Chloroplast DNA
proteins
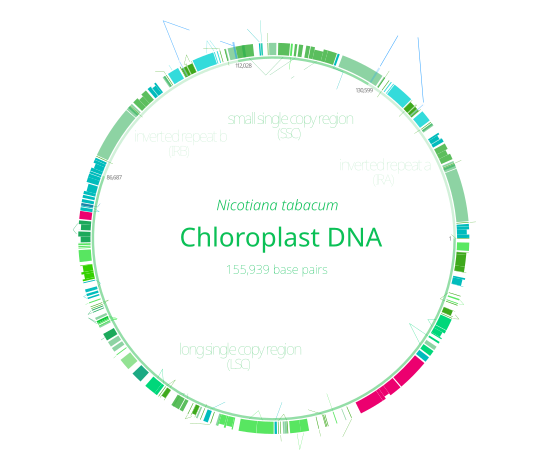
Chloroplast DNA (cpDNA) is the DNA located in chloroplasts, which are photosynthetic organelles located within the cells of some eukaryotic organisms. Chloroplasts, like other types of plastid, contain a genome separate from that in the cell nucleus. The existence of chloroplast DNA was identified biochemically in 1959,[1] and confirmed by electron microscopy in 1962.[2] The discoveries that the chloroplast contains ribosomes[3] and performs protein synthesis[4] revealed that the chloroplast is genetically semi-autonomous. The first complete chloroplast genome sequences were published in 1986, Nicotiana tabacum (tobacco) by Sugiura and colleagues and Marchantia polymorpha (liverwort) by Ozeki et al.[5][6] Since then, a great number of chloroplast DNAs from various species have been sequenced.
Molecular structure
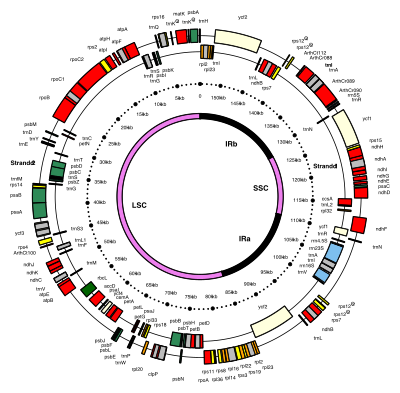
Most chloroplasts have their entire chloroplast genome combined into a single large ring, though those of
Chloroplast DNA has long been thought to have a circular structure, but some evidence suggests that chloroplast DNA more commonly takes a linear shape.
Inverted repeats
Many chloroplast DNAs contain two inverted repeats, which separate a long single copy section (LSC) from a short single copy section (SSC).[9]
The inverted repeats vary wildly in length, ranging from 4,000 to 25,000
The inverted repeat regions are highly
Nucleoids
Each chloroplast contains around 100 copies of its DNA in young leaves, declining to 15–20 copies in older leaves.
Though chloroplast DNA is not associated with true histones,[16] in red algae, a histone-like chloroplast protein (HC) coded by the chloroplast DNA that tightly packs each chloroplast DNA ring into a nucleoid has been found.[17]
In primitive red algae, the chloroplast DNA nucleoids are clustered in the center of a chloroplast, while in green plants and green algae, the nucleoids are dispersed throughout the stroma.[17]
Gene content and plastid gene expression
More than 5000 chloroplast genomes have been sequenced and are accessible via the NCBI organelle genome database.[18] The first chloroplast genomes were sequenced in 1986, from tobacco (Nicotiana tabacum)[19] and liverwort (Marchantia polymorpha).[20] Comparison of the gene sequences of the cyanobacteria Synechocystis to those of the chloroplast genome of Arabidopsis provided confirmation of the endosymbiotic origin of the chloroplast.[21][22] It also demonstrated the significant extent of gene transfer from the cyanobacterial ancestor to the nuclear genome.
In most plant species, the chloroplast genome encodes approximately 120 genes.
