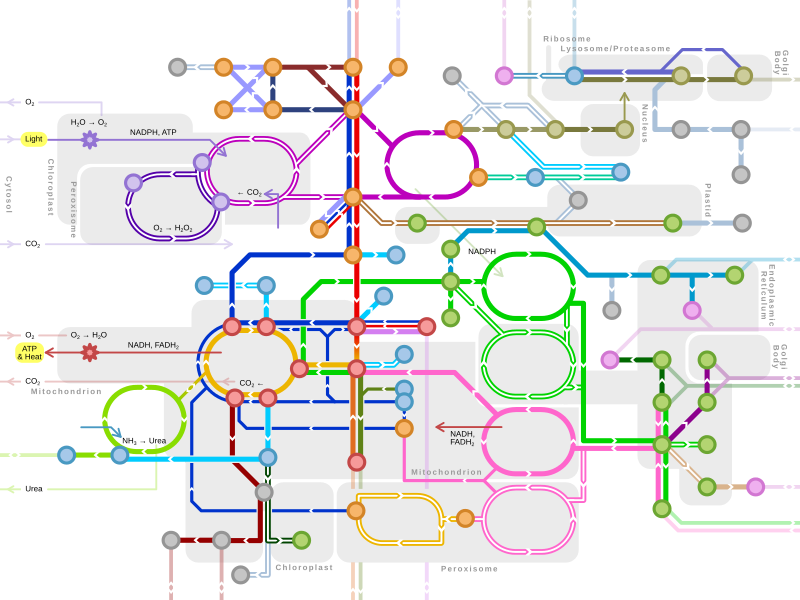
Oxidative phosphorylation
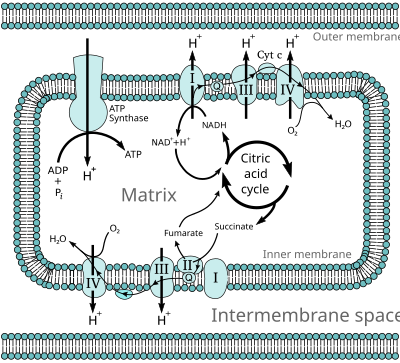
Oxidative phosphorylation (UK
The energy stored in the chemical bonds of
In eukaryotes, these redox reactions are catalyzed by a series of protein complexes within the inner membrane of the cell's mitochondria, whereas, in prokaryotes, these proteins are located in the cell's outer membrane. These linked sets of proteins are called the electron transport chain. In eukaryotes, five main protein complexes are involved, whereas in prokaryotes many different enzymes are present, using a variety of electron donors and acceptors.
The energy transferred by electrons flowing through this electron transport chain is used to transport
Although oxidative phosphorylation is a vital part of metabolism, it produces
Chemiosmosis
Oxidative phosphorylation works by using
ATP synthase releases this stored energy by completing the circuit and allowing protons to flow down the electrochemical gradient, back to the N-side of the membrane.[5] The electrochemical gradient drives the rotation of part of the enzyme's structure and couples this motion to the synthesis of ATP.
