Poliovirus
| Poliovirus | |
|---|---|

| |
| TEM micrograph of poliovirus virions. Scale bar (white): 50 nm | |

| |
| A type 3 poliovirus capsid, protein side chains coloured | |
| Virus classification | |
| (unranked): | Virus |
| Realm: | Riboviria |
| Kingdom: | Orthornavirae |
| Phylum: | Pisuviricota |
| Class: | Pisoniviricetes |
| Order: | Picornavirales |
| Family: | Picornaviridae
|
| Genus: | Enterovirus |
| Species: | |
| Virus: | Poliovirus
|
| Serotypes | |
| |
Poliovirus, the causative agent of
Poliovirus is composed of an
Poliovirus was first isolated in 1909 by
In 1981, the poliovirus genome was published by two different teams of researchers: by Vincent Racaniello and David Baltimore at MIT[8] and by Naomi Kitamura and Eckard Wimmer at Stony Brook University.[9]
The three-dimensional structure of poliovirus was determined in 1985 by
Poliovirus is one of the most well-characterized viruses, and has become a useful model system for understanding the biology of
Replication cycle
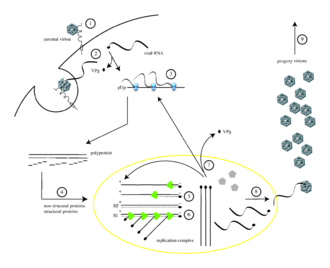
Poliovirus infects human cells by binding to an
Poliovirus is a positive-stranded RNA virus. Thus, the genome enclosed within the viral particle can be used as messenger RNA and immediately translated by the host cell. On entry, the virus hijacks the cell's translation machinery, causing inhibition of cellular protein synthesis in favor of virus-specific protein production.[19] Unlike the host cell's mRNAs, the 5' end of poliovirus RNA is extremely long—over 700 nucleotides—and highly structured. This region of the viral genome is called an internal ribosome entry site (IRES). This region consists of many secondary structures and 3 or 4 domains. Domain 3 is a self folding RNA element that contains conserved structural motifs in various stable stem loops linked by two four-way junctions. As IRES consists of many domains, these domains themselves consist of many loops that contribute to modified translation without a 5’ end cap by hijacking ribosomes. The interaction loop of domain 3 is known as GNRA tetraloop. The residues of adenosines A180 and A181 in the GUAA tetraloop form hydrogen bonds via non canonical base pairing interactions with the base pairs of the receptors C230/G242 and G231/C241, respectively.[20] Genetic mutations in this region prevent viral protein production.[21][22][23] The first IRES to be discovered was found in poliovirus RNA.[24]
Poliovirus mRNA is translated as one long

