Proteasome
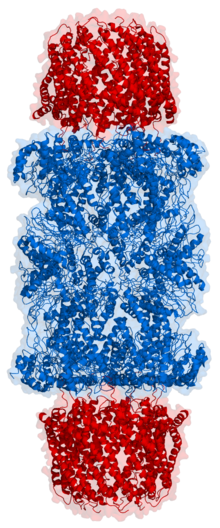
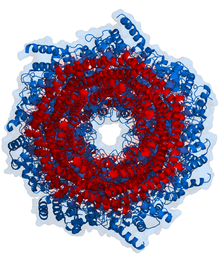


Proteasomes are protein complexes which degrade unneeded or damaged proteins by proteolysis, a chemical reaction that breaks peptide bonds. Enzymes that help such reactions are called proteases.
Proteasomes are part of a major mechanism by which cells regulate the concentration of particular proteins and degrade misfolded proteins. Proteins are tagged for degradation with a small protein called ubiquitin. The tagging reaction is catalyzed by enzymes called ubiquitin ligases. Once a protein is tagged with a single ubiquitin molecule, this is a signal to other ligases to attach additional ubiquitin molecules. The result is a polyubiquitin chain that is bound by the proteasome, allowing it to degrade the tagged protein.[1] The degradation process yields peptides of about seven to eight amino acids long, which can then be further degraded into shorter amino acid sequences and used in synthesizing new proteins.[1]
Proteasomes are found inside all eukaryotes and archaea, and in some bacteria. In eukaryotes, proteasomes are located both in the nucleus and in the cytoplasm.[2]
In structure, the proteasome is a cylindrical complex containing a "core" of four stacked rings forming a central pore. Each ring is composed of seven individual proteins. The inner two rings are made of seven β subunits that contain three to seven protease active sites. These sites are located on the interior surface of the rings, so that the target protein must enter the central pore before it is degraded. The outer two rings each contain seven α subunits whose function is to maintain a "gate" through which proteins enter the barrel. These α subunits are controlled by binding to "cap" structures or regulatory particles that recognize polyubiquitin tags attached to protein substrates and initiate the degradation process. The overall system of ubiquitination and proteasomal degradation is known as the ubiquitin–proteasome system.[3]
The proteasomal degradation pathway is essential for many cellular processes, including the cell cycle, the regulation of gene expression, and responses to oxidative stress. The importance of proteolytic degradation inside cells and the role of ubiquitin in proteolytic pathways was acknowledged in the award of the 2004 Nobel Prize in Chemistry to Aaron Ciechanover, Avram Hershko and Irwin Rose.[4]
Discovery
Before the discovery of the ubiquitin–proteasome system, protein degradation in cells was thought to rely mainly on
Much of the early work leading up to the discovery of the ubiquitin proteasome system occurred in the late 1970s and early 1980s at the
Although
Structure and organization
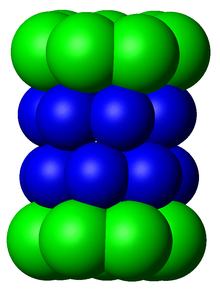
The proteasome subcomponents are often referred to by their
20S core particle
The number and diversity of subunits contained in the 20S core particle depends on the organism; the number of distinct and specialized subunits is larger in multicellular than unicellular organisms and larger in eukaryotes than in prokaryotes. All 20S particles consist of four stacked heptameric ring structures that are themselves composed of two different types of subunits; α subunits are structural in nature, whereas β subunits are predominantly catalytic. The α subunits are pseudoenzymes homologous to β subunits. They are assembled with their N-termini adjacent to that of the β subunits.[17] The outer two rings in the stack consist of seven α subunits each, which serve as docking domains for the regulatory particles and the alpha subunits N-termini (Pfam PF10584) form a gate that blocks unregulated access of substrates to the interior cavity.[18] The inner two rings each consist of seven β subunits and in their N-termini contain the protease active sites that perform the proteolysis reactions.[19] Three distinct catalytic activities were identified in the purified complex: chymotrypsin-like, trypsin-like and peptidylglutamyl-peptide hydrolyzing.[20] The size of the proteasome is relatively conserved and is about 150 angstroms (Å) by 115 Å. The interior chamber is at most 53 Å wide, though the entrance can be as narrow as 13 Å, suggesting that substrate proteins must be at least partially unfolded to enter.[21]
