular enzymes are then pumped into cells by active transport proteins.[40][41]

Energy from organic compounds
Carbohydrate catabolism is the breakdown of carbohydrates into smaller units. Carbohydrates are usually taken into cells after they have been digested into

Fats are catabolized by hydrolysis to free fatty acids and glycerol. The glycerol enters glycolysis and the fatty acids are broken down by beta oxidation to release acetyl-CoA, which then is fed into the citric acid cycle. Fatty acids release more energy upon oxidation than carbohydrates. Steroids are also broken down by some bacteria in a process similar to beta oxidation, and this breakdown process involves the release of significant amounts of acetyl-CoA, propionyl-CoA, and pyruvate, which can all be used by the cell for energy. M. tuberculosis can also grow on the lipid cholesterol as a sole source of carbon, and genes involved in the cholesterol-use pathway(s) have been validated as important during various stages of the infection lifecycle of M. tuberculosis.[45]
Energy transformations
Oxidative phosphorylation
In oxidative phosphorylation, the electrons removed from organic molecules in areas such as the citric acid cycle are transferred to oxygen and the energy released is used to make ATP. This is done in eukaryotes by a series of proteins in the membranes of mitochondria called the electron transport chain. In prokaryotes, these proteins are found in the cell's inner membrane.[49] These proteins use the energy from reduced molecules like NADH to pump protons across a membrane.[50]
Pumping protons out of the mitochondria creates a proton concentration difference across the membrane and generates an electrochemical gradient.[51] This force drives protons back into the mitochondrion through the base of an enzyme called ATP synthase. The flow of protons makes the stalk subunit rotate, causing the active site of the synthase domain to change shape and phosphorylate adenosine diphosphate – turning it into ATP.[20]
Energy from inorganic compounds
Energy from light
The energy in sunlight is captured by plants, cyanobacteria, purple bacteria, green sulfur bacteria and some protists. This process is often coupled to the conversion of carbon dioxide into organic compounds, as part of photosynthesis, which is discussed below. The energy capture and carbon fixation systems can, however, operate separately in prokaryotes, as purple bacteria and green sulfur bacteria can use sunlight as a source of energy, while switching between carbon fixation and the fermentation of organic compounds.[58][59]
In many organisms, the capture of solar energy is similar in principle to oxidative phosphorylation, as it involves the storage of energy as a proton concentration gradient. This proton motive force then drives ATP synthesis.[60] The electrons needed to drive this electron transport chain come from light-gathering proteins called photosynthetic reaction centres. Reaction centers are classified into two types depending on the nature of photosynthetic pigment present, with most photosynthetic bacteria only having one type, while plants and cyanobacteria have two.[61]
In plants, algae, and cyanobacteria, photosystem II uses light energy to remove electrons from water, releasing oxygen as a waste product. The electrons then flow to the cytochrome b6f complex, which uses their energy to pump protons across the thylakoid membrane in the chloroplast.[34] These protons move back through the membrane as they drive the ATP synthase, as before. The electrons then flow through photosystem I and can then be used to reduce the coenzyme NADP+.[62] This coenzyme can enter the Calvin cycle, which is discussed below, or be recycled for further ATP generation.[citation needed]
Anabolism
Anabolism is the set of constructive metabolic processes where the energy released by catabolism is used to synthesize complex molecules. In general, the complex molecules that make up cellular structures are constructed step-by-step from smaller and simpler precursors. Anabolism involves three basic stages. First, the production of precursors such as amino acids, monosaccharides, isoprenoids and nucleotides, secondly, their activation into reactive forms using energy from ATP, and thirdly, the assembly of these precursors into complex molecules such as proteins, polysaccharides, lipids and nucleic acids.[63]
Anabolism in organisms can be different according to the source of constructed molecules in their cells. Autotrophs such as plants can construct the complex organic molecules in their cells such as polysaccharides and proteins from simple molecules like carbon dioxide and water. Heterotrophs, on the other hand, require a source of more complex substances, such as monosaccharides and amino acids, to produce these complex molecules. Organisms can be further classified by ultimate source of their energy: photoautotrophs and photoheterotrophs obtain energy from light, whereas chemoautotrophs and chemoheterotrophs obtain energy from oxidation reactions.[63]
Carbon fixation

Photosynthesis is the synthesis of carbohydrates from sunlight and
In photosynthetic prokaryotes the mechanisms of carbon fixation are more diverse. Here, carbon dioxide can be fixed by the Calvin – Benson cycle, a reversed citric acid cycle,[66] or the carboxylation of acetyl-CoA.[67][68] Prokaryotic chemoautotrophs also fix CO2 through the Calvin–Benson cycle, but use energy from inorganic compounds to drive the reaction.[69]
