Coronavirus
| Orthocoronavirinae | |
|---|---|
| |
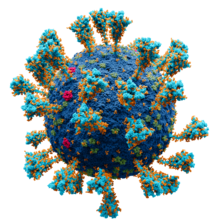
| Group member SARS-CoV-2
Illustration key:
| |
| Virus classification | |
| (unranked): | Virus |
| Realm: | Riboviria |
| Kingdom: | Orthornavirae |
| Phylum: | Pisuviricota |
| Class: | Pisoniviricetes |
| Order: | Nidovirales |
| Family: | Coronaviridae |
| Subfamily: | Orthocoronavirinae |
| Genera[1] | |
|
Information:
| |
| Synonyms[2][3] | |
| |
Coronaviruses are a group of related
Coronaviruses constitute the
Etymology
The name "coronavirus" is derived from Latin
The scientific name Coronavirus was accepted as a genus name by the International Committee for the Nomenclature of Viruses (later renamed
History
The earliest reports of a coronavirus infection in animals occurred in the late 1920s, when an acute respiratory infection of domesticated chickens emerged in North America.[15] Arthur Schalk and M.C. Hawn in 1931 made the first detailed report which described a new respiratory infection of chickens in North Dakota. The infection of new-born chicks was characterized by gasping and listlessness with high mortality rates of 40–90%.[16] Leland David Bushnell and Carl Alfred Brandly isolated the virus that caused the infection in 1933.[17] The virus was then known as infectious bronchitis virus (IBV). Charles D. Hudson and Fred Robert Beaudette cultivated the virus for the first time in 1937.[18] The specimen came to be known as the Beaudette strain. In the late 1940s, two more animal coronaviruses, JHM that causes brain disease (murine encephalitis) and mouse hepatitis virus (MHV) that causes hepatitis in mice were discovered.[19] It was not realized at the time that these three different viruses were related.[20][12]
Human coronaviruses were discovered in the 1960s[21][22] using two different methods in the United Kingdom and the United States.[23] E.C. Kendall, Malcolm Bynoe, and David Tyrrell working at the Common Cold Unit of the British Medical Research Council collected a unique common cold virus designated B814 in 1961.[24][25][26] The virus could not be cultivated using standard techniques which had successfully cultivated rhinoviruses, adenoviruses and other known common cold viruses. In 1965, Tyrrell and Bynoe successfully cultivated the novel virus by serially passing it through organ culture of human embryonic trachea.[27] The new cultivating method was introduced to the lab by Bertil Hoorn.[28] The isolated virus when intranasally inoculated into volunteers caused a cold and was inactivated by ether which indicated it had a lipid envelope.[24][29] Dorothy Hamre and John Procknow at the University of Chicago isolated a novel cold from medical students in 1962. They isolated and grew the virus in kidney tissue culture, designating it 229E. The novel virus caused a cold in volunteers and, like B814, was inactivated by ether.[30][31]
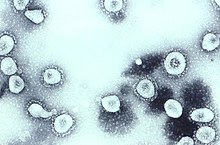
The IBV-like novel cold viruses were soon shown to be also morphologically related to the mouse hepatitis virus.
Microbiology
Structure
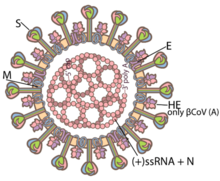
Coronaviruses are large, roughly spherical particles with unique surface projections.[43] Their size is highly variable with average diameters of 80 to 120 nm. Extreme sizes are known from 50 to 200 nm in diameter.[44] The total molecular mass is on average 40,000 kDa. They are enclosed in an envelope embedded with a number of protein molecules.[45] The lipid bilayer envelope, membrane proteins, and nucleocapsid protect the virus when it is outside the host cell.[46]
The viral envelope is made up of a lipid bilayer in which the membrane (M), envelope (E) and spike (S) structural proteins are anchored.[47] The molar ratio of E:S:M in the lipid bilayer is approximately 1:20:300.[48] The E and M protein are the structural proteins that combined with the lipid bilayer to shape the viral envelope and maintain its size.[49] S proteins are needed for interaction with the host cells. But human coronavirus NL63 is peculiar in that its M protein has the binding site for the host cell, and not its S protein.[50] The diameter of the envelope is 85 nm. The envelope of the virus in electron micrographs appears as a distinct pair of electron-dense shells (shells that are relatively opaque to the electron beam used to scan the virus particle).[51][49]
The
The E proteins are minor structural proteins and highly variable in different species.[44] There are only about 20 copies of the E protein molecule in a coronavirus particle.[48] They are 8.4 to 12 kDa in size and are composed of 76 to 109 amino acids.[44] They are integral proteins (i.e. embedded in the lipid layer) and have two domains namely a transmembrane domain and an extramembrane C-terminal domain. They are almost fully α-helical, with a single α-helical transmembrane domain, and form pentameric (five-molecular) ion channels in the lipid bilayer. They are responsible for virion assembly, intracellular trafficking and morphogenesis (budding).[45]

The spikes are the most distinguishing feature of coronaviruses and are responsible for the corona- or halo-like surface. On average a coronavirus particle has 74 surface spikes.