The two components of the proton-motive force are
The amount of energy released by oxidative phosphorylation is high, compared with the amount produced by
Electron and proton transfer molecules
The electron transport chain carries both protons and electrons, passing electrons from donors to acceptors, and transporting protons across a membrane. These processes use both soluble and protein-bound transfer molecules. In mitochondria, electrons are transferred within the intermembrane space by the water-
Within the inner mitochondrial membrane, the lipid-soluble electron carrier coenzyme Q10 (Q) carries both electrons and protons by a redox cycle.[10] This small benzoquinone molecule is very hydrophobic, so it diffuses freely within the membrane. When Q accepts two electrons and two protons, it becomes reduced to the ubiquinol form (QH2); when QH2 releases two electrons and two protons, it becomes oxidized back to the ubiquinone (Q) form. As a result, if two enzymes are arranged so that Q is reduced on one side of the membrane and QH2 oxidized on the other, ubiquinone will couple these reactions and shuttle protons across the membrane.[11] Some bacterial electron transport chains use different quinones, such as menaquinone, in addition to ubiquinone.[12]
Within proteins, electrons are transferred between flavin cofactors,[5][13] iron–sulfur clusters and cytochromes. There are several types of iron–sulfur cluster. The simplest kind found in the electron transfer chain consists of two iron atoms joined by two atoms of inorganic sulfur; these are called [2Fe–2S] clusters. The second kind, called [4Fe–4S], contains a cube of four iron atoms and four sulfur atoms. Each iron atom in these clusters is coordinated by an additional amino acid, usually by the sulfur atom of cysteine. Metal ion cofactors undergo redox reactions without binding or releasing protons, so in the electron transport chain they serve solely to transport electrons through proteins. Electrons move quite long distances through proteins by hopping along chains of these cofactors.[14] This occurs by quantum tunnelling, which is rapid over distances of less than 1.4×10−9 m.[15]
Eukaryotic electron transport chains
Many
In eukaryotes, the enzymes in this electron transport system use the energy released from O2 by NADH to pump protons across the inner membrane of the mitochondrion. This causes protons to build up in the intermembrane space, and generates an electrochemical gradient across the membrane. The energy stored in this potential is then used by ATP synthase to produce ATP. Oxidative phosphorylation in the eukaryotic mitochondrion is the best-understood example of this process. The mitochondrion is present in almost all eukaryotes, with the exception of anaerobic protozoa such as Trichomonas vaginalis that instead reduce protons to hydrogen in a remnant mitochondrion called a hydrogenosome.[16]
| Respiratory enzyme | Redox pair | Midpoint potential
(Volts) |
|---|---|---|
| NADH dehydrogenase | NAD+ / NADH | −0.32[17] |
| Succinate dehydrogenase | FMN or FAD / FMNH2 or FADH2 | −0.20[17] |
Cytochrome bc1 complex
|
Coenzyme Q10ox / Coenzyme Q10red | +0.06[17] |
| Cytochrome bc1 complex | Cytochrome box / Cytochrome bred | +0.12[17] |
| Complex IV | Cytochrome cox / Cytochrome cred | +0.22[17] |
| Complex IV | Cytochrome a ox / Cytochrome ared
|
+0.29[17] |
| Complex IV | O2 / HO− | +0.82[17] |
| Conditions: pH = 7[17] | ||
NADH-coenzyme Q oxidoreductase (complex I)

The reaction that is catalyzed by this enzyme is the two electron oxidation of NADH by coenzyme Q10 or ubiquinone (represented as Q in the equation below), a lipid-soluble quinone that is found in the mitochondrion membrane:
-
(1)
The start of the reaction, and indeed of the entire electron chain, is the binding of a NADH molecule to complex I and the donation of two electrons. The electrons enter complex I via a prosthetic group attached to the complex, flavin mononucleotide (FMN). The addition of electrons to FMN converts it to its reduced form, FMNH2. The electrons are then transferred through a series of iron–sulfur clusters: the second kind of prosthetic group present in the complex.[20] There are both [2Fe–2S] and [4Fe–4S] iron–sulfur clusters in complex I.
As the electrons pass through this complex, four protons are pumped from the matrix into the intermembrane space. Exactly how this occurs is unclear, but it seems to involve conformational changes in complex I that cause the protein to bind protons on the N-side of the membrane and release them on the P-side of the membrane.[24] Finally, the electrons are transferred from the chain of iron–sulfur clusters to a ubiquinone molecule in the membrane.[18] Reduction of ubiquinone also contributes to the generation of a proton gradient, as two protons are taken up from the matrix as it is reduced to ubiquinol (QH2).
Succinate-Q oxidoreductase (complex II)
-
(2)
In some eukaryotes, such as the
Electron transfer flavoprotein-Q oxidoreductase
Electron transfer flavoprotein-ubiquinone oxidoreductase (ETF-Q oxidoreductase), also known as electron transferring-flavoprotein dehydrogenase, is a third entry point to the electron transport chain. It is an enzyme that accepts electrons from electron-transferring flavoprotein in the mitochondrial matrix, and uses these electrons to reduce ubiquinone.[30] This enzyme contains a flavin and a [4Fe–4S] cluster, but, unlike the other respiratory complexes, it attaches to the surface of the membrane and does not cross the lipid bilayer.[31]
-
(3)
In mammals, this metabolic pathway is important in beta oxidation of fatty acids and catabolism of amino acids and choline, as it accepts electrons from multiple acetyl-CoA dehydrogenases.[32][33] In plants, ETF-Q oxidoreductase is also important in the metabolic responses that allow survival in extended periods of darkness.[34]
Q-cytochrome c oxidoreductase (complex III)
The reaction catalyzed by complex III is the oxidation of one molecule of ubiquinol and the reduction of two molecules of cytochrome c, a heme protein loosely associated with the mitochondrion. Unlike coenzyme Q, which carries two electrons, cytochrome c carries only one electron.
