Paleopolyploidy

Paleopolyploidy is the result of
Paleopolyploidy is extensively studied in plant lineages. It has been found that almost all flowering plants have undergone at least one round of genome duplication at some point during their evolutionary history. Ancient genome duplications are also found in the early ancestor of vertebrates (which includes the human lineage) near the origin of the
The term mesopolyploid is sometimes used for species that have undergone whole genome multiplication events (whole genome duplication, whole genome triplification, etc.) in more recent history, such as within the last 17 million years.[8]
Eukaryotes
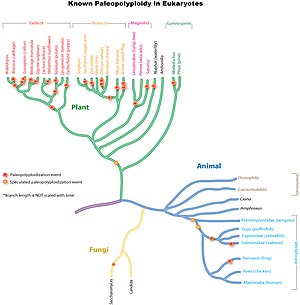
Ancient genome duplications are widespread throughout
A polyploidy event 160 million years ago is theorized to have created the ancestral line that led to all modern flowering plants.
The core eudicots also shared a common whole genome triplication (paleo-hexaploidy), which was estimated to have occurred after monocot-eudicot divergence but before the divergence of rosids and asterids.[13][14][15] Many eudicot species have experienced additional whole genome duplications or triplications. For example, the model plant Arabidopsis thaliana, the first plant to have its entire genome sequenced, has experienced at least two additional rounds of whole genome duplication since the duplication shared by the core eudicots.[2] The most recent event took place before the divergence of the Arabidopsis and Brassica lineages, about 20 million years ago to 45 million years ago. Other examples include the sequenced eudicot genomes of apple, soybean, tomato, cotton, etc.[citation needed]
Compared with plants, paleopolyploidy is much rarer in the animal kingdom. It has been identified mainly in amphibians and bony fishes. Although some studies suggested one or more common genome duplications are shared by all vertebrates (including humans), the evidence is not as strong as in the other cases because the duplications, if they exist, happened so long ago (about 400-500 Ma compared to less than 200 Ma in plants), and the matter is still under debate. The idea that vertebrates share a common whole genome duplication is known as the
A well-supported paleopolyploidy has been found in baker's yeast (Saccharomyces cerevisiae), despite its small, compact genome (~13Mbp), after the divergence from Kluyveromyces lactis and K. marxianus.[16] Through genome streamlining, yeast has lost 90% of the duplicated genome over evolutionary time and is now recognized as a diploid organism.[citation needed]
Detection method
Duplicated genes can be identified through sequence homology on the DNA or protein level. Paleopolyploidy can be identified as massive gene duplication at one time using a molecular clock. To distinguish between whole-genome duplication and a collection of (more common) single gene duplication events, the following rules are often applied:

- Duplicated genes are located in large duplicated blocks. Single gene duplication is a random process and tends to make duplicated genes scattered throughout the genome.
- Duplicated blocks are non-overlapping because they were created simultaneously. Segmental duplication within the genome can fulfill the first rule; but multiple independent segmental duplications could overlap each other.
In theory, the two duplicated genes should have the same "age"; that is, the divergence of the sequence should be equal between the two genes duplicated by paleopolyploidy (
However, using Ks plots to identify and document ancient polyploid events can be problematic, as the method fails to identify genome duplications that were followed by massive gene elimination and genome refinement. Other mixed model approaches that combined Ks plots with other methods are being developed to better understand paleopolyploidy.[17]
Duplication events that occurred a long time ago in the history of various evolutionary lineages can be difficult to detect because of subsequent diploidization (such that a polyploid starts to behave cytogenetically as a diploid over time) as mutations and gene translations gradually make one copy of each chromosome unlike its counterpart. This usually results in a low confidence for identifying a very ancient paleopolyploidy.