S1 proteins are the most critical components in terms of infection. They are also the most variable components as they are responsible for host cell specificity. They possess two major domains named N-terminal domain (S1-NTD) and C-terminal domain (S1-CTD), both of which serve as the receptor-binding domains. The NTDs recognize and bind sugars on the surface of the host cell. An exception is the
A subset of coronaviruses (specifically the members of betacoronavirus subgroup A) also has a shorter spike-like surface protein called hemagglutinin esterase (HE).[42] The HE proteins occur as homodimers composed of about 400 amino acid residues and are 40 to 50 kDa in size. They appear as tiny surface projections of 5 to 7 nm long embedded in between the spikes. They help in the attachment to and detachment from the host cell.[55]
Inside the envelope, there is the nucleocapsid, which is formed from multiple copies of the nucleocapsid (N) protein, which are bound to the positive-sense single-stranded RNA genome in a continuous beads-on-a-string type conformation.[49][56] N protein is a phosphoprotein of 43 to 50 kDa in size, and is divided into three conserved domains. The majority of the protein is made up of domains 1 and 2, which are typically rich in arginines and lysines. Domain 3 has a short carboxy terminal end and has a net negative charge due to excess of acidic over basic amino acid residues.[44]
Genome
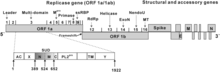

Coronaviruses contain a
The genome organization for a coronavirus is 5′-leader-UTR-replicase (ORF1ab)-spike (S)-envelope (E)-membrane (M)-nucleocapsid (N)-3′UTR-poly (A) tail. The open reading frames 1a and 1b, which occupy the first two-thirds of the genome, encode the replicase polyprotein (pp1ab). The replicase polyprotein self cleaves to form 16 nonstructural proteins (nsp1–nsp16).[49]
The later reading frames encode the four major structural proteins: spike, envelope, membrane, and nucleocapsid.[57] Interspersed between these reading frames are the reading frames for the accessory proteins. The number of accessory proteins and their function is unique depending on the specific coronavirus.[49]
Replication cycle
Cell entry
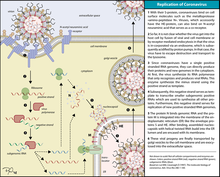
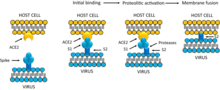
Infection begins when the viral spike protein attaches to its complementary host cell receptor. After attachment, a protease of the host cell cleaves and activates the receptor-attached spike protein. Depending on the host cell protease available, cleavage and activation allows the virus to enter the host cell by endocytosis or direct fusion of the viral envelope with the host membrane.[58]
Coronaviruses can enter cells by either fusing to their lipid envelope with the cell membrane on the cell surface or by internalization via endocytosis.[59]
Genome translation
On entry into the
The larger polyprotein pp1ab is a result of a -1 ribosomal frameshift caused by a slippery sequence (UUUAAAC) and a downstream RNA pseudoknot at the end of open reading frame ORF1a.[60] The ribosomal frameshift allows for the continuous translation of ORF1a followed by ORF1b.[49]
The polyproteins have their own
Replicase-transcriptase
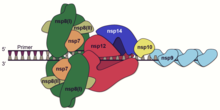