Carbohydrates and glycans
In carbohydrate anabolism, simple organic acids can be converted into
Although fat is a common way of storing energy, in
Polysaccharides and
Fatty acids, isoprenoids and sterol
Fatty acids are made by fatty acid synthases that polymerize and then reduce acetyl-CoA units. The acyl chains in the fatty acids are extended by a cycle of reactions that add the acyl group, reduce it to an alcohol, dehydrate it to an alkene group and then reduce it again to an alkane group. The enzymes of fatty acid biosynthesis are divided into two groups: in animals and fungi, all these fatty acid synthase reactions are carried out by a single multifunctional type I protein,[79] while in plant plastids and bacteria separate type II enzymes perform each step in the pathway.[80][81]
Proteins
Organisms vary in their ability to synthesize the 20 common amino acids. Most bacteria and plants can synthesize all twenty, but mammals can only synthesize eleven nonessential amino acids, so nine
Amino acids are made into proteins by being joined in a chain of
Nucleotide synthesis and salvage
Nucleotides are made from amino acids, carbon dioxide and
Xenobiotics and redox metabolism
All organisms are constantly exposed to compounds that they cannot use as foods and that would be harmful if they accumulated in cells, as they have no metabolic function. These potentially damaging compounds are called
A related problem for
Thermodynamics of living organisms
Living organisms must obey the laws of thermodynamics, which describe the transfer of heat and work. The second law of thermodynamics states that in any isolated system, the amount of entropy (disorder) cannot decrease. Although living organisms' amazing complexity appears to contradict this law, life is possible as all organisms are open systems that exchange matter and energy with their surroundings. Living systems are not in equilibrium, but instead are dissipative systems that maintain their state of high complexity by causing a larger increase in the entropy of their environments.[106] The metabolism of a cell achieves this by coupling the spontaneous processes of catabolism to the non-spontaneous processes of anabolism. In thermodynamic terms, metabolism maintains order by creating disorder.[107]
Regulation and control
As the environments of most organisms are constantly changing, the reactions of metabolism must be finely regulated to maintain a constant set of conditions within cells, a condition called homeostasis.[108][109] Metabolic regulation also allows organisms to respond to signals and interact actively with their environments.[110] Two closely linked concepts are important for understanding how metabolic pathways are controlled. Firstly, the regulation of an enzyme in a pathway is how its activity is increased and decreased in response to signals. Secondly, the control exerted by this enzyme is the effect that these changes in its activity have on the overall rate of the pathway (the flux through the pathway).[111] For example, an enzyme may show large changes in activity (i.e. it is highly regulated) but if these changes have little effect on the flux of a metabolic pathway, then this enzyme is not involved in the control of the pathway.[112]
There are multiple levels of metabolic regulation. In intrinsic regulation, the metabolic pathway self-regulates to respond to changes in the levels of substrates or products; for example, a decrease in the amount of product can increase the flux through the pathway to compensate.[111] This type of regulation often involves allosteric regulation of the activities of multiple enzymes in the pathway.[113] Extrinsic control involves a cell in a multicellular organism changing its metabolism in response to signals from other cells. These signals are usually in the form of water-soluble messengers such as hormones and growth factors and are detected by specific receptors on the cell surface.[114] These signals are then transmitted inside the cell by second messenger systems that often involved the phosphorylation of proteins.[115]
A very well understood example of extrinsic control is the regulation of glucose metabolism by the hormone
Evolution

The central pathways of metabolism described above, such as glycolysis and the citric acid cycle, are present in all
Many models have been proposed to describe the mechanisms by which novel metabolic pathways evolve. These include the sequential addition of novel enzymes to a short ancestral pathway, the duplication and then divergence of entire pathways as well as the recruitment of pre-existing enzymes and their assembly into a novel reaction pathway.[123] The relative importance of these mechanisms is unclear, but genomic studies have shown that enzymes in a pathway are likely to have a shared ancestry, suggesting that many pathways have evolved in a step-by-step fashion with novel functions created from pre-existing steps in the pathway.[124] An alternative model comes from studies that trace the evolution of proteins' structures in metabolic networks, this has suggested that enzymes are pervasively recruited, borrowing enzymes to perform similar functions in different metabolic pathways (evident in the MANET database)[125] These recruitment processes result in an evolutionary enzymatic mosaic.[126] A third possibility is that some parts of metabolism might exist as "modules" that can be reused in different pathways and perform similar functions on different molecules.[127]
