Phosphoinositide phospholipase C
This article needs additional citations for verification. (July 2011) |
| Phosphatidylinositol-specific phospholipase C | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| OPM superfamily | 118 | ||||||||
| OPM protein | 1djx | ||||||||
| CDD | cd00137 | ||||||||
| |||||||||
| phosphoinositide phospholipase C | |||||||||
|---|---|---|---|---|---|---|---|---|---|
ExPASy NiceZyme view | | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
Phosphoinositide phospholipase C (PLC, EC 3.1.4.11, triphosphoinositide phosphodiesterase, phosphoinositidase C, 1-phosphatidylinositol-4,5-bisphosphate phosphodiesterase, monophosphatidylinositol phosphodiesterase, phosphatidylinositol phospholipase C, PI-PLC, 1-phosphatidyl-D-myo-inositol-4,5-bisphosphate inositoltrisphosphohydrolase; systematic name 1-phosphatidyl-1D-myo-inositol-4,5-bisphosphate inositoltrisphosphohydrolase) is a family of eukaryotic intracellular enzymes that play an important role in signal transduction processes.
Phospholipase Cs participate in
Reaction and catalytic mechanism
All family members are capable of catalyzing the hydrolysis of PIP2, a
The chemical reaction may be expressed as:
- 1-phosphatidyl-1D-myo-inositol 4,5-bisphosphate + H2O
1D-myo-inositol 1,4,5-trisphosphate + diacylglycerol
PLCs catalyze the reaction in two sequential steps. The first reaction is a
Cell location
Phosphoinositide phospholipase C performs its catalytic function at the
Function

Phospholipase C performs a catalytic mechanism, depleting PIP2 and generating inositol trisphosphate (IP3) and diacylglycerol (DAG).
Depletion of PIP2 inactivates numerous effector molecules in the plasma membrane, most notably PIP2 dependent channels and transporters responsible for setting the cell's membrane potential.[5]
The hydrolytic products also go on to modulate the activity of downstream proteins important for cellular signaling. IP3 is soluble, and diffuses through the cytoplasm and interacts with IP3 receptors on the endoplasmic reticulum, causing the release of calcium and raising the level of intracellular calcium.
Further reading:
DAG remains within the inner leaflet of the plasma membrane due to its hydrophobic character, where it recruits protein kinase C (PKC), which becomes activated in conjunction with binding calcium ions. This results in a host of cellular responses through stimulation of calcium-sensitive proteins such as Calmodulin.
Further reading:
| PI-PLC-Y | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| OPM superfamily | 126 | ||||||||
| OPM protein | 2ptd | ||||||||
| |||||||||
Domain structure
In terms of domain organization, all family members possess homologous X and Y catalytic domains in the form of a distorted Triose Phosphate Isomerase (TIM) barrel with a highly disordered, charged, and flexible intervening linker region. Likewise, all isoforms possess four EF hand domains, and a single C2 domain that flank the X and Y catalytic core. An N-terminal PH domain is present in every family except for the sperm-specific ζ isoform.
Isoenzymes and activation
The phospholipase C family consists of 13 isoenzymes split between six subfamilies, PLC-δ (1,3 & 4), -β(1-4), -γ(
PLC-β
PLC-β(1-4) (120-155kDa) are activated by Gαq subunits through their C2 domain and long C-terminal extension. Gβγ subunits are known to activate the β2 and β3 isozymes only; however, this occurs through the PH domain and/or through interactions with the catalytic domain. The exact mechanism still requires further investigation. The PH domain of β2 and β3 plays a dual role, much like PLC-δ1, by binding to the plasma membrane, as well as being a site of interaction for the catalytic activator. However, PLC-β binds to the lipid surface independent of PIP2 with all isozymes preferring phosphoinositol-3-phosphate or neutral membranes.
Members of the Rho GTPase family (e.g., Rac1, Rac2, Rac3, and
PLC-γ
PLC-γ (120-155kDa) is activated by receptor and non-receptor
PLC-γ2
PLC-γ2 plays a major role in
In B cell signaling,
PLC-δ
The PLC-δ subfamily consists of three family members, δ1, 2, and 3. PLC-δ1 (85kDa) is the most well understood of the three. The enzyme is activated by high calcium levels generated by other PLC family members, and therefore functions as a calcium amplifier within the cell. Binding of its substrate PIP2 to the N-terminal
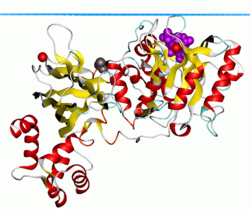
PLC-δ1 also possesses a classical
The widely expressed PLC-δ1 isoform is the best-characterized phospholipase family member, as it was the first to have high-resolution
Other PLC families
- PLC-ε (230-260kDa ) is activated by Ras and RhoGTPases.
- PLC-ζ (75kDa) is thought to play an important role in vertebrate fertilization by producing intracellular calcium oscillations important for the start of embryonic development. However, the mechanism of activation still remains unclear. This isoform is also capable of entering the early-formed pronucleusafter fertilization, which seems to coincide with the cessation of calcium mobilization. It, like PLC-δ1 and PLC-β, possesses nuclear export and localization sequences.
- PLC-η has been implicated in neuronal functioning.
Human proteins in this family
PLCB1; PLCB2; PLCB3; PLCB4; PLCD1; PLCD3; PLCD4; PLCE1; PLCG1; PLCG2; PLCH1; PLCH2; PLCL1; PLCL2; PLCZ1
See also
- Clostridium perfringens alpha toxin
- Lipid signaling
- PH domain, found in some phospholipases C
- Phospholipase
- Zinc-dependent phospholipase C, a different family of phospholipase C
References
- Downes CP, Michell RH (1981). "The polyphosphoinositide phosphodiesterase of erythrocyte membranes". Biochem. J. 198 (1): 133–40. PMID 6275838.
- Thompson W; Dawson RMC (1964). "The triphosphoinositide phosphodiesterase of brain tissue". Biochem. J. 91 (2): 237–243. PMID 4284484.
- Rhee SG, Bae YS (1997). "Regulation of phosphoinositide-specific phospholipase C isozymes". J. Biol. Chem. 272 (24): 15045–8. PMID 9182519.
- Phospholipase+C at the U.S. National Library of Medicine Medical Subject Headings (MeSH)