A number of the nonstructural proteins coalesce to form a multi-protein replicase-transcriptase complex (RTC). The main replicase-transcriptase protein is the RNA-dependent RNA polymerase (RdRp). It is directly involved in the replication and transcription of RNA from an RNA strand. The other nonstructural proteins in the complex assist in the replication and transcription process. The exoribonuclease nonstructural protein, for instance, provides extra fidelity to replication by providing a proofreading function which the RNA-dependent RNA polymerase lacks.[61]
Replication – One of the main functions of the complex is to replicate the viral genome. RdRp directly mediates the synthesis of negative-sense genomic RNA from the positive-sense genomic RNA. This is followed by the replication of positive-sense genomic RNA from the negative-sense genomic RNA.[49]
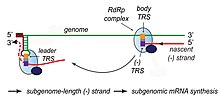
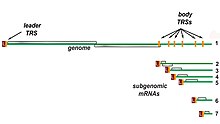
Transcription – The other important function of the complex is to transcribe the viral genome. RdRp directly mediates the synthesis of negative-sense subgenomic RNA molecules from the positive-sense genomic RNA. This process is followed by the transcription of these negative-sense subgenomic RNA molecules to their corresponding positive-sense mRNAs.[49] The subgenomic mRNAs form a "nested set" which have a common 5'-head and partially duplicate 3'-end.[62]
Recombination – The replicase-transcriptase complex is also capable of genetic recombination when at least two viral genomes are present in the same infected cell.[62] RNA recombination appears to be a major driving force in determining genetic variability within a coronavirus species, the capability of a coronavirus species to jump from one host to another and, infrequently, in determining the emergence of novel coronaviruses.[63] The exact mechanism of recombination in coronaviruses is unclear, but likely involves template switching during genome replication.[63]
Assembly and release
The replicated positive-sense genomic RNA becomes the genome of the
Transmission
Infected carriers are able to
Human coronaviruses infect the epithelial cells of the
Classification
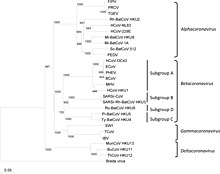
Coronaviruses form the subfamily Orthocoronavirinae,[2][3][4] which is one of two subfamilies in the family Coronaviridae, order Nidovirales, and realm Riboviria.[42][70] They are divided into the four genera: Alphacoronavirus, Betacoronavirus, Gammacoronavirus and Deltacoronavirus. Alphacoronaviruses and betacoronaviruses infect mammals, while gammacoronaviruses and deltacoronaviruses primarily infect birds.[71][72]
- Genus: Alphacoronavirus;[67]
- Genus Betacoronavirus;[68]
- Species: Severe acute respiratory syndrome–related coronavirus (SARS-CoV-1, SARS-CoV-2), Tylonycteris bat coronavirus HKU4
- Species:
- Genus Gammacoronavirus;[18]
- Genus Deltacoronavirus
- Species: Porcine coronavirus HKU15
- Species:
Origin

The
Many human coronaviruses have their origin in bats.
Unlike other betacoronaviruses,
Infection in humans
Coronaviruses vary significantly in risk factor. Some can kill more than 30% of those infected, such as
Six species of human coronaviruses are known, with one species subdivided into two different strains, making seven strains of human coronaviruses altogether.