-
(4)
As only one of the electrons can be transferred from the QH2 donor to a cytochrome c acceptor at a time, the reaction mechanism of complex III is more elaborate than those of the other respiratory complexes, and occurs in two steps called the
As coenzyme Q is reduced to ubiquinol on the inner side of the membrane and oxidized to ubiquinone on the other, a net transfer of protons across the membrane occurs, adding to the proton gradient.[5] The rather complex two-step mechanism by which this occurs is important, as it increases the efficiency of proton transfer. If, instead of the Q cycle, one molecule of QH2 were used to directly reduce two molecules of cytochrome c, the efficiency would be halved, with only one proton transferred per cytochrome c reduced.[5]
Cytochrome c oxidase (complex IV)

Cytochrome c oxidase, also known as complex IV, is the final protein complex in the electron transport chain.[40] The mammalian enzyme has an extremely complicated structure and contains 13 subunits, two heme groups, as well as multiple metal ion cofactors – in all, three atoms of copper, one of magnesium and one of zinc.[41]
This enzyme mediates the final reaction in the electron transport chain and transfers electrons to oxygen and hydrogen (protons), while pumping protons across the membrane.[42] The final electron acceptor oxygen is reduced to water in this step. Both the direct pumping of protons and the consumption of matrix protons in the reduction of oxygen contribute to the proton gradient. The reaction catalyzed is the oxidation of cytochrome c and the reduction of oxygen:
-
(5)
Alternative reductases and oxidases
Many eukaryotic organisms have electron transport chains that differ from the much-studied mammalian enzymes described above. For example, plants have alternative NADH oxidases, which oxidize NADH in the cytosol rather than in the mitochondrial matrix, and pass these electrons to the ubiquinone pool.[43] These enzymes do not transport protons, and, therefore, reduce ubiquinone without altering the electrochemical gradient across the inner membrane.[44]
Another example of a divergent electron transport chain is the alternative oxidase, which is found in plants, as well as some fungi, protists, and possibly some animals.[45][46] This enzyme transfers electrons directly from ubiquinol to oxygen.[47]
The electron transport pathways produced by these alternative NADH and ubiquinone oxidases have lower ATP yields than the full pathway. The advantages produced by a shortened pathway are not entirely clear. However, the alternative oxidase is produced in response to stresses such as cold, reactive oxygen species, and infection by pathogens, as well as other factors that inhibit the full electron transport chain.[48][49] Alternative pathways might, therefore, enhance an organism's resistance to injury, by reducing oxidative stress.[50]
Organization of complexes
The original model for how the respiratory chain complexes are organized was that they diffuse freely and independently in the mitochondrial membrane.
Prokaryotic electron transport chains
In contrast to the general similarity in structure and function of the electron transport chains in eukaryotes, bacteria and archaea possess a large variety of electron-transfer enzymes. These use an equally wide set of chemicals as substrates.[57] In common with eukaryotes, prokaryotic electron transport uses the energy released from the oxidation of a substrate to pump ions across a membrane and generate an electrochemical gradient. In the bacteria, oxidative phosphorylation in Escherichia coli is understood in most detail, while archaeal systems are at present poorly understood.[58]
The main difference between eukaryotic and prokaryotic oxidative phosphorylation is that bacteria and archaea use many different substances to donate or accept electrons. This allows prokaryotes to grow under a wide variety of environmental conditions.[59] In E. coli, for example, oxidative phosphorylation can be driven by a large number of pairs of reducing agents and oxidizing agents, which are listed below. The midpoint potential of a chemical measures how much energy is released when it is oxidized or reduced, with reducing agents having negative potentials and oxidizing agents positive potentials.