As well as the evolution of new metabolic pathways, evolution can also cause the loss of metabolic functions. For example, in some
Investigation and manipulation

Classically, metabolism is studied by a reductionist approach that focuses on a single metabolic pathway. Particularly valuable is the use of radioactive tracers at the whole-organism, tissue and cellular levels, which define the paths from precursors to final products by identifying radioactively labelled intermediates and products.[130] The enzymes that catalyze these chemical reactions can then be purified and their kinetics and responses to inhibitors investigated. A parallel approach is to identify the small molecules in a cell or tissue; the complete set of these molecules is called the metabolome. Overall, these studies give a good view of the structure and function of simple metabolic pathways, but are inadequate when applied to more complex systems such as the metabolism of a complete cell.[131]
An idea of the complexity of the metabolic networks in cells that contain thousands of different enzymes is given by the figure showing the interactions between just 43 proteins and 40 metabolites to the right: the sequences of genomes provide lists containing anything up to 26.500 genes.[132] However, it is now possible to use this genomic data to reconstruct complete networks of biochemical reactions and produce more holistic mathematical models that may explain and predict their behavior.[133] These models are especially powerful when used to integrate the pathway and metabolite data obtained through classical methods with data on gene expression from proteomic and DNA microarray studies.[134] Using these techniques, a model of human metabolism has now been produced, which will guide future drug discovery and biochemical research.[135] These models are now used in network analysis, to classify human diseases into groups that share common proteins or metabolites.[136][137]
Bacterial metabolic networks are a striking example of bow-tie[138][139][140] organization, an architecture able to input a wide range of nutrients and produce a large variety of products and complex macromolecules using a relatively few intermediate common currencies.[141]
A major technological application of this information is metabolic engineering. Here, organisms such as yeast, plants or bacteria are genetically modified to make them more useful in biotechnology and aid the production of drugs such as antibiotics or industrial chemicals such as 1,3-propanediol and shikimic acid.[142][143][144] These genetic modifications usually aim to reduce the amount of energy used to produce the product, increase yields and reduce the production of wastes.[145]
History
The term metabolism is derived from the Ancient Greek word μεταβολή – "Metabole" for "a change" which derived from μεταβάλλ –"Metaballein" means "To change"[146]

Greek philosophy
Application of the scientific method and Modern metabolic theories
The history of the scientific study of metabolism spans several centuries and has moved from examining whole animals in early studies, to examining individual metabolic reactions in modern biochemistry. The first controlled

In these early studies, the mechanisms of these metabolic processes had not been identified and a
and is notable for being the first organic compound prepared from wholly inorganic precursors. This proved that the organic compounds and chemical reactions found in cells were no different in principle than any other part of chemistry.It was the discovery of
See also
- Anthropogenic metabolism
- Antimetabolite – Chemical that inhibits the use of a metabolite
- Calorimetry – Determining heat transfer in a system by measuring its other properties
- Isothermal microcalorimetry – Measuring versus elapsed time the net rate of heat flow
- Inborn errors of metabolism – Class of genetic diseases
- origin of life
- Metabolic disorder – disease that involving errors in metabolic processes of building or degradation of molecules
- Microphysiometry
- Primary nutritional groups – Group of organisms
- Proto-metabolism
- Respirometry – Estimation of metabolic rates by measuring heat production
- Stream metabolism
- Sulfur metabolism – Set of chemical reactions involving sulfur in living organisms
- Thermic effect of food – effect of food
- Urban metabolism – description and analysis of the flows of materials and energy within cities
- Water metabolism – Aspect of homeostasis concerning control of the amount of water in an organism
- Overflow metabolism – Cellular phenomena
- Oncometabolism
- Reactome – Database of biological pathways
- KEGG – Collection of bioinformatics databases
References
- ^ )
- PMID 11158550.
- ^ PMID 15340153.
- ^ S2CID 44260374.
- ^ S2CID 19107073.
- PMID 29697773.
- PMID 28187287.
- ^ Cooper GM (2000). "The Molecular Composition of Cells". The Cell: A Molecular Approach (2nd ed.). Archived from the original on 27 August 2020. Retrieved 25 June 2020.
- S2CID 4550126.
- ^ ISBN 978-0-7167-4339-2.