Four human coronaviruses produce symptoms that are generally mild, even though it is contended they might have been more aggressive in the past:[93]
- Human coronavirus OC43 (HCoV-OC43), β-CoV
- Human coronavirus HKU1 (HCoV-HKU1), β-CoV
- Human coronavirus 229E (HCoV-229E), α-CoV
- Human coronavirus NL63 (HCoV-NL63), α-CoV–
Three human coronaviruses produce potentially severe symptoms:
- Severe acute respiratory syndrome coronavirus(SARS-CoV), β-CoV (identified in 2003)
- Middle East respiratory syndrome-related coronavirus(MERS-CoV), β-CoV (identified in 2012)
- Severe acute respiratory syndrome coronavirus 2(SARS-CoV-2), β-CoV (identified in 2019)
These cause the diseases commonly called
Common cold
Although the common cold is usually caused by rhinoviruses,[94] in about 15% of cases the cause is a coronavirus.[95] The human coronaviruses HCoV-OC43, HCoV-HKU1, HCoV-229E, and HCoV-NL63 continually circulate in the human population in adults and children worldwide and produce the generally mild symptoms of the common cold.[88] The four mild coronaviruses have a seasonal incidence occurring in the winter months in temperate climates.[96][97] There is no preponderance in any season in tropical climates.[98]
Severe acute respiratory syndrome (SARS)
MERS-CoV |
SARS-CoV |
SARS-CoV-2 | |
|---|---|---|---|
| Disease | MERS | SARS | COVID-19 |
| Outbreaks | 2012 MERS outbreak |
2002–2004 | COVID-19 pandemic |
| Epidemiology | |||
| Date of first identified case |
June 2012 |
November 2002 |
December 2019[99] |
| Location of first identified case |
Jeddah, Saudi Arabia |
Shunde , China |
Wuhan, China |
| Age average | 56 | 44[100][a] | 56[101] |
| Sex ratio (M:F) | 3.3:1 | 0.8:1[102] | 1.6:1[101] |
| Confirmed cases | 2494 | 8096[103] | 676,609,955[104][b] |
| Deaths | 858 | 774[103] | 6,881,955[104][b] |
| Case fatality rate | 37% | 9.2% | 1.02%[104] |
| Symptoms | |||
| Fever | 98% | 99–100% | 87.9%[105] |
| Dry cough | 47% | 29–75% | 67.7%[105] |
Dyspnea |
72% | 40–42% | 18.6%[105] |
| Diarrhea | 26% | 20–25% | 3.7%[105] |
| Sore throat | 21% | 13–25% | 13.9%[105] |
| Ventilatory use | 24.5%[106] | 14–20% | 4.1%[107] |
| Notes | |||
In 2003, following the outbreak of severe acute respiratory syndrome (SARS) which had begun the prior year in Asia, and secondary cases elsewhere in the world, the World Health Organization (WHO) issued a press release stating that a novel coronavirus identified by several laboratories was the causative agent for SARS. The virus was officially named the SARS coronavirus (SARS-CoV). More than 8,000 people from 29 countries and territories were infected, and at least 774 died.[108][69]
Middle East respiratory syndrome (MERS)
In September 2012, a new type of coronavirus was identified, initially called Novel Coronavirus 2012, and now officially named Middle East respiratory syndrome coronavirus (MERS-CoV).[109][110] The World Health Organization issued a global alert soon after.[111] The WHO update on 28 September 2012 said the virus did not seem to pass easily from person to person.[112] However, on 12 May 2013, a case of human-to-human transmission in France was confirmed by the French Ministry of Social Affairs and Health.[113] In addition, cases of human-to-human transmission were reported by the Ministry of Health in Tunisia. Two confirmed cases involved people who seemed to have caught the disease from their late father, who became ill after a visit to Qatar and Saudi Arabia. Despite this, it appears the virus had trouble spreading from human to human, as most individuals who are infected do not transmit the virus.[114] By 30 October 2013, there were 124 cases and 52 deaths in Saudi Arabia.[115]
After the Dutch Erasmus Medical Centre sequenced the virus, the virus was given a new name, Human Coronavirus–Erasmus Medical Centre (HCoV-EMC). The final name for the virus is Middle East respiratory syndrome coronavirus (MERS-CoV). The only U.S. cases (both survived) were recorded in May 2014.[116]
In May 2015, an outbreak of MERS-CoV occurred in the
In December 2019, a pneumonia outbreak was reported in Wuhan, China.[119] On 31 December 2019, the outbreak was traced to a novel strain of coronavirus,[120] which was given the interim name 2019-nCoV by the World Health Organization,[121][122][123] later renamed SARS-CoV-2 by the International Committee on Taxonomy of Viruses.
As of 10 March 2023, there were at least 6,881,955[104] confirmed deaths and more than 676,609,955[104] confirmed cases in the COVID-19 pandemic. The Wuhan strain has been identified as a new strain of Betacoronavirus from group 2B with approximately 70% genetic similarity to the SARS-CoV.[124] The virus has a 96% similarity to a bat coronavirus, so it is widely suspected to originate from bats as well.[125][126]
During a surveillance study of archived samples of Malaysian viral pneumonia patients, virologists identified a strain of canine coronavirus which has infected humans in 2018.
