Protein structure
This article needs additional citations for verification. (May 2018) |


Protein structure is the
Protein structures range in size from tens to several thousand amino acids.
A protein usually undergoes reversible structural changes in performing its biological function. The alternative structures of the same protein are referred to as different conformations, and transitions between them are called conformational changes.
Levels of protein structure
There are four distinct levels of protein structure.
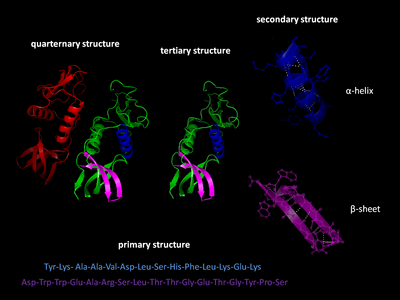
Primary structure
The
Secondary structure

Tertiary structure
Quaternary structure
Quaternary structure is the three-dimensional structure consisting of the aggregation of two or more individual polypeptide chains (subunits) that operate as a single functional unit (
Domains, motifs, and folds in protein structure

Proteins are frequently described as consisting of several structural units. These units include domains,
Structural domain
A
Structural and sequence motifs
The structural and sequence motifs refer to short segments of protein three-dimensional structure or amino acid sequence that were found in a large number of different proteins
Supersecondary structure
Tertiary protein structures can have multiple secondary elements on the same polypeptide chain. The
Protein fold
A protein fold refers to the general protein architecture, like a helix bundle, β-barrel, Rossmann fold or different "folds" provided in the Structural Classification of Proteins database.[9] A related concept is protein topology.
Protein dynamics and conformational ensembles
Proteins are not static objects, but rather populate ensembles of

Proteins are often thought of as relatively stable tertiary structures that experience conformational changes after being affected by interactions with other proteins or as a part of enzymatic activity. However, proteins may have varying degrees of stability, and some of the less stable variants are intrinsically disordered proteins. These proteins exist and function in a relatively 'disordered' state lacking a stable tertiary structure. As a result, they are difficult to describe by a single fixed tertiary structure. Conformational ensembles have been devised as a way to provide a more accurate and 'dynamic' representation of the conformational state of intrinsically disordered proteins.[15][14]
Protein ensemble files are a representation of a protein that can be considered to have a flexible structure. Creating these files requires determining which of the various theoretically possible protein conformations actually exist. One approach is to apply computational algorithms to the protein data in order to try to determine the most likely set of conformations for an ensemble file. There are multiple methods for preparing data for the Protein Ensemble Database that fall into two general methodologies – pool and molecular dynamics (MD) approaches (diagrammed in the figure). The pool based approach uses the protein's amino acid sequence to create a massive pool of random conformations. This pool is then subjected to more computational processing that creates a set of theoretical parameters for each conformation based on the structure. Conformational subsets from this pool whose average theoretical parameters closely match known experimental data for this protein are selected. The alternative molecular dynamics approach takes multiple random conformations at a time and subjects all of them to experimental data. Here the experimental data is serving as limitations to be placed on the conformations (e.g. known distances between atoms). Only conformations that manage to remain within the limits set by the experimental data are accepted. This approach often applies large amounts of experimental data to the conformations which is a very computationally demanding task.[14]
The conformational ensembles were generated for a number of highly dynamic and partially unfolded proteins, such as Sic1/Cdc4,[16] p15 PAF,[17] MKK7,[18] Beta-synuclein[19] and P27[20]
Protein folding
This section needs expansion. You can help by adding to it. (April 2019) |
As it is translated, polypeptides exit the ribosome mostly as a random coil and folds into its native state.[21][22] The final structure of the protein chain is generally assumed to be determined by its amino acid sequence (Anfinsen's dogma).[23]
Protein stability
Thermodynamic stability of proteins represents the free energy difference between the folded and unfolded protein states. This free energy difference is very sensitive to temperature, hence a change in temperature may result in unfolding or denaturation. Protein denaturation may result in loss of function, and loss of native state. The free energy of stabilization of soluble globular proteins typically does not exceed 50 kJ/mol.[citation needed] Taking into consideration the large number of hydrogen bonds that take place for the stabilization of secondary structures, and the stabilization of the inner core through hydrophobic interactions, the free energy of stabilization emerges as small difference between large numbers.[24]
Protein structure determination


Around 90% of the protein structures available in the
General secondary structure composition can be determined via
Protein structure databases
A protein structure database is a database that is modeled around the various experimentally determined protein structures. The aim of most protein structure databases is to organize and annotate the protein structures, providing the biological community access to the experimental data in a useful way. Data included in protein structure databases often includes 3D coordinates as well as experimental information, such as unit cell dimensions and angles for x-ray crystallography determined structures. Though most instances, in this case either proteins or a specific structure determinations of a protein, also contain sequence information and some databases even provide means for performing sequence based queries, the primary attribute of a structure database is structural information, whereas sequence databases focus on sequence information, and contain no structural information for the majority of entries. Protein structure databases are critical for many efforts in computational biology such as structure based drug design, both in developing the computational methods used and in providing a large experimental dataset used by some methods to provide insights about the function of a protein.[32]
Structural classifications of proteins
Protein structures can be grouped based on their structural similarity,
Computational prediction of protein structure
The generation of a
See also
- Biomolecular structure
- Gene structure
- Nucleic acid structure
- PCRPi-DB
- Ribbon diagram 3D schematic representation of proteins
References
- ISBN 978-1-305-68645-8.
- PMID 15951512.
- PMID 14886310.
- PMID 13658959.
- PMID 14816373.
- S2CID 29904865.
- PMID 19059267.
- PMID 25694109.
- S2CID 7147867. Archived from the originalon 5 January 2013.
- PMID 21570668.
- ^
Fraser JS, Clarkson MW, Degnan SC, Erion R, Kern D, Alber T (December 2009). "Hidden alternative structures of proline isomerase essential for catalysis". Nature. 462 (7273): 669–673. PMID 19956261.
- OCLC 690489261.
- PMID 18365235. 1432-119X.
- ^ PMID 26301226.
- ^ Protein Ensemble Database
- PMID 20399186.
- PMID 24559989.
- PMID 25737554.
- PMID 25389903.
- PMID 16214166.
- PMID 21111607.
- ISBN 978-0-8153-3218-3.
- PMID 4565129.
- PMID 1969647.
- S2CID 4162786.
- ^ "PDB Statistics". 1 October 2022.
- )
- PMID 22356513.
- PMID 17981904.
- PMID 23056252.
- PMID 28637405.
- S2CID 45184564.
- PMID 7723011. Archived from the original(PDF) on 26 April 2012.
- PMID 9309224.
- ^ PMID 19325884.
- PMID 20457744.
- PMID 18436442.
Further reading
- 50 Years of Protein Structure Determination Timeline - HTML Version - National Institute of General Medical Sciences Archived 29 October 2018 at the NIH
External links
 Media related to Protein structures at Wikimedia Commons
Media related to Protein structures at Wikimedia Commons
- Protein Structure drugdesign.org