| Respiratory enzyme | Redox pair | Midpoint potential
(Volts) |
|---|---|---|
| Formate dehydrogenase | Bicarbonate / Formate | −0.43 |
| Hydrogenase | Proton / Hydrogen | −0.42 |
| NADH dehydrogenase | NAD+ / NADH | −0.32 |
| Glycerol-3-phosphate dehydrogenase | DHAP / Gly-3-P | −0.19 |
| Pyruvate oxidase | Acetate + Carbon dioxide / Pyruvate | ? |
| Lactate dehydrogenase | Pyruvate / Lactate | −0.19 |
| D-amino acid dehydrogenase | 2-oxoacid + ammonia / D-amino acid
|
? |
| Glucose dehydrogenase | Gluconate / Glucose | −0.14 |
Succinate dehydrogenase
|
Fumarate / Succinate | +0.03 |
| Ubiquinol oxidase | Oxygen / Water | +0.82 |
| Nitrate reductase | Nitrate / Nitrite | +0.42 |
| Nitrite reductase | Nitrite / Ammonia | +0.36 |
| Dimethyl sulfoxide reductase | DMSO / DMS | +0.16 |
| Trimethylamine N-oxide reductase | TMAO / TMA | +0.13 |
| Fumarate reductase | Fumarate / Succinate | +0.03 |
As shown above, E. coli can grow with reducing agents such as formate, hydrogen, or lactate as electron donors, and nitrate, DMSO, or oxygen as acceptors.
Some prokaryotes use redox pairs that have only a small difference in midpoint potential. For example, nitrifying bacteria such as Nitrobacter oxidize nitrite to nitrate, donating the electrons to oxygen. The small amount of energy released in this reaction is enough to pump protons and generate ATP, but not enough to produce NADH or NADPH directly for use in anabolism.[62] This problem is solved by using a nitrite oxidoreductase to produce enough proton-motive force to run part of the electron transport chain in reverse, causing complex I to generate NADH.[63][64]
Prokaryotes control their use of these electron donors and acceptors by varying which enzymes are produced, in response to environmental conditions.[65] This flexibility is possible because different oxidases and reductases use the same ubiquinone pool. This allows many combinations of enzymes to function together, linked by the common ubiquinol intermediate.[60] These respiratory chains therefore have a modular design, with easily interchangeable sets of enzyme systems.
In addition to this metabolic diversity, prokaryotes also possess a range of isozymes – different enzymes that catalyze the same reaction. For example, in E. coli, there are two different types of ubiquinol oxidase using oxygen as an electron acceptor. Under highly aerobic conditions, the cell uses an oxidase with a low affinity for oxygen that can transport two protons per electron. However, if levels of oxygen fall, they switch to an oxidase that transfers only one proton per electron, but has a high affinity for oxygen.[66]
ATP synthase (complex V)
ATP synthase, also called complex V, is the final enzyme in the oxidative phosphorylation pathway. This enzyme is found in all forms of life and functions in the same way in both prokaryotes and eukaryotes.[67] The enzyme uses the energy stored in a proton gradient across a membrane to drive the synthesis of ATP from ADP and phosphate (Pi). Estimates of the number of protons required to synthesize one ATP have ranged from three to four,[68][69] with some suggesting cells can vary this ratio, to suit different conditions.[70]
-
(6)
This phosphorylation reaction is an equilibrium, which can be shifted by altering the proton-motive force. In the absence of a proton-motive force, the ATP synthase reaction will run from right to left, hydrolyzing ATP and pumping protons out of the matrix across the membrane. However, when the proton-motive force is high, the reaction is forced to run in the opposite direction; it proceeds from left to right, allowing protons to flow down their concentration gradient and turning ADP into ATP.[67] Indeed, in the closely related vacuolar type H+-ATPases, the hydrolysis reaction is used to acidify cellular compartments, by pumping protons and hydrolysing ATP.[71]
ATP synthase is a massive protein complex with a mushroom-like shape. The mammalian enzyme complex contains 16 subunits and has a mass of approximately 600
As protons cross the membrane through the channel in the base of ATP synthase, the FO proton-driven motor rotates.
