Protease
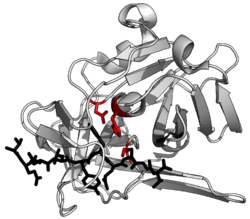
A protease (also called a peptidase, proteinase, or proteolytic enzyme)
In the absence of functional accelerants, proteolysis would be very slow, taking hundreds of
Classification
Based on catalytic residue
Proteases can be classified into seven broad groups:[6]
- Serine proteases - using a serine alcohol
- Cysteine proteases - using a cysteine thiol
- secondary alcohol
- Aspartic proteases - using an aspartate carboxylic acid
- Glutamic proteases - using a glutamate carboxylic acid
- Asparagine peptide lyases - using an asparagine to perform an elimination reaction(not requiring water)
Proteases were first grouped into 84 families according to their evolutionary relationship in 1993, and classified under four catalytic types:
Peptide lyases
A seventh catalytic type of proteolytic enzymes, asparagine peptide lyase, was described in 2011. Its proteolytic mechanism is unusual since, rather than hydrolysis, it performs an elimination reaction.[9] During this reaction, the catalytic asparagine forms a cyclic chemical structure that cleaves itself at asparagine residues in proteins under the right conditions. Given its fundamentally different mechanism, its inclusion as a peptidase may be debatable.[9]
Based on evolutionary phylogeny
An up-to-date classification of protease evolutionary
Currently more than 50 clans are known, each indicating an independent evolutionary origin of proteolysis.[10]
Based on optimal pH
Alternatively, proteases may be classified by the optimal pH in which they are active:
- Acid proteases
- Neutral proteases involved in calpains.
- Basic proteases(or alkaline proteases)
Enzymatic function and mechanism
Proteases are involved in
).Catalysis
Catalysis is achieved by one of two mechanisms:
- Aspartic, glutamic, and metallo-proteases activate a water molecule, which performs a nucleophilic attack on the peptide bond to hydrolyze it.
- Serine, threonine, and cysteine proteases use a nucleophilic residue (usually in a covalentlylink the protease to the substrate protein, releasing the first half of the product. This covalent acyl-enzyme intermediate is then hydrolyzed by activated water to complete catalysis by releasing the second half of the product and regenerating the free enzyme
Specificity
Proteolysis can be highly promiscuous such that a wide range of protein substrates are hydrolyzed. This is the case for digestive enzymes such as trypsin, which have to be able to cleave the array of proteins ingested into smaller peptide fragments. Promiscuous proteases typically bind to a single amino acid on the substrate and so only have specificity for that residue. For example, trypsin is specific for the sequences ...K\... or ...R\... ('\'=cleavage site).[12]
Conversely some proteases are highly specific and only cleave substrates with a certain sequence. Blood clotting (such as thrombin) and viral polyprotein processing (such as TEV protease) requires this level of specificity in order to achieve precise cleavage events. This is achieved by proteases having a long binding cleft or tunnel with several pockets that bind to specified residues. For example, TEV protease is specific for the sequence ...ENLYFQ\S... ('\'=cleavage site).[13]
Degradation and autolysis
Proteases, being themselves proteins, are cleaved by other protease molecules, sometimes of the same variety. This acts as a method of regulation of protease activity. Some proteases are less active after autolysis (e.g. TEV protease) whilst others are more active (e.g. trypsinogen).
Biodiversity of proteases
Proteases occur in all organisms, from
Plants
Plant genomes encode hundreds of proteases, largely of unknown function. Those with known function are largely involved in
Animals
Proteases are used throughout an organism for various metabolic processes. Acid proteases secreted into the stomach (such as
By a complex cooperative action, proteases can catalyze cascade reactions, which result in rapid and efficient amplification of an organism's response to a physiological signal.
Bacteria
Bacteria contain proteases responsible for general protein quality control (e.g. the AAA+
A secreted bacterial protease may also act as an exotoxin, and be an example of a virulence factor in bacterial pathogenesis (for example, exfoliative toxin). Bacterial exotoxic proteases destroy extracellular structures.
Viruses
The genomes of some
Archaea
Archaea use proteases to regulate various cellular processes from cell-signaling, metabolism, secretion and protein quality control.[21][22] Only two ATP-dependent proteases are found in archaea: the membrane associated LonB protease and a soluble 20S proteosome complex .[21]
Uses
The field of protease research is enormous. Since 2004, approximately 8000
Digestive proteases are part of many
Inhibitors
The activity of proteases is inhibited by
Natural protease inhibitors include the family of
Other natural protease inhibitors are used as defense mechanisms. Common examples are the trypsin inhibitors found in the seeds of some plants, most notable for humans being soybeans, a major food crop, where they act to discourage predators. Raw soybeans are toxic to many animals, including humans, until the protease inhibitors they contain have been denatured.
See also
- Ligase
- Protease
- cysteine-
- serine-
- threonine-
- aspartic-
- glutamic-
- metallo-
- PA clan
- Convergent evolution
- Proteolysis
- Catalytic triad
- The Proteolysis Map
- Proteases in angiogenesis
- Intramembrane proteases
- Protease inhibitor (pharmacology)
- Protease inhibitor (biology)
- TopFIND - database of protease specificity, substrates, products and inhibitors
- MEROPS - Database of protease evolutionary groups
References
- ^ "Proteolytic enzyme | Description, Types, & Functions | Britannica".
- PMID 18650443.
- ^ PMID 24931469.
- ^ PMID 17051221.
- .
To assess the relative proficiencies of enzymes that catalyze the hydrolysis of internal and C-terminal peptide bonds [...]
- PMID 22016395.
- PMID 8439290.
- ^ Sanman, Laura E. (June 2014). "Activity-Based Profiling of Proteases". Annual Review of Biochemistry. 83: 249–273.
- ^ PMID 21832066.
- ^ PMID 19892822.
- ISBN 978-1-4160-2973-1.
- PMID 18067249.
- PMID 23826349.
- PMID 18257708.
- PMID 15535125.
- doi:10.1016/j.soilbio.2006.01.006. Archived from the originalon 2021-04-28. Retrieved 2018-12-29.
- ^ Sims GK, Wander MM (2002). "Proteolytic activity under nitrogen or sulfur limitation". Appl. Soil Ecol. 568: 1–5.
- PMID 12475203.
- PMID 23016690.
- PMID 27090931.
- ^ PMID 25774151.
- PMID 32953999.
- ISBN 978-0-12-079610-6.
- ISBN 978-1-85578-147-4.
- S2CID 84748291.
- PMID 11427378.
- PMID 15060002.
External links
- International Proteolysis Society
- MEROPS - the peptidase database Archived 2006-11-14 at the Wayback Machine
- List of protease inhibitors
- Protease cutting predictor
- List of proteases and their specificities (see also [1] Archived 2011-04-30 at the Wayback Machine)
- Proteolysis MAP from Center for Proteolytic Pathways
- Proteolysis Cut Site database - curated expert annotation from users
- Protease cut sites graphical interface
- TopFIND protease database covering cut sites, substrates and protein termini
- Proteases at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
