Proline
| |||
| |||
| Names | |||
|---|---|---|---|
| IUPAC name
Proline
| |||
| Systematic IUPAC name
Pyrrolidine-2-carboxylic acid[1] | |||
| Identifiers | |||
3D model (
JSmol ) |
|||
| 80812 | |||
| ChEBI |
| ||
| ChEMBL |
| ||
| ChemSpider | |||
| DrugBank |
| ||
ECHA InfoCard
|
100.009.264 | ||
| EC Number |
| ||
| 26927 | |||
| KEGG |
| ||
| MeSH | Proline | ||
PubChem CID
|
|||
RTECS number
|
| ||
| UNII |
| ||
CompTox Dashboard (EPA)
|
|||
| |||
| |||
| Properties | |||
| C5H9NO2 | |||
| Molar mass | 115.132 g·mol−1 | ||
| Appearance | Transparent crystals | ||
| Melting point | 205 to 228 °C (401 to 442 °F; 478 to 501 K) (decomposes) | ||
| Solubility | 1.5g/100g ethanol 19 degC[2] | ||
| log P | -0.06 | ||
| Acidity (pKa) | 1.99 (carboxyl), 10.96 (amino)[3] | ||
| Supplementary data page | |||
| Proline (data page) | |||
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa).
| |||
Proline (symbol Pro or P)
Proline is the only proteinogenic secondary amino acid which is a secondary amine, as the nitrogen atom is attached both to the α-carbon and to a chain of three carbons that together form a five-membered ring.
History and etymology
Proline was first isolated in 1900 by
The name proline comes from pyrrolidine, one of its constituents.[7]
Biosynthesis
Proline is

Biological activity
L-Proline has been found to act as a weak
A diet rich in proline was linked to an increased risk of depression in humans in a study from 2022 that was tested on a limited pre-clinical trial on humans and primarily in other organisms. Results were significant in the other organisms.[16]
Properties in protein structure
The distinctive cyclic structure of proline's side chain gives proline an exceptional conformational rigidity compared to other amino acids. It also affects the rate of peptide bond formation between proline and other amino acids. When proline is bound as an amide in a peptide bond, its nitrogen is not bound to any hydrogen, meaning it cannot act as a hydrogen bond donor, but can be a hydrogen bond acceptor.
Peptide bond formation with incoming Pro-tRNAPro in the ribosome is considerably slower than with any other tRNAs, which is a general feature of N-alkylamino acids.[17] Peptide bond formation is also slow between an incoming tRNA and a chain ending in proline; with the creation of proline-proline bonds slowest of all.[18]
The exceptional conformational rigidity of proline affects the
Proline acts as a structural disruptor in the middle of regular
Multiple prolines and/or
Cis–trans isomerization
From a kinetic standpoint, cis–trans proline isomerization is a very slow process that can impede the progress of protein folding by trapping one or more proline residues crucial for folding in the non-native isomer, especially when the native protein requires the cis isomer. This is because proline residues are exclusively synthesized in the ribosome as the trans isomer form. All organisms possess prolyl isomerase enzymes to catalyze this isomerization, and some bacteria have specialized prolyl isomerases associated with the ribosome. However, not all prolines are essential for folding, and protein folding may proceed at a normal rate despite having non-native conformers of many X–Pro peptide bonds.
Uses
Proline and its derivatives are often used as asymmetric catalysts in
In brewing, proteins rich in proline combine with polyphenols to produce haze (turbidity).[25]
L-Proline is an osmoprotectant and therefore is used in many pharmaceutical and biotechnological applications.
The
Specialties
Proline is one of the two amino acids that do not follow along with the typical Ramachandran plot, along with glycine. Due to the ring formation connected to the beta carbon, the ψ and φ angles about the peptide bond have fewer allowable degrees of rotation. As a result, it is often found in "turns" of proteins as its free entropy (ΔS) is not as comparatively large to other amino acids and thus in a folded form vs. unfolded form, the change in entropy is smaller. Furthermore, proline is rarely found in α and β structures as it would reduce the stability of such structures, because its side chain α-nitrogen can only form one nitrogen bond.
Additionally, proline is the only amino acid that does not form a red-purple colour when developed by spraying with ninhydrin for uses in chromatography. Proline, instead, produces an orange-yellow colour.
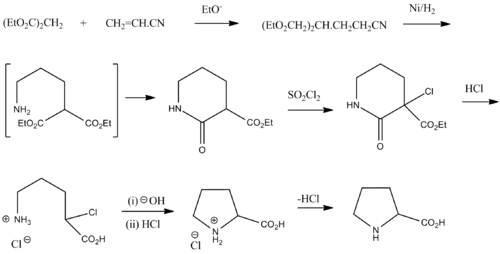
Synthesis
See also
- Hyperprolinemia
- Inborn error of metabolism
- Prolidase deficiency
- Prolinol
References
- ^ "Proline". PubChem. U.S. National Library of Medicine. Archived from the original on 16 January 2014. Retrieved 8 May 2018.
- ISBN 978-3-540-69933-0. Archivedfrom the original on 2016-05-15.
- ^ Nelson DL, Cox MM. Principles of Biochemistry. New York: W.H. Freeman and Company.
- ^ "Nomenclature and Symbolism for Amino Acids and Peptides". IUPAC-IUB Joint Commission on Biochemical Nomenclature. 1983. Archived from the original on 9 October 2008. Retrieved 5 March 2018.
- ^ Plimmer RH (1912) [1908], Plimmer RH, Hopkins FG (eds.), The chemical composition of the proteins, Monographs on biochemistry, vol. Part I. Analysis (2nd ed.), London: Longmans, Green and Co., p. 130, retrieved September 20, 2010
- ^ "Proline". Amino Acids Guide. Archived from the original on 2015-11-27.
- ^ "Proline". American Heritage Dictionary of the English Language, 4th edition. Archived from the original on 2015-09-15. Retrieved 2015-12-06.
- ISBN 1-57259-153-6..
- ^ ISBN 978-0-08-053519-7. Archivedfrom the original on 26 April 2016.
- ^ PMID 1349155.
- ^ ISBN 978-1-4398-4833-3. Archivedfrom the original on 14 May 2016.
- S2CID 21788988.
- S2CID 252438394.
- S2CID 233397785.
- PMID 30131422.
- S2CID 248528026.
- PMID 19104062..
- PMID 23288527.
- S2CID 940786.
- from the original on 2012-01-18.
- PMID 29064600.
- PMID 6589627.
- PMID 16478167.
- PMID 28506462.
- ^ Siebert KJ. "Haze and Foam". Cornell AgriTech. Archived from the original on 2010-07-11. Retrieved 2010-07-13. Accessed July 12, 2010.
- S2CID 84495391.
- ^ Vogel, Practical Organic Chemistry 5th edition
Further reading
- Balbach J, Schmid FX (2000). "Proline isomerization and its catalysis in protein folding". In Pain RH (ed.). Mechanisms of Protein Folding (2nd ed.). Oxford University Press. pp. 212–249. ISBN 978-0-19-963788-1..