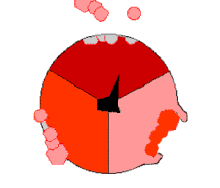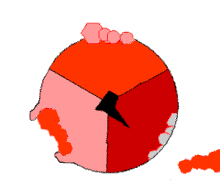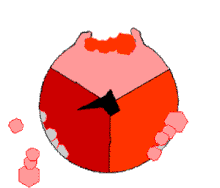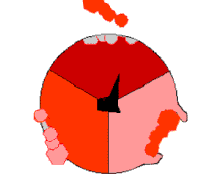
This ATP synthesis reaction is called the binding change mechanism and involves the active site of a β subunit cycling between three states.[77] In the "open" state, ADP and phosphate enter the active site (shown in brown in the diagram). The protein then closes up around the molecules and binds them loosely – the "loose" state (shown in red). The enzyme then changes shape again and forces these molecules together, with the active site in the resulting "tight" state (shown in pink) binding the newly produced ATP molecule with very high affinity. Finally, the active site cycles back to the open state, releasing ATP and binding more ADP and phosphate, ready for the next cycle.
In some bacteria and archaea, ATP synthesis is driven by the movement of sodium ions through the cell membrane, rather than the movement of protons.[78][79] Archaea such as Methanococcus also contain the A1Ao synthase, a form of the enzyme that contains additional proteins with little similarity in sequence to other bacterial and eukaryotic ATP synthase subunits. It is possible that, in some species, the A1Ao form of the enzyme is a specialized sodium-driven ATP synthase,[80] but this might not be true in all cases.[79]
Oxidative phosphorylation - energetics
The transport of electrons from redox pair NAD+/ NADH to the final redox pair 1/2 O2/ H2O can be summarized as
1/2 O2 + NADH + H+ → H2O + NAD+
The potential difference between these two redox pairs is 1.14 volt, which is equivalent to -52 kcal/mol or -2600 kJ per 6 mol of O2.
When one NADH is oxidized through the electron transfer chain, three ATPs are produced, which is equivalent to 7.3 kcal/mol x 3 = 21.9 kcal/mol.
The conservation of the energy can be calculated by the following formula
Efficiency = (21.9 x 100%) / 52 = 42%
So we can conclude that when NADH is oxidized, about 42% of energy is conserved in the form of three ATPs and the remaining (58%) energy is lost as heat (unless the chemical energy of ATP under physiological conditions was underestimated).
Reactive oxygen species
Molecular oxygen is a good terminal electron acceptor because it is a strong oxidizing agent. The reduction of oxygen does involve potentially harmful intermediates.[81] Although the transfer of four electrons and four protons reduces oxygen to water, which is harmless, transfer of one or two electrons produces superoxide or peroxide anions, which are dangerously reactive.
-
(7)
These
The cytochrome c oxidase complex is highly efficient at reducing oxygen to water, and it releases very few partly reduced intermediates; however small amounts of superoxide anion and peroxide are produced by the electron transport chain.
To counteract these reactive oxygen species, cells contain numerous
Oxidative phosphorylation in hypoxic conditions
As oxygen is fundamental for oxidative phosphorylation, a shortage in O2 level likely alters ATP production rates. However, proton motive force and ATP production can be maintained by intracellular acidosis.[88] Cytosolic protons that have accumulated with ATP hydrolysis and lactic acidosis can freely diffuse across the mitochondrial outer-membrane and acidify the inter-membrane space, hence directly contributing to the proton motive force and ATP production.
Inhibitors
There are several well-known drugs and toxins that inhibit oxidative phosphorylation. Although any one of these toxins inhibits only one enzyme in the electron transport chain, inhibition of any step in this process will halt the rest of the process. For example, if oligomycin inhibits ATP synthase, protons cannot pass back into the mitochondrion.[89] As a result, the proton pumps are unable to operate, as the gradient becomes too strong for them to overcome. NADH is then no longer oxidized and the citric acid cycle ceases to operate because the concentration of NAD+ falls below the concentration that these enzymes can use.