- PMID 3120698.
- PMID 23431419.
- PMID 15722563.
- ^ "Lipid nomenclature Lip-1 & Lip-2". qmul.ac.uk. Archived from the original on 6 June 2020. Retrieved 6 June 2020.
- OCLC 913469736.
- S2CID 4644919.
- PMID 16198625.
- ^ PMID 354490.
- PMID 378655.
- ^ PMID 16607397.
- PMID 22528680.
- ^ Berg JM, Tymoczko JL, Stryer L (2002). "Vitamins Are Often Precursors to Coenzymes". Biochemistry. 5th Edition. Archived from the original on 15 December 2020. Retrieved 9 June 2020.
- PMID 17295611.
- OCLC 607553259.
- PMID 1872381.
- ^ "Electrolyte Balance". Anatomy and Physiology. OpenStax. Archived from the original on 2 June 2020. Retrieved 23 June 2020.
- ^ Lodish H, Berk A, Zipursky SL, Matsudaira P, Baltimore D, Darnell J (2000). "The Action Potential and Conduction of Electric Impulses". Molecular Cell Biology (4th ed.). Archived from the original on 30 May 2020. Retrieved 23 June 2020 – via NCBI.
- S2CID 37462321.
- ISBN 978-3-319-73975-5.
- from the original on 25 June 2020. Retrieved 24 June 2020.
- PMID 17194590.
- ^ a b Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2002). "How Cells Obtain Energy from Food". Molecular Biology of the Cell (4th ed.). Archived from the original on 5 July 2021. Retrieved 25 June 2020 – via NCBI.
- from the original on 25 June 2020. Retrieved 25 June 2020.
- ^ S2CID 5686066.
- OCLC 162303067.
- OCLC 945435943.
- PMID 8302217.
- S2CID 206934353.
- PMID 9470654.
- PMID 1494216.
- PMID 10449337.
- PMID 8366068.
- ^ PMID 15466941.
- PMID 25621294.
- PMID 24611808.
- PMID 14144484.
- PMID 10736367.
- PMID 11533293.
- PMID 16756489.
- (PDF) from the original on 22 January 2020. Retrieved 11 November 2019.
- PMID 11893513.
- PMID 8257102.
- from the original on 2 May 2019. Retrieved 6 October 2019.
- PMID 9990725.
- PMID 12165429.
- PMID 8987358.
- PMID 15911555.
- PMID 16000812.
- PMID 11591679.
- ^ Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2002). "Energy Conversion: Mitochondria and Chloroplasts". Molecular Biology of the Cell (4th ed.). Archived from the original on 15 December 2020. Retrieved 3 July 2020.
- S2CID 21596537.
- S2CID 4421776.
- ^ a b Mandal A (26 November 2009). "What is Anabolism?". News-Medical.net. Archived from the original on 5 July 2020. Retrieved 4 July 2020.
- PMID 6351728.
- PMID 11886877.
- PMID 15838028.
- PMID 8354269.
- S2CID 45967404.
- PMID 9891798.
- PMID 6115423.
- S2CID 44741368.
- ^ S2CID 39986630.
- PMID 16815497.
- ^ S2CID 40858130.
- from the original on 31 October 2020. Retrieved 28 August 2020.
- )
- S2CID 10388991.
- PMID 10824734.
- S2CID 4043407.
- PMID 15952903.
- S2CID 46348092.
- S2CID 27523830. Archived from the original(PDF) on 15 April 2007.
- ^ PMID 12735695.
- PMID 16621811.
- PMID 15012203.
- ^ PMID 7023367.
- S2CID 4019443.
- PMID 8948633.
- ISBN 978-0-7216-0240-0.
- PMID 11375928. Archived from the originalon 1 May 2011.
- PMID 4896351.
- ^ PMID 16669783.
- PMID 14658380.
- (PDF) from the original on 24 October 2020. Retrieved 18 September 2019.
- PMID 8749362.
- S2CID 28872968.
- PMID 12369887.
- PMID 11465080.
- PMID 11695986.
- PMID 16125262.
- (PDF) from the original on 11 November 2019. Retrieved 11 November 2019.
- PMID 8660387.
- PMID 14757749.
- S2CID 20240552.
- S2CID 43713549.
- PMID 10482783.
- from the original on 4 August 2020. Retrieved 22 September 2019.
- S2CID 3001195.
- from the original on 29 March 2007. Retrieved 12 March 2007.
- PMID 16045939.
- ^ PMID 7925313.
