Polyadenylation

Polyadenylation is the addition of a poly(A) tail to an RNA transcript, typically a messenger RNA (mRNA). The poly(A) tail consists of multiple adenosine monophosphates; in other words, it is a stretch of RNA that has only adenine bases. In eukaryotes, polyadenylation is part of the process that produces mature mRNA for translation. In many bacteria, the poly(A) tail promotes degradation of the mRNA. It, therefore, forms part of the larger process of gene expression.
The process of polyadenylation begins as the
The poly(A) tail is important for the nuclear export, translation and stability of mRNA. The tail is shortened over time, and, when it is short enough, the mRNA is enzymatically degraded.[2] However, in a few cell types, mRNAs with short poly(A) tails are stored for later activation by re-polyadenylation in the cytosol.[3] In contrast, when polyadenylation occurs in bacteria, it promotes RNA degradation.[4] This is also sometimes the case for eukaryotic non-coding RNAs.[5][6]
mRNA molecules in both prokaryotes and eukaryotes have polyadenylated 3′-ends, with the prokaryotic poly(A) tails generally shorter and fewer mRNA molecules polyadenylated.[7]
Background on RNA
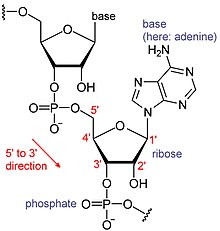
RNAs are a type of large biological molecules, whose individual building blocks are called nucleotides. The name poly(A) tail (for polyadenylic acid tail)
Messenger RNA (mRNA) is RNA that has a coding region that acts as a template for protein synthesis (translation). The rest of the mRNA, the untranslated regions, tune how active the mRNA is.[10] There are also many RNAs that are not translated, called non-coding RNAs. Like the untranslated regions, many of these non-coding RNAs have regulatory roles.[11]
Nuclear polyadenylation
Function
In nuclear polyadenylation, a poly(A) tail is added to an RNA at the end of transcription. On mRNAs, the poly(A) tail protects the mRNA molecule from enzymatic degradation in the cytoplasm and aids in transcription termination, export of the mRNA from the nucleus, and translation.[2] Almost all eukaryotic mRNAs are polyadenylated,[12] with the exception of animal replication-dependent histone mRNAs.[13] These are the only mRNAs in eukaryotes that lack a poly(A) tail, ending instead in a stem-loop structure followed by a purine-rich sequence, termed histone downstream element, that directs where the RNA is cut so that the 3′ end of the histone mRNA is formed.[14]
Many eukaryotic non-coding RNAs are always polyadenylated at the end of transcription. There are small RNAs where the poly(A) tail is seen only in intermediary forms and not in the mature RNA as the ends are removed during processing, the notable ones being
Mechanism
| Proteins involved:[12][18] CPSF: cleavage/polyadenylation specificity factor CstF : cleavage stimulation factorPAP: polyadenylate polymerase PABII: polyadenylate binding protein 2 CFI: cleavage factor I CFII: cleavage factor II |
The
The RNA is typically cleaved before transcription termination, as CstF also binds to RNA polymerase II.[28] Through a poorly understood mechanism (as of 2002), it signals for RNA polymerase II to slip off of the transcript.[29] Cleavage also involves the protein CFII, though it is unknown how.[30] The cleavage site associated with a polyadenylation signal can vary up to some 50 nucleotides.[31]
When the RNA is cleaved, polyadenylation starts, catalysed by polyadenylate polymerase.