Infection in animals
Coronaviruses have been recognized as causing pathological conditions in
Farm animals
Coronaviruses infect domesticated birds.
Coronaviruses also affect other branches of animal husbandry such as pig farming and cattle raising.[127] Swine acute diarrhea syndrome coronavirus (SADS-CoV), which is related to bat coronavirus HKU2, causes diarrhea in pigs.[134] Porcine epidemic diarrhea virus (PEDV) is a coronavirus that has recently emerged and similarly causes diarrhea in pigs.[135] Transmissible gastroenteritis virus (TGEV), which is a member of the species Alphacoronavirus 1,[136] is another coronavirus that causes diarrhea in young pigs.[137][138] In the cattle industry bovine coronavirus (BCV), which is a member of the species Betacoronavirus 1 and related to HCoV-OC43,[139] is responsible for severe profuse enteritis in young calves.[127]
Domestic pets
Coronaviruses infect domestic pets such as cats, dogs, and ferrets.[130] There are two forms of feline coronavirus which are both members of the species Alphacoronavirus 1.[136] Feline enteric coronavirus is a pathogen of minor clinical significance, but spontaneous mutation of this virus can result in feline infectious peritonitis (FIP), a disease with high mortality.[127] There are two different coronaviruses that infect dogs. Canine coronavirus (CCoV), which is a member of the species Alphacoronavirus 1,[136] causes mild gastrointestinal disease.[127] Canine respiratory coronavirus (CRCoV), which is a member of the species Betacoronavirus 1 and related to HCoV-OC43,[139] cause respiratory disease.[127] Similarly, there are two types of coronavirus that infect ferrets.[140] Ferret enteric coronavirus causes a gastrointestinal syndrome known as epizootic catarrhal enteritis (ECE), and a more lethal systemic version of the virus (like FIP in cats) known as ferret systemic coronavirus (FSC).[141][142]
Laboratory animals
Coronaviruses infect laboratory animals.
Prevention and treatment
A number of vaccines using different methods have been developed against human coronavirus SARS-CoV-2.[146][147] Antiviral targets against human coronaviruses have also been identified such as viral proteases, polymerases, and entry proteins. Drugs are in development which target these proteins and the different steps of viral replication.[148][147]
Vaccines are available for animal coronaviruses IBV, TGEV, and Canine CoV, although their effectiveness is limited. In the case of outbreaks of highly contagious animal coronaviruses, such as PEDV, measures such as destruction of entire herds of pigs may be used to prevent transmission to other herds.[49]
See also
References
- ^ "Virus Taxonomy: 2018b Release". International Committee on Taxonomy of Viruses (ICTV). March 2019. Archived from the original on 2018-03-04. Retrieved 2020-01-24.
- ^ a b "2017.012-015S" (xlsx). International Committee on Taxonomy of Viruses (ICTV). October 2018. Archived from the original on 2019-05-14. Retrieved 2020-01-24.
- ^ PMID 30832341.
- ISBN 978-0-323-39281-5.
- ^ PMID 21994708.
Coronaviruses possess the largest genomes [26.4 kb (ThCoV HKU12) to 31.7 kb (SW1)] among all known RNA viruses (Figure 1) [2,13,16].
- ^ PMC 7086490.
[T]here is also a characteristic "fringe" of projections 200 A long, which are rounded or petal shaped ... This appearance, recalling the solar corona, is shared by mouse hepatitis virus and several viruses recently recovered from man, namely strain B814, 229E and several others.
- ^ "Definition of Coronavirus by Merriam-Webster". Merriam-Webster. Archived from the original on 2020-03-23. Retrieved 2020-03-24.
- ^ "Definition of Corona by Merriam-Webster". Merriam-Webster. Archived from the original on 2020-03-24. Retrieved 2020-03-24.
- ^ ISBN 978-0-19-263285-2.
We looked more closely at the appearance of the new viruses and noticed that they had a kind of halo surrounding them. Recourse to a dictionary produced the Latin equivalent, corona, and so the name coronavirus was born.
- PMID 6362367.
