Mobile genetic elements

Mobile genetic elements (MGEs), sometimes called selfish genetic elements,
Types
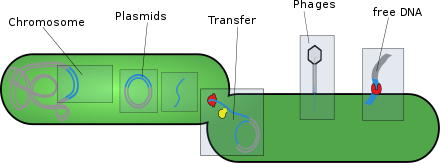

- giant viruses.[8]
- DNA transposons: These are transposons that move directly from one position to another in the genome using a transposase to cut and stick at another locus.[9] These genetic elements are cleaved at four single stranded sites in DNA by transposase. In order to achieve max stability of the intermediate transposon, one single strand cleavage at the target DNA occurs. Simultaneously the donor strand is ligated to the target strand after cleavage leaving a single strand overhang on either end of the target sequence. These sites usually contain a 5 to 9 base pair overhang that can create a cohesive end.[10] Transposase then holds the sequence in a crossed formation and ligates the donor strand to the target strand. The structure formed by the duplex of DNA and transposase in replicative transposons is known as the Shapiro Intermediate.[11] The 5 to 9 base pair overhang is left on either side of the target sequence allowing it to join to its target sequence in either orientation. The sequence of these overhangs can determine joining orientation.[10] Before site specific recombination can occur, the oligonucleotide ends must be filled. The ligation of these ends generates a replication fork at each end of the transposable element. The single strand displacement causes synthesis from the un-ligated 3' hydroxyl group to form long single stranded sections adjacent to the 5' end. Therefore, the opposite strand is sequenced discontinuously as both replication forks approach the center of the transposable element. This results in two recombinant duplexes containing the semi conserved transposable element flanked by the previous 5 to 9 base pair overhang. Site specific reciprocal recombination takes place between the two transposable elements facilitated by proteins. This reciprocal replication overlaps in time and occurs between duplicated segments of the replication element before replication is completed.[10]The target molecule as a result contains the inserted element flanked by the 5 to 9 base pair sequences. Transposition of these elements duplicates the transposition element leaving a transposition element in its original location and a new transposon at the reciprocal replication site. In doing so, organisms total base pairs in their genomes are increased. Transposition occurrences increase over time and as organisms age.