Downstream effects
The poly(A) tail acts as the binding site for
Deadenylation
In eukaryotic
In animals, poly(A) ribonuclease (
Cytoplasmic polyadenylation
There is polyadenylation in the cytosol of some animal cell types, namely in the
In the early mouse embryo, cytoplasmic polyadenylation of maternal RNAs from the egg cell allows the cell to survive and grow even though transcription does not start until the middle of the 2-cell stage (4-cell stage in human).[51][52] In the brain, cytoplasmic polyadenylation is active during learning and could play a role in long-term potentiation, which is the strengthening of the signal transmission from a nerve cell to another in response to nerve impulses and is important for learning and memory formation.[3][53]
Cytoplasmic polyadenylation requires the RNA-binding proteins CPSF and CPEB, and can involve other RNA-binding proteins like Pumilio.[54] Depending on the cell type, the polymerase can be the same type of polyadenylate polymerase (PAP) that is used in the nuclear process, or the cytoplasmic polymerase GLD-2.[55]
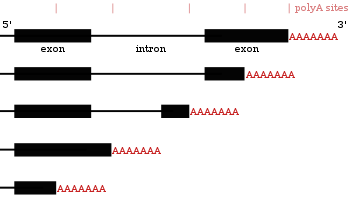
Alternative polyadenylation
Many protein-coding genes have more than one polyadenylation site, so a gene can code for several mRNAs that differ in their
Since alternative polyadenylation changes the length of the 3' UTR,[62] it can also change which binding sites are available for microRNAs in the 3′ UTR.[19][63] MicroRNAs tend to repress translation and promote degradation of the mRNAs they bind to, although there are examples of microRNAs that stabilise transcripts.[64][65] Alternative polyadenylation can also shorten the coding region, thus making the mRNA code for a different protein,[66][67] but this is much less common than just shortening the 3′ untranslated region.[27]
The choice of poly(A) site can be influenced by extracellular stimuli and depends on the expression of the proteins that take part in polyadenylation.
Tagging for degradation in eukaryotes
For many
In prokaryotes and organelles

In many bacteria, both mRNAs and non-coding RNAs can be polyadenylated. This poly(A) tail promotes degradation by the
In as different groups as animals and
While many bacteria and mitochondria have polyadenylate polymerases, they also have another type of polyadenylation, performed by polynucleotide phosphorylase itself. This enzyme is found in bacteria,[85] mitochondria,[86] plastids[87] and as a constituent of the archaeal exosome (in those archaea that have an exosome).[88] It can synthesise a 3′ extension where the vast majority of the bases are adenines. Like in bacteria, polyadenylation by polynucleotide phosphorylase promotes degradation of the RNA in plastids[89] and likely also archaea.[83]
Evolution
Although polyadenylation is seen in almost all organisms, it is not universal.
The most ancient polyadenylating enzyme is polynucleotide phosphorylase. This enzyme is part of both the bacterial degradosome and the archaeal exosome,[93] two closely related complexes that recycle RNA into nucleotides. This enzyme degrades RNA by attacking the bond between the 3′-most nucleotides with a phosphate, breaking off a diphosphate nucleotide. This reaction is reversible, and so the enzyme can also extend RNA with more nucleotides. The heteropolymeric tail added by polynucleotide phosphorylase is very rich in adenine. The choice of adenine is most likely the result of higher ADP concentrations than other nucleotides as a result of using ATP as an energy currency, making it more likely to be incorporated in this tail in early lifeforms. It has been suggested that the involvement of adenine-rich tails in RNA degradation prompted the later evolution of polyadenylate polymerases (the enzymes that produce poly(A) tails with no other nucleotides in them).[94]
Polyadenylate polymerases are not as ancient. They have separately evolved in both bacteria and eukaryotes from CCA-adding enzyme, which is the enzyme that completes the 3′ ends of tRNAs. Its catalytic domain is homologous to that of other polymerases.[79] It is presumed that the horizontal transfer of bacterial CCA-adding enzyme to eukaryotes allowed the archaeal-like CCA-adding enzyme to switch function to a poly(A) polymerase.[82] Some lineages, like archaea and cyanobacteria, never evolved a polyadenylate polymerase.[94]
Polyadenylate tails are observed in several
History
Poly(A)polymerase was first identified in 1960 as an enzymatic activity in extracts made from cell nuclei that could polymerise ATP, but not ADP, into polyadenine.[100][101] Although identified in many types of cells, this activity had no known function until 1971, when poly(A) sequences were found in mRNAs.[102][103] The only function of these sequences was thought at first to be protection of the 3′ end of the RNA from nucleases, but later the specific roles of polyadenylation in nuclear export and translation were identified. The polymerases responsible for polyadenylation were first purified and characterized in the 1960s and 1970s, but the large number of accessory proteins that control this process were discovered only in the early 1990s.[102]
See also
References
- ^ S2CID 478260.
