Porphobilinogen deaminase
| HMBS | |||
|---|---|---|---|
Gene ontology | |||
| Molecular function | |||
| Cellular component | |||
| Biological process | |||
| Sources:Amigo / QuickGO | |||
Ensembl | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| UniProt | |||||||||
| RefSeq (mRNA) | |||||||||
| RefSeq (protein) | |||||||||
| Location (UCSC) | Chr 11: 119.08 – 119.09 Mb | Chr 9: 44.25 – 44.26 Mb | |||||||
| PubMed search | [3] | [4] | |||||||
| View/Edit Human | View/Edit Mouse |
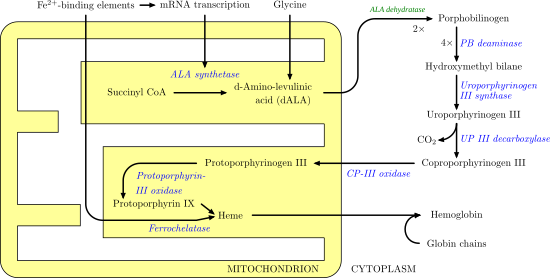
Porphobilinogen deaminase (hydroxymethylbilane synthase, or uroporphyrinogen I synthase) is an enzyme (EC 2.5.1.61) that in humans is encoded by the HMBS gene. Porphobilinogen deaminase is involved in the third step of the heme biosynthetic pathway. It catalyzes the head to tail condensation of four porphobilinogen molecules into the linear hydroxymethylbilane while releasing four ammonia molecules:
- 4 porphobilinogen + H2O hydroxymethylbilane + 4 NH3
Structure and function
Functionally, porphobilinogen deaminase catalyzes the loss of ammonia from the porphobilinogen monomer (deamination) and its subsequent polymerization to a linear tetrapyrrole, which is released as hydroxymethylbilane:

The structure of 40-42 kDa porphobilinogen deaminase, which is highly conserved amongst organisms, consists of three domains.[5][6] Domains 1 and 2 are structurally very similar: each consisting of five beta-sheets and three alpha helices in humans.[7] Domain 3 is positioned between the other two and has a flattened beta-sheet geometry. A dipyrrole, a cofactor of this enzyme consisting of two condensed porphobilinogen molecules, is covalently attached to domain 3 and extends into the active site, the cleft between domains 1 and 2.[8] Several positively charged arginine residues, positioned to face the active site from domains 1 and 2, have been shown to stabilize the carboxylate functionalities on the incoming porphobilinogen as well as the growing pyrrole chain. These structural features presumably favor the formation of the final hydroxymethylbilane product.[9] Porphobilinogen deaminase usually exists in dimer units in the cytoplasm of the cell.
Reaction mechanism

The first step is believed to involve an
Pathology
The most well-known health issue involving porphobilinogen deaminase is acute intermittent porphyria, an autosomal dominant genetic disorder where insufficient hydroxymethylbilane is produced, leading to a build-up of porphobilinogen in the cytoplasm. This is caused by a gene mutation that, in 90% of cases, causes decreased amounts of enzyme. However, mutations where less-active enzymes and/or different isoforms have been described.[13][14][15] At least 115 disease-causing mutations in this gene have been discovered.[16]
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000256269 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000032126 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- PMID 2609111.
- S2CID 4264432.
- PMID 19207107.
- S2CID 13483654.
- PMID 2025226.
- .
- PMID 11152419.
- PMID 2664584.
- ^ "Entrez Gene: HMBS hydroxymethylbilane synthase".
- PMID 2789372.
- S2CID 24402776.
- PMID 31819097.
Further reading
- Deybach JC, Puy H (1995). "Porphobilinogen deaminase gene structure and molecular defects". J. Bioenerg. Biomembr. 27 (2): 197–205. S2CID 41764609.
- Astrin KH, Desnick RJ (1995). "Molecular basis of acute intermittent porphyria: mutations and polymorphisms in the human hydroxymethylbilane synthase gene". Hum. Mutat. 4 (4): 243–52. S2CID 24402776.