- eukaryotes.[12] Retrotransposons consist of two major types, long terminal repeats (LTRs) and Non-LTR transposons. Non-LTR transposons can be further classified into Long interspersed nuclear element (LINEs) and Short interspersed nuclear element (SINEs).[13] These retrotransposons are regulated by a family of short non-coding RNAs termed as PIWI [P-element induced wimpy testis]-interacting RNAs (piRNAs).[14] piRNA is a recently discovered class of ncRNAs, which are in the length range of ~24-32 nucleotides. Initially, piRNAs were described as repeat-associated siRNAs (rasiRNAs) because of their origin from the repetitive elements such as transposable sequences of the genome. However, later it was identified that they acted via PIWI-protein. In addition to having a role in the suppression of genomic transposons, various roles of piRNAs have been recently reported like regulation of 3’ UTR of protein-coding genes via RNAi, transgenerational epigenetic inheritance to convey a memory of past transposon activity, and RNA-induced epigenetic silencing.[14]
- Integrons: These are gene cassettes that usually carry antibiotic resistance genes to bacterial plasmids and transposons.[15]
- proteins. They are present in all cellular organisms and viruses.[16]
- Introners: Sequences similar to transposons that can jump in the genome leaving new introns where they were, they have been pointed as a possible mechanism of intron gain in the evolution of eukaryotes where they are present in at least 5% of all species, specially in the aquatic taxa due possibly to horizontal gene transfer that occurs more frequently in these animals.[17][18] They were first described in 2009 in the unicellular green algae micromonas.[19]
- living beings and should be better conceived as mobile genetic elements. Viral agents are evolutionarily connected with various mobile genetic elements.[20] These viral agents are thought to have arisen from secreted or ejected plasmids of other organisms. Transposons also provide insight into how these elements may have originally started. This theory is known as the vagrancy hypothesis proposed by Barbara McClintock in 1950.[21][1][22][4][23]
- Satellite nucleic acids: These are DNA or RNA molecules, which are encapsulated as a stowaway in the virions of certain helper viruses and which depend on these to be able to replicate. Although they are sometimes considered genetic elements of their helper viruses, they are not always found within their helper viruses.[20][21][25]
- endogenous retroviruses.[27]
Research examples
CRISPR-Cas systems in bacteria and archaea are adaptive immune systems to protect against deadly consequences from MGEs. Using comparative genomic and phylogenetic analysis, researchers found that CRISPR-Cas variants are associated with distinct types of MGEs such as transposable elements. In CRISPR-associated transposons, CRISPR-Cas controls transposable elements for their propagation.[28]
MGEs such as plasmids by a horizontal transmission are generally beneficial to an organism. The ability of transferring plasmids (sharing) is important in an evolutionary perspective. Tazzyman and Bonhoeffer found that fixation (receiving) of the transferred plasmids in a new organism is just as important as the ability to transfer them.[29] Beneficial rare and transferable plasmids have a higher fixation probability, whereas deleterious transferable genetic elements have a lower fixation probability because they are lethal to the host organisms.
One type of MGEs, namely the Integrative Conjugative Elements (ICEs) are central to horizontal gene transfer shaping the genomes of prokaryotes enabling rapid acquisition of novel adaptive traits.[30][31]
As a representative example of ICEs, the ICEBs1 is well-characterized for its role in the global DNA damage SOS response of Bacillus subtilis[32] and also its potential link to the radiation and desiccation resistance of Bacillus pumilus SAFR-032 spores,[33] isolated from spacecraft cleanroom facilities.[34][35][36]
Transposition by transposable elements is mutagenic. Thus, organisms have evolved to repress the transposition events, and failure to repress the events causes cancers in somatic cells. Cecco et al. found that during early age transcription of retrotransposable elements are minimal in mice, but in advanced age the transcription level increases.[37] This age-dependent expression level of transposable elements is reduced by calorie restriction diet. Replication of transposable elements often results in repeated sequences being added into the genome. These sequences are often non coding but can interfere with coding sequences of DNA. Though mutagenetic by nature, transposons increase the genome of an organism that they transpose into. More research should be conducted into how these elements may serve as a rapid adaptation tool employed by organisms to generate variability. Many transposition elements are dormant or require activation. should also be noted that current values for coding sequences of DNA would be higher if transposition elements that code for their own transposition machinery were considered as coding sequences.
Some others researched examples include Mavericks,[38][39][40] Starships[41][40] and Space invaders (or SPINs)[42][43]
Diseases
The consequence of mobile genetic elements can alter the transcriptional patterns, which frequently leads to genetic disorders such as immune disorders, breast cancer, multiple sclerosis, and amyotrophic lateral sclerosis. In humans, stress can lead to transactional activation of MGEs such as
Other notes
The total of all mobile genetic elements in a genome may be referred to as the mobilome.
Barbara McClintock was awarded the 1983 Nobel Prize in Physiology or Medicine "for her discovery of mobile genetic elements" (transposable elements).[45]
Mobile genetic elements play a critical role in the spread of virulence factors, such as exotoxins and exoenzymes, among bacteria. Strategies to combat certain bacterial infections by targeting these specific virulence factors and mobile genetic elements have been proposed.[46]
See also
- ACLAME (The CLAssification of Mobile genetic Elements) database
- De novo gene birth
- Exon shuffling
- Gene fusion
- Gene duplication
- Horizontal gene transfer
- Virulence factors
- Miniature Inverted-repeat Transposable Elements (MITEs)
References
- ^ S2CID 3907750.
- PMID 27769507.
- PMID 30117500.
- ^ PMID 18948295.
- ^ PMID 20805406.
- ISBN 978-0632034369.
- ISBN 9781555816124.
- S2CID 195814061.
- PMID 20885819.
- ^ PMID 287033.
- OCLC 47283049.
- PMID 26104698.
- S2CID 195814061.
- ^ PMID 32655289.
- S2CID 2078235.
- PMID 24612670.
- PMID 36417430.
- ^ Buehler J (2023-03-30). "How a DNA 'Parasite' May Have Fragmented Our Genes". Quanta Magazine. Retrieved 2023-03-31.
- S2CID 206516961.
- ^ .
- ^ PMID 34468181.
- PMID 24847023.
- PMID 20332804.
- ISBN 978-0199574858.
- ^ Briddon RW, Ghabrial S, Lin NS, Palukaitis P, Scholthof KB, Vetten HJ. "3 - Satellites and Other Virus-dependent Nucleic Acids - Subviral Agents - Subviral Agents (2011)". International Committee on Taxonomy of Viruses (ICTV). Archived from the original on 13 January 2019.
- ^ Di Serio F, Owens RA, Li SF, Matoušek J, Pallás V, Randles JW, et al. (November 2020). "Viroids". International Committee on Taxonomy of Viruses (ICTV). Archived from the original on 2 December 2020.
- S2CID 205485232.
- PMID 28811374.
- PMID 24080312.
- PMID 21876676.
- S2CID 21460836.
- PMID 17511812.
- S2CID 8675124.
- S2CID 13416635.
- PMID 16332797.
- PMID 15941382.
- PMID 24323947.
- PMID 33481003.
- S2CID 250645873.
- ^ a b Bolakhe S (2023-08-03). "Selfish, Virus-Like DNA Can Carry Genes Between Species". Quanta Magazine. Retrieved 2023-08-06.
- PMID 35588244.
- PMID 18936483.
- ^ "Space Invader DNA jumped across mammalian genomes". Science. 2008-11-03. Archived from the original on May 4, 2021. Retrieved 2023-08-06.
- S2CID 9882712.
- ^ "The Nobel Prize in Physiology or Medicine 1983". nobelprize.org. Retrieved 14 July 2010.
- PMID 23248780.
Bibliography
- Miller WJ, Capy P, eds. (2004). Mobile genetic elements: protocols and genomic applications. Humana Press. ISBN 978-1-58829-007-6.
- ISBN 978-0-12-638680-6.
