Sense (molecular biology)
In
DNA sense
Because of the
Sometimes the phrases coding strand and template strand are encountered in place of sense and antisense, respectively, and in the context of a double-stranded DNA molecule the usage of these terms is essentially equivalent. However, the coding/sense strand need not always contain a code that is used to make a protein; both protein-coding and non-coding RNAs may be transcribed.
The terms "sense" and "antisense" are relative only to the particular RNA transcript in question, and not to the DNA strand as a whole. In other words, either DNA strand can serve as the sense or antisense strand. Most organisms with sufficiently large genomes make use of both strands, with each strand functioning as the template strand for different RNA transcripts in different places along the same DNA molecule. In some cases, RNA transcripts can be transcribed in both directions (i.e. on either strand) from a common
Sense DNA
The DNA sense strand looks like the
Hence, a base triplet 3′-TAC-5′ in the DNA antisense strand (complementary to the 5′-ATG-3′ of the DNA sense strand) is used as the template which results in a 5′-AUG-3′ base triplet in the mRNA. The DNA sense strand will have the triplet ATG, which looks similar to the mRNA triplet AUG but will not be used to make methionine because it will not be directly used to make mRNA. The DNA sense strand is called a "sense" strand not because it will be used to make protein (it won't be), but because it has a sequence that corresponds directly to the RNA codon sequence. By this logic, the RNA transcript itself is sometimes described as "sense".
Example with double-stranded DNA
- DNA strand 1: antisense strand (transcribed to) → RNA strand (sense)
- DNA strand 2: sense strand
Some regions within a double-stranded DNA molecule code for genes, which are usually instructions specifying the order in which amino acids are assembled to make proteins, as well as regulatory sequences, splicing sites, non-coding introns, and other gene products. For a cell to use this information, one strand of the DNA serves as a template for the synthesis of a complementary strand of RNA. The transcribed DNA strand is called the template strand, with antisense sequence, and the mRNA transcript produced from it is said to be sense sequence (the complement of antisense). The untranscribed DNA strand, complementary to the transcribed strand, is also said to have sense sequence; it has the same sense sequence as the mRNA transcript (though T bases in DNA are substituted with U bases in RNA).
3′CGCTATAGCGTTT 5′ |
DNA antisense strand (template/noncoding) | Used as a template for transcription. |
5′GCGATATCGCAAA 3′ |
DNA sense strand (nontemplate/coding) | Complementary to the template strand. |
5′GCGAUAUCGCAAA 3′ |
mRNA sense transcript | RNA strand that is transcribed from the noncoding (template/antisense) strand. Note1: Except for the fact that all thymines are now uracils (T → U), it is complementary to the noncoding (template/antisense) DNA strand and identical to the coding (nontemplate/sense) DNA strand.
|
3′CGCUAUAGCGUUU 5′ |
mRNA antisense transcript | RNA strand that is transcribed from the coding (nontemplate/sense) strand. Note: Except for the fact that all thymines are now uracils (T → U), it is complementary to the coding (nontemplate/sense) DNA strand and identical to the noncoding (template/antisense) DNA strand.
|
The names assigned to each strand actually depend on which direction you are writing the sequence that contains the information for proteins (the "sense" information), not on which strand is depicted as "on the top" or "on the bottom" (which is arbitrary). The only biological information that is important for labeling strands is the relative locations of the terminal 5′ phosphate group and the terminal 3′ hydroxyl group (at the ends of the strand or sequence in question), because these ends determine the direction of transcription and translation. A sequence written 5′-CGCTAT-3′ is equivalent to a sequence written 3′-TATCGC-5′ as long as the 5′ and 3′ ends are noted. If the ends are not labeled, convention is to assume that both sequences are written in the 5′-to-3′ direction. The "Watson strand" refers to 5′-to-3′ top strand (5′→3′), whereas the "Crick strand" refers to the 5′-to-3′ bottom strand (3′←5′).[4] Both Watson and Crick strands can be either sense or antisense strands depending on the specific gene product made from them.
For example, the notation "YEL021W", an alias of the URA3 gene used in the National Center for Biotechnology Information (NCBI) database, denotes that this gene is in the 21st open reading frame (ORF) from the centromere of the left arm (L) of Yeast (Y) chromosome number V (E), and that the expression coding strand is the Watson strand (W). "YKL074C" denotes the 74th ORF to the left of the centromere of chromosome XI and that the coding strand is the Crick strand (C). Another confusing term referring to "Plus" and "Minus" strand is also widely used. Whether the strand is sense (positive) or antisense (negative), the default query sequence in NCBI BLAST alignment is "Plus" strand.
Ambisense
A single-stranded genome that is used in both positive-sense and negative-sense capacities is said to be ambisense. Some viruses have ambisense genomes.
Antisense RNA
An RNA sequence that is complementary to an
Some
RNA sense in viruses
In virology, the term "sense" has a slightly different meaning. The genome of an RNA virus can be said to be either positive-sense, also known as a "plus-strand", or negative-sense, also known as a "minus-strand". In most cases, the terms "sense" and "strand" are used interchangeably, making terms such as "positive-strand" equivalent to "positive-sense", and "plus-strand" equivalent to "plus-sense". Whether a viral genome is positive-sense or negative-sense can be used as a basis for classifying viruses.
Positive-sense
Positive-sense (
Negative-sense
Negative-sense (3′-to-5′) viral RNA is complementary to the viral mRNA, thus a positive-sense RNA must be produced by an RNA-dependent RNA polymerase from it prior to translation. Like DNA, negative-sense RNA has a nucleotide sequence complementary to the mRNA that it encodes; also like DNA, this RNA cannot be translated into protein directly. Instead, it must first be transcribed into a positive-sense RNA that acts as an mRNA. Some viruses (e.g. influenza viruses) have negative-sense genomes and so must carry an RNA polymerase inside the virion.
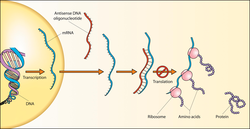
Antisense oligonucleotides
If the antisense oligonucleotide contains a stretch of DNA or a DNA mimic (phosphorothioate DNA, 2′F-ANA, or others) it can recruit
Other antisense mechanisms are not enzyme-dependent, but involve steric blocking of their target RNA (e.g. to prevent translation or to induce alternative splicing). Steric blocking antisense mechanisms often use oligonucleotides that are heavily modified. Since there is no need for RNase H recognition, this can include chemistries such as 2′-O-alkyl, peptide nucleic acid (PNA), locked nucleic acid (LNA), and Morpholino oligomers.
See also
- Antisense therapy
- Directionality (molecular biology)
- DNA codon table
- RNA virus
- Transcription (genetics)
- Translation (genetics)
- Viral replication
References
- PMID 12782362.
- PMID 1993885.
- PMID 2016591.
- PMID 21303550.
- ^ "FDA approves fomivirsen for CMV". healio. 1 October 1998. Retrieved 18 September 2020.
- ^ "FDA approves orphan drug for inherited cholesterol disorder". Drug Topics. 30 January 2013. Retrieved 18 September 2020.
- PMID 75545.
- PMID 22069063.
- ^ PMID 30577479.
- PMID 29322273.
- S2CID 24496875.
- S2CID 45686564.