Many site-specific inhibitors of the electron transport chain have contributed to the present knowledge of mitochondrial respiration. Synthesis of ATP is also dependent on the electron transport chain, so all site-specific inhibitors also inhibit ATP formation. The fish poison rotenone, the barbiturate drug amytal, and the antibiotic piericidin A inhibit NADH and coenzyme Q.[90]
Carbon monoxide, cyanide, hydrogen sulphide and azide effectively inhibit cytochrome oxidase. Carbon monoxide reacts with the reduced form of the cytochrome while cyanide and azide react with the oxidised form. An antibiotic, antimycin A, and British anti-Lewisite, an antidote used against chemical weapons, are the two important inhibitors of the site between cytochrome B and C1.[90]
| Compounds | Use | Site of action | Effect on oxidative phosphorylation |
|---|---|---|---|
| Cyanide Carbon monoxide Azide Hydrogen sulfide |
Poisons | Complex IV | Inhibit the electron transport chain by binding more strongly than oxygen to the Fe–Cu center in cytochrome c oxidase, preventing the reduction of oxygen.[91] |
| Oligomycin | Antibiotic | Complex V | Inhibits ATP synthase by blocking the flow of protons through the Fo subunit.[89] |
| CCCP 2,4-Dinitrophenol |
Poisons, weight-loss[N 1] | Inner membrane | uncouples proton pumping from ATP synthesis because it carries protons across the inner mitochondrial membrane.[92]
|
| Rotenone | Pesticide | Complex I | Prevents the transfer of electrons from complex I to ubiquinone by blocking the ubiquinone-binding site.[93] |
oxaloacetate
|
Poisons | Complex II | Competitive inhibitors of succinate dehydrogenase (complex II).[94] |
| Antimycin A | Piscicide | Complex III | Binds to the Qi site of oxidation of ubiquinol .
|
Not all inhibitors of oxidative phosphorylation are toxins. In
History
The field of oxidative phosphorylation began with the report in 1906 by
For another twenty years, the mechanism by which ATP is generated remained mysterious, with scientists searching for an elusive "high-energy intermediate" that would link oxidation and phosphorylation reactions.
See also
- Respirometry
- TIM/TOM Complex
Notes
- ^ DNP was extensively used as an anti-obesity medication in the 1930s but was ultimately discontinued due to its dangerous side effects. However, illicit use of the drug for this purpose continues today. See 2,4-Dinitrophenol#Dieting aid for more information.
References
- ^ "oxidative Meaning in the Cambridge English Dictionary". dictionary.cambridge.org. Archived from the original on 24 January 2018. Retrieved 28 April 2018.
- ^ Voet, D.; Voet, J. G. (2004). "Biochemistry", 3rd ed., p. 804, Wiley.ISBN 0-471-19350-X.
- S2CID 4149605.
- ^ from the original on 30 September 2007.
- ^ PMID 11340051.
- PMID 14641005.
- PMID 7654171.
- PMID 3881803.
- S2CID 7685958.
- S2CID 28013583.
- PMID 388618.
- (PDF) from the original on 2008-05-29.
- PMID 15952888.
- S2CID 4431405.
- PMID 15582386.
- S2CID 4401178.
- ^ a b c d e f g h Medical CHEMISTRY Compendium. By Anders Overgaard Pedersen and Henning Nielsen. Aarhus University. 2008
- ^ PMID 15916556.
- ^ PMID 16828051.
- ^ S2CID 1892332.
- S2CID 4372778.
- PMID 17157874.
- PMID 14741580.
- PMID 20025615.
- PMID 14527321.
- S2CID 29222766.
- PMID 15078221.
- PMID 11803022.
- S2CID 4421676.
- PMID 3593226.
- PMID 17050691.
- from the original on 29 September 2007.
- (PDF) from the original on 2007-09-27.
- PMID 16055629.
- (PDF) from the original on 2015-12-28.
- PMID 14977419.
- PMID 9651245.
- (PDF) from the original on 2007-09-27.
- S2CID 13942619.
- PMID 7940677.
- S2CID 20860573.
- PMID 16904626.
- PMID 15725055.
- S2CID 893754.
- PMID 15370881.
- PMID 9698817.
- PMID 1883834.
- PMID 15012279.