- Helliwell JR, Nieh YP, Habash J, et al. (2003). "Time-resolved and static-ensemble structural chemistry of hydroxymethylbilane synthase". Faraday Discussions. 122: 131–44, discussion 171–90. PMID 12555854.
- Hessels J, Voortman G, van der Wagen A, et al. (2004). "Homozygous acute intermittent porphyria in a 7-year-old boy with massive excretions of porphyrins and porphyrin precursors". J. Inherit. Metab. Dis. 27 (1): 19–27. S2CID 9504522.
- Kauppinen R (2004). "Molecular diagnostics of acute intermittent porphyria". Expert Rev. Mol. Diagn. 4 (2): 243–9. S2CID 21392441.
- Hrdinka M, Puy H, Martasek P (2007). "May 2006 update in porphobilinogen deaminase gene polymorphisms and mutations causing acute intermittent porphyria: comparison with the situation in Slavic population". Physiological Research. 55 (Suppl 2): S119–36. PMID 17298216.
- Kauppinen R, Peltonen L, Pihlaja H, Mustajoki P (1993). "CRIM-positive mutations of acute intermittent porphyria in Finland". Hum. Mutat. 1 (5): 392–6. S2CID 10575526.
- Mgone CS, Lanyon WG, Moore MR, Connor JM (1992). "Detection of seven point mutations in the porphobilinogen deaminase gene in patients with acute intermittent porphyria, by direct sequencing of in vitro amplified cDNA". Hum. Genet. 90 (1–2): 12–6. S2CID 19680295.
- Gu XF, de Rooij F, Voortman G, et al. (1992). "High frequency of mutations in exon 10 of the porphobilinogen deaminase gene in patients with a CRIM-positive subtype of acute intermittent porphyria". Am. J. Hum. Genet. 51 (3): 660–5. PMID 1496994.
- Delfau MH, Picat C, De Rooij F, et al. (1991). "Molecular heterogeneity of acute intermittent porphyria: identification of four additional mutations resulting in the CRIM-negative subtype of the disease". Am. J. Hum. Genet. 49 (2): 421–8. PMID 1714233.
- Namba H, Narahara K, Tsuji K, et al. (1991). "Assignment of human porphobilinogen deaminase to 11q24.1----q24.2 by in situ hybridization and gene dosage studies". Cytogenet. Cell Genet. 57 (2–3): 105–8. PMID 1914516.
- Lee JS, Anvret M (1992). "Identification of the most common mutation within the porphobilinogen deaminase gene in Swedish patients with acute intermittent porphyria". Proc. Natl. Acad. Sci. U.S.A. 88 (23): 10912–5. PMID 1961762.
- Tunnacliffe A, McGuire RS (1991). "A physical linkage group in human chromosome band 11q23 covering a region implicated in leukocyte neoplasia". Genomics. 8 (3): 447–53. PMID 1981047.
- Scobie GA, Llewellyn DH, Urquhart AJ, et al. (1990). "Acute intermittent porphyria caused by a C----T mutation that produces a stop codon in the porphobilinogen deaminase gene". Hum. Genet. 85 (6): 631–4. S2CID 28858687.
- Delfau MH, Picat C, de Rooij FW, et al. (1990). "Two different point G to A mutations in exon 10 of the porphobilinogen deaminase gene are responsible for acute intermittent porphyria". J. Clin. Invest. 86 (5): 1511–6. PMID 2243128.
- Raich N, Romeo PH, Dubart A, et al. (1986). "Molecular cloning and complete primary sequence of human erythrocyte porphobilinogen deaminase". Nucleic Acids Res. 14 (15): 5955–68. PMID 2875434.
- Vidaud M, Gattoni R, Stevenin J, et al. (1989). "A 5' splice-region G----C mutation in exon 1 of the human beta-globin gene inhibits pre-mRNA splicing: a mechanism for beta+-thalassemia". Proc. Natl. Acad. Sci. U.S.A. 86 (3): 1041–5. PMID 2915972.
External links