[T]hese viruses displayed a characteristic fringe of large, distinctive, petal-shaped peplomers or spikes which resembled a crown, like the corona spinarum in religious art; hence the name coronaviruses.
- ^ .
- PMID 19960211.
- ^ "International Committee on Taxonomy of Viruses (ICTV)". talk.ictvonline.org. Retrieved 2020-09-14.
- PMID 4316767.
- PMID 9876830.
- .
- ^ PMC 7176155.
- ^ ISBN 978-3-642-65775-7.
- ^ "Il était une fois les coronavirus". Réalités Biomédicales (in French). 2020-03-27. Retrieved 2020-04-18.
- PMID 16378050.
- PMID 32299810.
- )
- ^ PMID 14455113.
- PMC 558394.
- S2CID 220187042.
- PMID 14288084.
- ISBN 978-0-19-263285-2.
- ISBN 978-0-8014-1896-9.
- S2CID 1314901.
- ^ Knapp A. "The Secret History Of The First Coronavirus". Forbes. Retrieved 2020-05-06.
- ^ "The woman who discovered the first coronavirus". BBC News. 2020-04-14.
- PMC 2440895.
- PMID 4293939.
- PMID 4298953.
- PMID 5231356.
- ISSN 0362-4331. Retrieved 2020-04-25.
- S2CID 80726096.
- PMID 23202515.
- PMC 7122465,
The other OC strains and B814 that could not be adapted to mouse brain resisted adaptation to cell culture as well; these distinct viruses have since been lost and may actually have been rediscovered recently
- PMID 31978945.
- ^ S2CID 212719285.
- PMID 15030705.
Virions acquired an envelope by budding into the cisternae and formed mostly spherical, sometimes pleomorphic, particles that averaged 78 nm in diameter (Figure 1A).
- ^ PMID 16877062.
- ^ .
- PMID 21130884.
See Figure 10.
- PMID 9233431.
- ^ PMID 1316677.
- ^ PMID 25720466.
See section: Virion Structure.
- PMID 31315999.
- PMID 16873249.
Particle diameters ranged from 50 to 150 nm, excluding the spikes, with mean particle diameters of 82 to 94 nm; Also See Figure 1 for double shell.
- PMID 31133031.
- PMID 21130884.
- PMID 32201500.
- PMID 18550812.
- PMID 24418573.
See Figure 4c.
- PMID 12927536.
See Figure 1.
- PMID 24121034.
See Figure 2.
- PMID 35756892.
- PMID 16877062.
See Figure 8.
- PMID 27279608.
Finally, these results, combined with those from previous work (33, 44), suggest that CoVs encode at least three proteins involved in fidelity (nsp12-RdRp, nsp14-ExoN, and nsp10), supporting the assembly of a multiprotein replicase-fidelity complex, as described previously (38).
- ^ S2CID 91572610.
- ^ PMID 27012512.
- PMID 25720466.
See section: Coronavirus Life Cycle—Assembly and Release
- PMID 16877062.
Nevertheless, the interaction between S protein and receptor remains the principal, if not sole, determinant of coronavirus host species range and tissue tropism.
- PMID 30531947.
Different SARS-CoV strains isolated from several hosts vary in their binding affinities for human ACE2 and consequently in their infectivity of human cells 76, 78 (Fig. 6b)
- ^ PMC 7176201.
- ^ PMC 7176184.
- ^ S2CID 12438123.
- ^ International Committee on Taxonomy of Viruses (2010-08-24). "ICTV Master Species List 2009—v10". Archived from the original (xls) on 2013-04-15.
- PMID 23596293.
Alphacoronaviruses and betacoronaviruses are found exclusively in mammals, whereas gammacoronaviruses and deltacoronaviruses primarily infect birds.
- ^ "Nextstrain, phylogenetic tree of Beta-CoV". nextstrain.org.
- PMID 23596293.
- PMID 22278237.
- ^ PMID 27743750.
Specifically, all HCoVs are thought to have a bat origin, with the exception of lineage A beta-CoVs, which may have reservoirs in rodents [2].
- PMID 22993147.
If these predictions are correct, this observation suggests that HCoV-NL63 may have originated from bats between 1190 and 1449 CE.
- PMID 19788804.