- S2CID 27791605.
- PMID 7575476.
- S2CID 39154236.
- PMID 11116185.
- PMID 1734513.
- PMID 11949930.
- (PDF) from the original on 19 June 2007. Retrieved 25 March 2007.
- PMID 9084754.
- )
- S2CID 39128865.
- PMID 17517598.
- PMID 12826406.
- PMID 12095253.
- PMID 16854231.
- PMID 11711174.
- PMID 16731630.
- PMID 16230003.
- S2CID 4424895.
- PMID 10817161.
- PMID 9439549.
- PMID 17289424.
- PMID 15961036.
- PMID 16616498.
- PMID 17267599.
- PMID 17502601.
- PMID 18599447.
- PMID 15331224.
- PMID 12874056.
- PMID 16916470.
- ^ "Macromolecules: Nutrients, Metabolism, and Digestive Processes | Virtual High School - KeepNotes". keepnotes.com. Retrieved 29 December 2023.
- PMID 12749845.
- PMID 16095939.
- PMID 14642355.
- S2CID 11814282.
- ^ "metabolism | Origin and meaning of metabolism by Online Etymology Dictionary". www.etymonline.com. Archived from the original on 21 September 2017. Retrieved 23 July 2020.
- ISBN 978-1-4088-3622-4.
- ^ Al-Roubi AS (1982). Ibn Al-Nafis as a philosopher. Symposium on Ibn al-Nafis, Second International Conference on Islamic Medicine. Kuwait: Islamic Medical Organization.
- S2CID 32900603.
- ^ Williams HA (1904). Modern Development of the Chemical and Biological Sciences. A History of Science: in Five Volumes. Vol. IV. New York: Harper and Brothers. pp. 184–185. Retrieved 26 March 2007.
- PMID 8595136.
- S2CID 71727190.
- ^ Eduard Buchner's 1907 Nobel lecture Archived 8 July 2017 at the Wayback Machine at http://nobelprize.org Archived 5 April 2006 at the Wayback Machine Accessed 20 March 2007
- S2CID 28092593.
- .
- PMID 16746382.
Further reading
Introductory
- Rose S, Mileusnic R (1999). The Chemistry of Life. Penguin Press Science. ISBN 0-14-027273-9.
- Schneider EC, Sagan D (2005). Into the Cool: Energy Flow, Thermodynamics, and Life. University of Chicago Press. ISBN 0-226-73936-8.
- Lane N (2004). Oxygen: The Molecule that Made the World. USA: Oxford University Press. ISBN 0-19-860783-0.
Advanced
- Price N, Stevens L (1999). Fundamentals of Enzymology: Cell and Molecular Biology of Catalytic Proteins. Oxford University Press. ISBN 0-19-850229-X.
- Berg J, Tymoczko J, Stryer L (2002). Biochemistry. W. H. Freeman and Company. ISBN 0-7167-4955-6.
- Cox M, Nelson DL (2004). Lehninger Principles of Biochemistry. Palgrave Macmillan. ISBN 0-7167-4339-6.
- ISBN 0-13-066271-2.
- Da Silva JJ, Williams RJ (1991). The Biological Chemistry of the Elements: The Inorganic Chemistry of Life. Clarendon Press. ISBN 0-19-855598-9.
- Nicholls DG, Ferguson SJ (2002). Bioenergetics. Academic Press Inc. ISBN 0-12-518121-3.
- Wood HG (February 1991). "Life with CO or CO2 and H2 as a source of carbon and energy". FASEB Journal. 5 (2): 156–63. S2CID 45967404.
External links
General information
- The Biochemistry of Metabolism (archived 8 March 2005)
- Sparknotes SAT biochemistry Overview of biochemistry. School level.
- MIT Biology Hypertextbook Archived 19 May 2016 at the Portuguese Web Archive Undergraduate-level guide to molecular biology.
Human metabolism
- Topics in Medical Biochemistry Guide to human metabolic pathways. School level.
- THE Medical Biochemistry Page Comprehensive resource on human metabolism.
Databases
- Flow Chart of Metabolic Pathways at ExPASy
- IUBMB-Nicholson Metabolic Pathways Chart
- SuperCYP: Database for Drug-Cytochrome-Metabolism Archived 3 November 2011 at the Wayback Machine
Metabolic pathways
- Metabolism reference Pathway Archived 23 February 2009 at the Wayback Machine
- The Nitrogen cycle and Nitrogen fixation at the Wayback Machine (archive index)