- 3Dpol, an RNA dependent RNA polymerasewhose function is to make multiple copies of the viral RNA genome
- 2Apro and 3Cpro/3CDpro, proteases which cleave the viral polypeptide
- VPg (3B), a small protein that binds viral RNA and is necessary for synthesis of viral positive and negative strand RNA
- 2BC, 2B, 2C (an ATPase)[28], 3AB, 3A, 3B proteins which comprise the protein complex needed for virus replication.
- VP0, which is further cleaved into VP2 and VP4, VP1 and VP3, proteins of the viral capsid
After translation, transcription and genome replication which involve a single process, synthesis of (+) RNA) is realized. For the infecting (+)RNA to be replicated, multiple copies of (−)RNA must be transcribed and then used as templates for (+)RNA synthesis. Replicative intermediates (RIs), which are an association of RNA molecules consisting of a template RNA and several growing RNAs of varying length, are seen in both the replication complexes for (−)RNAs and (+)RNAs. For synthesis of each negative-strand and positive-strand RNAs, VPg protein in the poliovirus works as a primer. RNA-dependent RNA polymerase of the poliovirus adds two uracil nucleotides (UU) to VPg protein utilizing the poly(A) tail at the 3′-end of the +ssRNA genome as a pattern for synthesis of the negative-strand antigenomic RNA. To initiate this −ssRNA synthesis, the tyrosine hydroxyl of VPg is needed. But for the initiation of positive strand RNA synthesis, CRE-dependent VPg uridylylation is needed. Which means that VPg is once more utilized as a primer however this time it adds the two uridine triphosphates using a cis-acting replication element (CRE) as a template.[29][30]
The CRE of poliovirus is identified as an unachieved base-paired stem and a final loop consisting of 61 nt. The CRE is found in enteroviruses. It is a highly preserved secondary RNA structural element and bedded in the genome's polyprotein-coding region. The complex can be translocated to the 5' region of the genome that have no coding activity, at least 3.7-kb distant from the initial location. This process can occurs without negatively influencing activity. CRE copies do not influence replication negatively. Uridylylation process of VPg that takes place at CRE needs the presence of 3CDpro that is an RNA binding protein. It is attached to the CRE directly and specifically. Because of its presence VPg can bind the CRE properly and primary production proceeds without problems.[31]
Some of the (+) RNA molecules are used as templates for further (−) RNA synthesis, some function as mRNA, and some are destined to be the genomes of progeny virions.[25]
In the assembly of new virus particles (i.e. the packaging of progeny genome into a procapsid which can survive outside the host cell), including, respectively:[32]
- Five copies each of VP0, VP3, and VP1 whose N termini and VP4 form interior surface of capsid, assemble into a 'pentamer' and 12 pentamers form a procapsid. (The outer surface of capsid is consisting of VP1, VP2, VP3; C termini of VP1 and VP3 form the canyons which around each of the vertices; around this time, the 60 copies of VP0 are cleaved into VP4 and VP2.)
- Each procapsid acquires a copy of the virus genome, with VPg still attached at the 5' end.
Fully assembled poliovirus leaves the confines of its host cell by
Drake demonstrated that poliovirus is able to undergo multiplicity reactivation.
Origin and serotypes
Poliovirus is structurally similar to other human enteroviruses (
The mutation rate in the virus is relatively high even for an RNA virus with a synonymous substitution rate of 1.0 x 10−2 substitutions/site/year and non synonymous substitution rate of 3.0 x 10−4 substitutions/site/year.
The three
Specific strains of each serotype are used to prepare
Polioviruses were formerly classified as a distinct species belonging to the genus Enterovirus in the family Picornaviridae. In 2008, the Poliovirus species was eliminated and the three serotypes were assigned to the species Human enterovirus C (later renamed Enterovirus C), in the genus Enterovirus in the family Picornaviridae. The type species of the genus Enterovirus was changed from Poliovirus to (Human) Enterovirus C.[45]
Pathogenesis
The primary determinant of infection for any virus is its ability to enter a cell and produce additional infectious particles. The presence of CD155 is thought to define the animals and tissues that can be infected by poliovirus. CD155 is found (outside of laboratories) only on the cells of humans, higher
The CD155 gene appears to have been subject to
Poliovirus is an
In many respects, the
Poliomyelitis is a disease of the central nervous system. However, CD155 is believed to be present on the surface of most or all human cells. Therefore, receptor expression does not explain why poliovirus preferentially infects certain tissues. This suggests that tissue tropism is determined after cellular infection. Recent work has suggested that the type I interferon response (specifically that of interferon alpha and beta) is an important factor that defines which types of cells support poliovirus replication.[56] In mice expressing CD155 (through genetic engineering) but lacking the type I interferon receptor, poliovirus not only replicates in an expanded repertoire of tissue types, but these mice are also able to be infected orally with the virus.[57]
Immune system avoidance

Poliovirus uses two key mechanisms to evade the
Individuals who are exposed to poliovirus, either through infection or by
PVR transgenic mouse
Although humans are the only known natural hosts of poliovirus, monkeys can be experimentally infected and they have long been used to study poliovirus. In 1990–91, a small animal model of poliomyelitis was developed by two laboratories. Mice were engineered to express a human receptor to poliovirus (hPVR).[61][62]
Unlike normal mice,
Three distinct types of TgPVR mice have been well studied:[65]
- In TgPVR1 mice, the transgene encoding the human PVR was incorporated into mouse chromosome 4. These mice express the highest levels of the transgene and the highest sensitivity to poliovirus. TgPVR1 mice are susceptible to poliovirus through the intraspinal, intracerebral, intramuscular, and intravenous pathways, but not through the oral route.
- TgPVR21 mice have incorporated the human PVR at chromosome 13. These mice are less susceptible to poliovirus infection through the intracerebral route, possibly because they express decreased levels of hPVR. TgPVR21 mice have been shown to be susceptible to poliovirus infection through intranasal inoculation, and may be useful as a mucosal infection model.[66]
- In TgPVR5 mice, the human transgene is located on chromosome 12. These mice exhibit the lowest levels of hPVR expression and are the least susceptible to poliovirus infection.
Recently, a fourth TgPVR mouse model was developed. These "cPVR" mice carry hPVR
The development of the TgPVR mouse has had a profound effect on oral
Cloning and synthesis