- ^ PMID 11255003.
- ^ PMID 10357857.
- PMID 10943888.
- S2CID 19863143.
- S2CID 19003617.
- ^ PMID 9242905.
- PMID 14140701.
- ISBN 978-0-87901-500-8.[page needed]
- PMID 18212021.
- PMID 16651366.
- ^ PMID 18479511.
- ^ PMID 17998288.
- PMID 12408966.
- PMID 17965236.
- PMID 16131612.
- S2CID 206956408.
- ^ PMID 8440247.
- ^ PMID 17158511.
- PMID 18817380.
- ^ PMID 10899149.
- PMID 14690600.
- PMID 20479262.
- PMID 21295486.
- PMID 15937220.
- ^ PMID 17024186.
- ^ PMID 18411206.
- PMID 18157150.
- ^ Molecular Biology of the Cell, Chapter 6, "From DNA to RNA". 4th edition. Alberts B, Johnson A, Lewis J, et al. New York: Garland Science; 2002.
- S2CID 34840144.
- PMID 12097343.
- PMID 17850751.
- PMID 18304944.
- PMID 7852352.
- PMID 12145212.
- S2CID 5777074.
- S2CID 2237085.
- PMID 9784497.
- ^ PMID 17595167.
- PMID 15145353.
- PMID 10970864.
- PMID 16714281.
- PMID 28334977.
- ^ PMID 17933768.
- PMID 15231747.
- PMID 16495412.
- PMID 18430932.
- S2CID 231195473.
- S2CID 9734550.
- PMID 16705177.
- PMID 16169522.
- PMID 18023855.
- PMID 17481902.
- S2CID 16092673.
- PMID 18434412.
- PMID 15647503.
- PMID 18256699.
- ^ PMID 27677860.
- PMID 16356263.
- PMID 22685694.
- PMID 18757892.
- PMID 27220521.
- PMID 18566288.
- PMID 18392144.
- PMID 18835850.
- S2CID 7448467.
- PMID 17210931.
- ^ PMID 16207706.
- PMID 30552333.
- PMID 18978773.
- PMID 17507659.
- PMID 17464285.
- PMID 21292162.
- PMID 18451104.
- PMID 32976578.
- PMID 17395456.
- PMID 21663793.
- S2CID 14898055.
- ^ PMID 17872511.
- PMID 16738135.
- S2CID 26109164.
- ^ PMID 11917006.
- ^ S2CID 86607431.
- PMID 22172994.
- PMID 17965156.
- PMID 18312863.
- PMID 12486011.
- PMID 17065466.
- PMID 12953107.
- PMID 19161858.
- PMID 16282984.
- PMID 18399989.
- PMID 19053279.
- ^ PMID 18177749.
- PMID 10074205.
- PMID 23923003.
- PMID 11717411.
- PMID 30319572.
- ^ "Inhibition of host poly(A)-binding protein by virus ~ ViralZone". viralzone.expasy.org.
- .
- PMID 9353246.
- ^ PMID 12102557.
- PMID 5288383.
Further reading
- Danckwardt S, Hentze MW, Kulozik AE (February 2008). "3′ end mRNA processing: molecular mechanisms and implications for health and disease". The EMBO Journal. 27 (3): 482–98. PMID 18256699.
External links
 Media related to Polyadenylation at Wikimedia Commons
Media related to Polyadenylation at Wikimedia Commons