In
19S regulatory particle
The 19S particle in eukaryotes consists of 19 individual proteins and is divisible into two subassemblies, a 9-subunit base that binds directly to the α ring of the 20S core particle, and a 10-subunit lid. Six of the nine base proteins are ATPase subunits from the AAA Family, and an evolutionary homolog of these ATPases exists in archaea, called PAN (proteasome-activating nucleotidase).[26] The association of the 19S and 20S particles requires the binding of ATP to the 19S ATPase subunits, and ATP hydrolysis is required for the assembled complex to degrade folded and ubiquitinated proteins. Note that only the step of substrate unfolding requires energy from ATP hydrolysis, while ATP-binding alone can support all the other steps required for protein degradation (e.g., complex assembly, gate opening, translocation, and proteolysis).[27][28] In fact, ATP binding to the ATPases by itself supports the rapid degradation of unfolded proteins. However, while ATP hydrolysis is required for unfolding only, it is not yet clear whether this energy may be used in the coupling of some of these steps.[28][29]
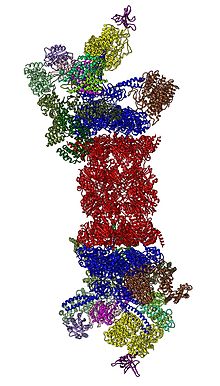
In 2012, two independent efforts have elucidated the molecular architecture of the 26S proteasome by single particle electron microscopy.[31][32] In 2016, three independent efforts have determined the first near-atomic resolution structure of the human 26S proteasome in the absence of substrates by cryo-EM.[33][34][35] In 2018, a major effort has elucidated the detailed mechanisms of deubiquitylation, initiation of translocation and processive unfolding of substrates by determining seven atomic structures of substrate-engaged 26S proteasome simultaneously.[15] In the heart of the 19S, directly adjacent to the 20S, are the AAA-ATPases (AAA proteins) that assemble to a heterohexameric ring of the order Rpt1/Rpt2/Rpt6/Rpt3/Rpt4/Rpt5. This ring is a trimer of dimers: Rpt1/Rpt2, Rpt6/Rpt3, and Rpt4/Rpt5 dimerize via their N-terminal coiled-coils. These coiled-coils protrude from the hexameric ring. The largest regulatory particle non-ATPases Rpn1 and Rpn2 bind to the tips of Rpt1/2 and Rpt6/3, respectively. The ubiquitin receptor Rpn13 binds to Rpn2 and completes the base sub-complex. The lid covers one half of the AAA-ATPase hexamer (Rpt6/Rpt3/Rpt4) and, unexpectedly, directly contacts the 20S via Rpn6 and to lesser extent Rpn5. The subunits Rpn9, Rpn5, Rpn6, Rpn7, Rpn3, and Rpn12, which are structurally related among themselves and to subunits of the COP9 complex and eIF3 (hence called PCI subunits) assemble to a horseshoe-like structure enclosing the Rpn8/Rpn11 heterodimer. Rpn11, the deubiquitinating enzyme, is placed at the mouth of the AAA-ATPase hexamer, ideally positioned to remove ubiquitin moieties immediately before translocation of substrates into the 20S. The second ubiquitin receptor identified to date, Rpn10, is positioned at the periphery of the lid, near subunits Rpn8 and Rpn9.
Conformational changes of 19S
The 19S regulatory particle within the 26S proteasome holoenzyme has been observed in six strongly differing conformational states in the absence of substrates to date.[36][37] A hallmark of the AAA-ATPase configuration in this predominant low-energy state is a staircase- or lockwasher-like arrangement of the AAA-domains.[30][31] In the presence of ATP but absence of substrate three alternative, less abundant conformations of the 19S are adopted primarily differing in the positioning of the lid with respect to the AAA-ATPase module.[33][37] In the presence of ATP-γS or a substrate, considerably more conformations have been observed displaying dramatic structural changes of the AAA-ATPase module.[15][36][38][39] Some of the substrate-bound conformations bear high similarity to the substrate-free ones, but they are not entirely identical, particularly in the AAA-ATPase module.[15][36] Prior to the 26S assembly, the 19S regulatory particle in a free form has also been observed in seven conformational states.[40] Notably, all these conformers are somewhat different and present distinct features. Thus, the 19S regulatory particle can sample at least 20 conformational states under different physiological conditions.