In 1981, Racaniello and Baltimore used recombinant DNA technology to generate the first infectious clone of an animal RNA virus, poliovirus. DNA encoding the RNA genome of poliovirus was introduced into cultured mammalian cells and infectious poliovirus was produced.[68] Creation of the infectious clone propelled understanding of poliovirus biology, and has become a standard technology used to study many other viruses.
In 2002,
Modification for therapies
A modification of the poliovirus, called PVSRIPO, was tested in early clinical trials as a possible treatment for cancer.[71][needs update]
References
- ISBN 978-0-8385-8529-0.[page needed]
- ^ PMID 12142481.
- ^ ISBN 978-0-387-98273-1.[page needed]
- ^ ISBN 978-0-300-01324-5.[page needed]
- ^ "Behind the picture: Rosalind Franklin and the polio model". Medical Research Council. 2019-03-14. Archived from the original on 2018-10-30. Retrieved 4 September 2019.
- ISBN 0-00-655211-0. Archivedfrom the original on 2023-01-14. Retrieved 2020-05-15.
- ISBN 978-0-19-920565-3.
- PMID 6272282.
- S2CID 4352308.
- PMID 2994218.
- ^ PMID 17623069.
- S2CID 44296539.
- ^ PMID 12663789.
- PMID 4309884.
- PMID 2168959.
- PMID 8212594.
- ^ PMID 15885840.
- PMID 17622193.
- PMID 13965389.
- PMID 26778619.
- PMID 7536344.
- S2CID 4327857.
- PMID 2839690.
- ISBN 978-0-470-02386-0.
- ^ ISBN 978-0-470-02386-0. Archivedfrom the original on 2023-01-14. Retrieved 2016-10-19.
- ISBN 978-0-8153-4150-5.[page needed]
- ^ "Poliomyelitis". Brown University. Archived from the original on February 22, 2007.
- ISBN 978-0-470-02386-0.
- ISBN 978-0-12-800947-5.
- PMID 12663781.
- PMID 12554882.
- ISBN 978-0-470-02386-0. Archivedfrom the original on 2023-01-14. Retrieved 2016-10-19.
- ISBN 978-0-470-02386-0.
- ^ from the original on 2020-07-09. Retrieved 2018-10-26.
- PMID 13581529.
- PMID 3021340.
- PMID 20335491.
- PMID 29111273.
- PMID 17517601.
- PMID 18287242.
- PMID 6275352.
- PMID 21450075.
- ^ "Global eradication of wild poliovirus type 2 declared". Global Polio Eradication Initiative. 2015-09-20. Archived from the original on 2015-09-28. Retrieved 2015-09-30.
- ^ "GPEI-Two out of three wild poliovirus strains eradicated". Archived from the original on 2019-11-07. Retrieved 2019-11-02.
- PMID 19495937.
- PMID 12759359.
- PMID 16500041.
- ^ Bodian D, Horstmann DH (1969). Polioviruse. Philadelphia, Penn: Lippincott. pp. 430–73.
- PMID 13337331.
- OCLC 124160564.[page needed]
- ^ Acute Poliomyelitis at eMedicine
Pediatric Poliomyelitis at eMedicine - PMID 9126254.
- PMID 9770421.
- PMID 1326581.
- PMID 20221252.
- PMID 15767446.
- PMID 17507470.
- PMID 16364730.
- ISBN 978-0-470-02386-0.[page needed]
- ^ "WHO | Poliomyelitis". Archived from the original on May 17, 2014.
- S2CID 11946113.
- PMID 1846972.
- PMID 8289371.
- PMID 11597452.
- S2CID 11070024.
- ^ PMID 15033568.
- from the original on 2021-02-24. Retrieved 2022-04-01.
- PMID 6272391.
- S2CID 5810309.
- S2CID 83531627.
- PMID 24939611.
External links
- ICTVdb virus classification 2006
- Home of Picornaviruses (latest updates of species, serotypes, & proposed changes) Archived 2010-11-04 at the Wayback Machine
- Goodsell D. "Poliovirus and Rhinovirus". August 2001 Molecule of the Month. Archived from the original on 2011-03-03. Retrieved 2010-01-07.
- 3D macromolecular structures of the Poliovirus archived in the EM Data Bank(EMDB)
- "Human poliovirus 1". NCBI Taxonomy Browser. 12080.
- "Human poliovirus 3". NCBI Taxonomy Browser. 12086.