Chloroplast genome reduction and gene transfer
Over time, many parts of the chloroplast genome were transferred to the
Endosymbiotic gene transfer is how we know about the
In land plants, some 11–14% of the DNA in their nuclei can be traced back to the chloroplast,[32] up to 18% in Arabidopsis, corresponding to about 4,500 protein-coding genes.[33] There have been a few recent transfers of genes from the chloroplast DNA to the nuclear genome in land plants.[8]
Proteins encoded by the chloroplast
Of the approximately three-thousand proteins found in chloroplasts, some 95% of them are encoded by nuclear genes. Many of the chloroplast's protein complexes consist of subunits from both the chloroplast genome and the host's nuclear genome. As a result,
Protein synthesis
Protein synthesis within chloroplasts relies on an RNA polymerase coded by the chloroplast's own genome, which is related to RNA polymerases found in bacteria. Chloroplasts also contain a mysterious second RNA polymerase that is encoded by the plant's nuclear genome. The two RNA polymerases may recognize and bind to different kinds of promoters within the chloroplast genome.[35] The ribosomes in chloroplasts are similar to bacterial ribosomes.[36]
This section needs expansion with: Genome size differences between algae and land plants, chloroplast stuff coded by the nucleus, DNA replication, NADPH redox, special tRNA synthetases, etc.. You can help by adding to it. (January 2013) |
RNA editing in plastids
RNA editing is the insertion, deletion, and substitution of nucleotides in a mRNA transcript prior to translation to protein. The highly oxidative environment inside chloroplasts increases the rate of mutation so post-transcription repairs are needed to conserve functional sequences. The chloroplast editosome substitutes C -> U and U -> C at very specific locations on the transcript. This can change the codon for an amino acid or restore a non-functional pseudogene by adding an AUG start codon or removing a premature UAA stop codon.[37]
The editosome recognizes and binds to cis sequence upstream of the editing site. The distance between the binding site and editing site varies by gene and proteins involved in the editosome. Hundreds of different
Basal land plants such as liverworts, mosses and ferns have hundreds of different editing sites while flowering plants typically have between thirty and forty. Parasitic plants such as
DNA replication
Leading model of cpDNA replication
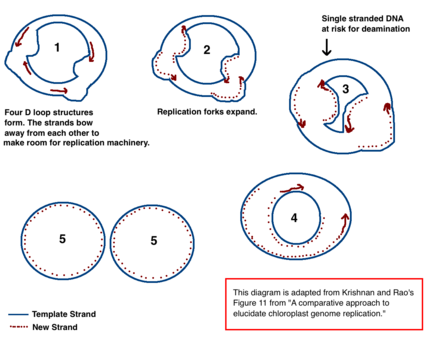
The mechanism for chloroplast DNA (cpDNA) replication has not been conclusively determined, but two main models have been proposed. Scientists have attempted to observe chloroplast replication via
In addition to the early microscopy experiments, this model is also supported by the amounts of

In cpDNA, there are several A → G deamination gradients. DNA becomes susceptible to deamination events when it is single stranded. When replication forks form, the strand not being copied is single stranded, and thus at risk for A → G deamination. Therefore, gradients in deamination indicate that replication forks were most likely present and the direction that they initially opened (the highest gradient is most likely nearest the start site because it was single stranded for the longest amount of time).[39] This mechanism is still the leading theory today; however, a second theory suggests that most cpDNA is actually linear and replicates through homologous recombination. It further contends that only a minority of the genetic material is kept in circular chromosomes while the rest is in branched, linear, or other complex structures.[39][12]
Alternative model of replication
One of the main competing models for cpDNA asserts that most cpDNA is linear and participates in
Protein targeting and import
The movement of so many chloroplast genes to the nucleus means that many chloroplast
Curiously, around half of the protein products of transferred genes aren't even targeted back to the chloroplast. Many became
Because the cell acquiring a chloroplast
Cytoplasmic translation and N-terminal transit sequences
Chloroplast transit peptides exhibit huge variation in length and
Not all chloroplast proteins include a N-terminal cleavable transit peptide though.
Phosphorylation, chaperones, and transport
After a chloroplast
Phosphorylation changes the polypeptide's shape,
The heat shock protein and the 14-3-3 proteins together form a cytosolic guidance complex that makes it easier for the chloroplast polypeptide to get imported into the chloroplast.[42]
Alternatively, if a chloroplast preprotein's transit peptide is not phosphorylated, a chloroplast preprotein can still attach to a heat shock protein or
The translocon on the outer chloroplast membrane (TOC)
The
The first three proteins form a core complex that consists of one Toc159, four to five Toc34s, and four Toc75s that form four holes in a disk 13