The most recent common ancestor of hCoV-229E and GhanaBt-CoVGrp1 existed in ≈1686–1800 AD.
- PMID 23235471.
- PMID 27743750.
- PMID 23720729.
- PMID 17267506.
- PMID 21763784.
- PMID 18258002.
- PMID 25552712.
- ^ PMID 23804565.
See Table 1
- PMID 15650185.
- PMID 15650185.
However, it is tempting to speculate about an alternative hypothesis, that the 1889-1890 pandemic may have been the result of interspecies transmission of bovine coronaviruses to humans, resulting in the subsequent emergence of HCoV-OC43.
- ^ PMID 29551135.
- PMID 21849456.
- PMID 18508518.
- PMID 29166910.
- ^ S2CID 260316838.
- PMID 32501321.
- ISBN 978-1-4377-1604-7. Archivedfrom the original on 2016-05-04.
- ISBN 978-0-07-015147-5. Archivedfrom the original on 2016-05-16.
- PMID 30541871.
See Figure 1.
- PMID 32246136.
- PMID 20700397.)
{{cite journal}}: CS1 maint: DOI inactive as of March 2024 (link - PMID 31986257.
- PMID 20205928.
- ^ CIDRAP, University of Minnesota. 2020-03-10. Retrieved 2020-03-29.
- PMID 14742282.
- ^ a b "Summary of probable SARS cases with onset of illness from 1 November 2002 to 31 July 2003". World Health Organization. April 2004.
- ^ a b c d e "COVID-19 Dashboard by the Center for Systems Science and Engineering (CSSE) at Johns Hopkins University (JHU)". ArcGIS. Johns Hopkins University. Retrieved 2023-03-10.
- ^ a b c d e "Report of the WHO-China Joint Mission on Coronavirus Disease 2019 (COVID-19)" (PDF). World Health Organization. February 2020.
- PMID 29506344.
- PMID 32145829.
- ^ Pasley J. "How SARS terrified the world in 2003, infecting more than 8,000 people and killing 774". Business Insider. Retrieved 2020-11-08.
- ^ Doucleef M (2012-09-26). "Scientists Go Deep On Genes Of SARS-Like Virus". Associated Press. Archived from the original on 2012-09-27. Retrieved 2012-09-27.
- ^ Falco M (2012-09-24). "New SARS-like virus poses medical mystery". CNN Health. Archived from the original on 2013-11-01. Retrieved 2013-03-16.
- ^ "New SARS-like virus found in Middle East". Al-Jazeera. 2012-09-24. Archived from the original on 2013-03-09. Retrieved 2013-03-16.
- ^ Kelland K (2012-09-28). "New virus not spreading easily between people: WHO". Reuters. Archived from the original on 2012-11-24. Retrieved 2013-03-16.
- ^ Nouveau coronavirus—Point de situation : Un nouveau cas d'infection confirmé Archived 8 June 2013 at the Wayback Machine (Novel coronavirus—Status report: A new case of confirmed infection) 12 May 2013, social-sante.gouv.fr
- ^ "MERS Transmission". Centers for Disease Control and Prevention (CDC). 2019-08-02. Archived from the original on 2019-12-07. Retrieved 2019-12-10.
- ^ "Novel coronavirus infection". World Health Association. 2013-05-22. Archived from the original on 2013-06-07. Retrieved 2013-05-23.
- ^ "MERS in the U.S." Center for Disease Control. 2019-08-02. Archived from the original on 2019-12-15. Retrieved 2019-12-10.
- ^ Sang-Hun C (2015-06-08). "MERS Virus's Path: One Man, Many South Korean Hospitals". The New York Times. Archived from the original on 2017-07-15. Retrieved 2017-03-01.
- ^ "Middle East respiratory syndrome coronavirus (MERS-CoV)". WHO. Archived from the original on 2019-10-18. Retrieved 2019-12-10.
- ^ The Editorial Board (2020-01-29). "Is the World Ready for the Coronavirus?—Distrust in science and institutions could be a major problem if the outbreak worsens". The New York Times. Retrieved 2020-01-30.
- ^ "WHO Statement Regarding Cluster of Pneumonia Cases in Wuhan, China". www.who.int. 2020-01-09. Archived from the original on 2020-01-14. Retrieved 2020-01-10.