Evolutionary importance
Paleopolyploidization events lead to massive cellular changes, including doubling of the genetic material, changes in gene expression and increased cell size. Gene loss during diploidization is not completely random, but heavily selected. Genes from large gene families are duplicated. On the other hand, individual genes are not duplicated.[clarification needed] Overall, paleopolyploidy can have both short-term and long-term evolutionary effects on an organism's fitness in the natural environment.[citation needed]
Enhanced phenotypic evolution
Whole genome duplication may increase the rates and efficiency by which organisms acquire new biological traits. However, one test of this hypothesis, which compared evolutionary rates in innovation in early teleost fishes (with duplicate genomes) to early holostean fishes (without duplicated genomes) found little difference between the two.[7]
Genome diversity
Genome doubling provided the organism with redundant alleles that can evolve freely with little selection pressure. The duplicated genes can undergo neofunctionalization or subfunctionalization which could help the organism adapt to the new environment or survive different stress conditions.[citation needed]
Hybrid vigor
Polyploids often have larger cells and even larger organs. Many important crops, including wheat, maize and cotton, are paleopolyploids which were selected for domestication by ancient peoples.[citation needed]
Speciation
It has been suggested that many polyploidization events created new species, via a gain of adaptive traits, or by sexual incompatibility with their diploid counterparts. An example would be the recent
Allopolyploidy and autopolyploidy
There are two major divisions of
Following polyploidy events, there are several possible fates for duplicated
Vertebrates as paleopolyploid
The hypothesis of vertebrate paleopolyploidy originated as early as the 1970s, proposed by the biologist Susumu Ohno. He reasoned that the vertebrate genome could not achieve its complexity without large scale whole-genome duplications. The "two rounds of genome duplication" hypothesis (2R hypothesis) came about, and gained in popularity, especially among developmental biologists.[citation needed]
Some researchers have questioned the 2R hypothesis because it predicts that vertebrate genomes should have a 4:1 gene ratio compared with invertebrate genomes, and this is not supported by findings from the 48 vertebrate genome projects available in mid-2011. For example, the human genome consists of ~21,000 protein coding genes according to June, 2011 counts at UCSC and Ensembl genome analysis centers[
See also
References
- S2CID 4422074.
- ^ S2CID 4423658.
- PMID 23435085.
- S2CID 20796914.
- PMID 15208399.
- PMID 15208398.
- ^ PMID 27671652.
- S2CID 205358099.
- PMID 15161969.
- PMID 19966307.
- S2CID 88293665.
- S2CID 206553839.
- PMID 18832442.
- PMID 17721507.
- S2CID 206510918.
- PMID 12093907.
- PMID 30239709.
- PMID 22040744.
- ^ PMID 10860970.
- ^ PMID 20070540.
- )
- ^ PMID 20671102.
- PMID 18563158.
- PMID 12028790.
- PMID 26048246.
Further reading
- Adams KL, Wendel JF (April 2005). "Polyploidy and genome evolution in plants". Current Opinion in Plant Biology. 8 (2): 135–41. PMID 15752992.
- Cui L, Wall PK, Leebens-Mack JH, Lindsay BG, Soltis DE, Doyle JJ, et al. (June 2006). "Widespread genome duplications throughout the history of flowering plants". Genome Research. 16 (6): 738–49. PMID 16702410.
- Comai L (November 2005). "The advantages and disadvantages of being polyploid". Nature Reviews. Genetics. 6 (11): 836–46. S2CID 3329282.
- Eckardt NA (July 2004). "Two genomes are better than one: widespread paleopolyploidy in plants and evolutionary effects". The Plant Cell. 16 (7): 1647–1649. PMID 15272471.
- Otto SP, Whitton J (2000). "Polyploid incidence and evolution". Annual Review of Genetics. 34 (1): 401–437. PMID 11092833.
- Makalowski W (May 2001). "Are we polyploids? A brief history of one hypothesis". Genome Research. 11 (5): 667–70. PMID 11337465.
- Kellis M, Birren BW, Lander ES (April 2004). "Proof and evolutionary analysis of ancient genome duplication in the yeast Saccharomyces cerevisiae". Nature. 428 (6983): 617–24. S2CID 4422074.