- PMID 9426242.
- PMID 10393984.
- S2CID 46138989.
- S2CID 18123642.
- PMID 10775262.
- PMID 12206908.
- from the original on 2007-09-29.
- PMID 6326133.
- S2CID 12289639.
- PMID 10477309.
- ^ PMID 6387427.
- ^ PMID 9230919.
- PMID 11803023.
- .
- PMID 16517654.
- PMID 2856189.
- S2CID 39165641.
- (PDF) from the original on 2007-09-27.
- ^ PMID 9242922.
- S2CID 35989618.
- S2CID 3926411.
- PMID 9620972.
- from the original on 30 September 2007.
- PMID 14633978.
- PMID 10836500.
- S2CID 30953216.
- PMID 11893513.
- PMID 16607397.
- ^ from the original on 29 September 2007.
- S2CID 23763996.
- ^ PMID 8169202.
- S2CID 24887884.
- ^ PMID 8660387.
- S2CID 11125090. Archived from the original(PDF) on 2014-06-14. Retrieved 2017-10-27.
- PMID 16978905.
- PMID 11050436.
- S2CID 2502238.
- PMID 19409964.
- S2CID 4349744.
- PMID 30713504.
- ^ PMID 1831660.
- ^ OCLC 71209231.
- PMID 8380331.
- PMID 156853.
- S2CID 26620903.
- PMID 14249115.
- PMID 10620491.
- S2CID 18598358.
- .
- S2CID 26999163.
- ISBN 9780674366701.
- from the original on 16 December 2008.
- PMID 1883194.
- ^ Belitser VA, Tsibakova ET (1939). "About phosphorilation mechanism coupled with respiration". Biokhimiya. 4: 516–534.
- S2CID 4153659.
- S2CID 1784050.
- OCLC 55202414.
- ^ Mitchell P (1978). "David Keilin's Respiratory Chain Concept and Its Chemiosmotic Consequences" (PDF). Nobel lecture. Nobel Foundation. Archived (PDF) from the original on 2007-09-27. Retrieved 2007-07-21.
- from the original on 29 September 2007.
- PMID 4517936.
- ^ "The Nobel Prize in Chemistry 1997". Nobel Foundation. Archived from the original on 2017-03-25. Retrieved 2007-07-21.
Further reading
Introductory
- Nelson DL, Cox MM (2004). Lehninger Principles of Biochemistry (4th ed.). W. H. Freeman. ISBN 0-7167-4339-6.
- Schneider ED, Sagan D (2006). Into the Cool: Energy Flow, Thermodynamics and Life (1st ed.). University of Chicago Press. ISBN 0-226-73937-6.
- ISBN 0-19-920564-7.
Advanced
- Nicholls DG, Ferguson SJ (2002). Bioenergetics 3 (1st ed.). Academic Press. ISBN 0-12-518121-3.
- Haynie D (2001). Biological Thermodynamics (1st ed.). Cambridge University Press. ISBN 0-521-79549-4.
- Rajan SS (2003). Introduction to Bioenergetics (1st ed.). Anmol. ISBN 81-261-1364-2.
- Wikstrom M, ed. (2005). Biophysical and Structural Aspects of Bioenergetics (1st ed.). Royal Society of Chemistry. ISBN 0-85404-346-2.
General resources
- Animated diagrams illustrating oxidative phosphorylation Wiley and CoConcepts in Biochemistry
- On-line biophysics lectures Antony Crofts, University of Illinois at Urbana–Champaign
- ATP Synthase Graham Johnson
Structural resources
- PDB molecule of the month:
- ATP synthase Archived 2020-07-24 at the Wayback Machine
- Cytochrome c Archived 2020-07-24 at the Wayback Machine
- Cytochrome c oxidase Archived 2020-07-24 at the Wayback Machine
- Interactive molecular models at Universidade Fernando Pessoa:







![{\displaystyle {\ce {O2->[{\ce {e^{-}}}]{\underset {Superoxide}{O2^{\underline {\bullet }}}}->[{\ce {e^{-}}}]{\underset {Peroxide}{O2^{2-}}}}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/a3d9bf9d3a61736aa6207fa53b8ce0165b9eebb6)