- ^ "Laboratory testing of human suspected cases of novel coronavirus (nCoV) infection. Interim guidance, 10 January 2020" (PDF). Archived (PDF) from the original on 2020-01-20. Retrieved 2020-01-14.
- ^ "Novel Coronavirus 2019, Wuhan, China". www.cdc.gov (CDC). 2020-01-23. Archived from the original on 2020-01-20. Retrieved 2020-01-23.
- ^ "2019 Novel Coronavirus infection (Wuhan, China): Outbreak update". Canada.ca. 2020-01-21.
- PMID 31953166.
- ^ Cohen J (2020-01-26). "Wuhan seafood market may not be source of novel virus spreading globally". ScienceMag American Association for the Advancement of Science. (AAAS). Archived from the original on 2020-01-27. Retrieved 2020-01-29.
- ^ Eschner K (2020-01-28). "We're still not sure where the COVID-19 really came from". Popular Science. Archived from the original on 2020-01-30. Retrieved 2020-01-30.
- ^ S2CID 219575461.
- ISBN 978-0-12-511340-3.
- ^ PMID 20627412.
- ^ a b "Merck Veterinary Manual". Merck Veterinary Manual. Retrieved 2020-06-08.
- ^ PMID 25954763.
- PMID 17296157.

- ^ "Taxonomy browser (Avian coronavirus)". www.ncbi.nlm.nih.gov. Retrieved 2020-06-03.
- PMID 29618817.
- PMID 32041637.
- ^ a b c "Taxonomy browser (Alphacoronavirus 1)". www.ncbi.nlm.nih.gov. Retrieved 2020-06-08.
- PMID 21695242.
- PMID 23824792.
- ^ a b "Taxonomy browser (Betacoronavirus 1)". www.ncbi.nlm.nih.gov. Retrieved 2020-06-08.
- ^ "Taxonomy browser (Alphacoronavirus)". www.ncbi.nlm.nih.gov. Retrieved 2020-06-08.
- ^ Murray J (2014-04-16). "What's New With Ferret FIP-like Disease?" (xls). Archived from the original on 2014-04-24. Retrieved 2014-04-24.
- ^ "Infectious Diseases of Ferrets - Exotic and Laboratory Animals". Merck Veterinary Manual. Retrieved 2020-06-08.
- ^ a b "Taxonomy browser (Embecovirus)". www.ncbi.nlm.nih.gov. Retrieved 2020-06-08.
- PMID 16339739.
- ^ "Enteric Coronavirus". Diseases of Research Animals. Archived from the original on 2019-07-01. Retrieved 2020-01-24.
- ^ "COVID-19 vaccine and treatments tracker (Choose vaccines or treatments tab, apply filters to view select data)". Milken Institute. 2020-11-03. Retrieved 2020-11-03.
- ^ a b "COVID-19 vaccine and therapeutics tracker". BioRender. 2020-10-30. Retrieved 2020-11-03.
- PMID 32147628.
Further reading
- Acheson NH (2011). "Chapter 14: Coronaviruses". Fundamentals of molecular virology. Hoboken, NJ: John Wiley & Sons. pp. 159–171. ISBN 978-0-470-90059-8.
- Alwan A, Mahjour J, Memish ZA (2013). "Novel coronavirus infection: time to stay ahead of the curve". Eastern Mediterranean Health Journal. 19 (Suppl 1): S3–4. ]
- Laude H, Rasschaert D, Delmas B, Godet M, Gelfi J, Charley B (June 1990). "Molecular biology of transmissible gastroenteritis virus". Veterinary Microbiology. 23 (1–4): 147–54. PMID 2169670.
- Sola I, Alonso S, Zúñiga S, Balasch M, Plana-Durán J, Enjuanes L (April 2003). "Engineering the transmissible gastroenteritis virus genome as an expression vector inducing lactogenic immunity". Journal of Virology. 77 (7): 4357–69. PMID 12634392.
- Tajima M (1970). "Morphology of transmissible gastroenteritis virus of pigs. A possible member of coronaviruses. Brief report". Archiv für die Gesamte Virusforschung. 29 (1): 105–08. S2CID 42104521.