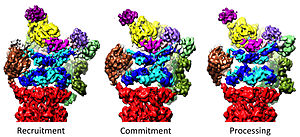
Regulation of the 20S by the 19S
The 19S regulatory particle is responsible for stimulating the 20S to degrade proteins. A primary function of the 19S regulatory ATPases is to open the gate in the 20S that blocks the entry of substrates into the degradation chamber.[41] The mechanism by which the proteasomal ATPase open this gate has been recently elucidated.[18] 20S gate opening, and thus substrate degradation, requires the C-termini of the proteasomal ATPases, which contains a specific motif (i.e., HbYX motif). The ATPases C-termini bind into pockets in the top of the 20S, and tether the ATPase complex to the 20S proteolytic complex, thus joining the substrate unfolding equipment with the 20S degradation machinery. Binding of these C-termini into these 20S pockets by themselves stimulates opening of the gate in the 20S in much the same way that a "key-in-a-lock" opens a door.[18] The precise mechanism by which this "key-in-a-lock" mechanism functions has been structurally elucidated in the context of human 26S proteasome at near-atomic resolution, suggesting that the insertion of five C-termini of ATPase subunits Rpt1/2/3/5/6 into the 20S surface pockets are required to fully open the 20S gate.[36][15][33]
Other regulatory particles
20S proteasomes can also associate with a second type of regulatory particle, the 11S regulatory particle, a heptameric structure that does not contain any ATPases and can promote the degradation of short peptides but not of complete proteins. It is presumed that this is because the complex cannot unfold larger substrates. This structure is also known as PA28, REG, or PA26.[17] The mechanisms by which it binds to the core particle through the C-terminal tails of its subunits and induces α-ring conformational changes to open the 20S gate suggest a similar mechanism for the 19S particle.[42] The expression of the 11S particle is induced by interferon gamma and is responsible, in conjunction with the immunoproteasome β subunits, for the generation of peptides that bind to the major histocompatibility complex.[16]
Yet another type of non-ATPase regulatory particle is the Blm10 (yeast) or PA200/PSME4 (human). It opens only one α subunit in the 20S gate and itself folds into a dome with a very small pore over it.[17]
Assembly
The assembly of the proteasome is a complex process due to the number of subunits that must associate to form an active complex. The β subunits are synthesized with
Only recently, the assembly process of the 19S regulatory particle has been elucidated to considerable extent. The 19S regulatory particle assembles as two distinct subcomponents, the base and the lid. Assembly of the base complex is facilitated by four assembly
Protein degradation process

Ubiquitination and targeting
Proteins are targeted for degradation by the proteasome with covalent modification of a lysine residue that requires the coordinated reactions of three
The mechanism by which a polyubiquitinated protein is targeted to the proteasome is not fully understood. A few high-resolution snapshots of the proteasome bound to a polyubiquitinated protein suggest that ubiquitin receptors might be coordinated with deubiquitinase Rpn11 for initial substrate targeting and engagement.
The
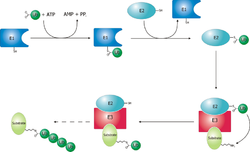
Deubiquitylation
Ubiquitin chains conjugated to a protein targeted for proteasomal degradation are normally removed by any one of the three proteasome-associated deubiquitylating enzymes (DUBs), which are Rpn11, Ubp6/USP14 and UCH37. This process recycles ubiquitin and is essential to maintain the ubiquitin reservoir in cells.[56] Rpn11 is an intrinsic, stoichiometric subunit of the 19S regulatory particle and is essential for the function of 26S proteasome. The DUB activity of Rpn11 is enhanced in the proteasome as compared to its monomeric form. How Rpn11 removes a ubiquitin chain en bloc from a protein substrate was captured by an atomic structure of the substrate-engaged human proteasome in a conformation named EB.[15] Interestingly, this structure also shows how the DUB activity is coupled to the substrate recognition by the proteasomal AAA-ATPase. In contrast to Rpn11, USP14 and UCH37 are the DUBs that do not always associated with the proteasome. In cells, about 10-40% of the proteasomes were found to have USP14 associated. Both Ubp6/USP14 and UCH37 are largely activated by the proteasome and exhibit a very low DUB activity alone. Once activated, USP14 was found to suppress proteasome function by its DUB activity and by inducing parallel pathways of proteasome conformational transitions, one of which turned out to directly prohibit substrate insertion into the AAA-ATPase, as intuitively observed by time-resolved cryogenic electron microscopy. [57] It appears that USP14 regulates proteasome function at multiple checkpoints by both catalytically competing with Rpn11 and allosterically reprogramming the AAA-ATPase states, which is rather unexpected for a DUB.[57] These observations imply that the proteasome regulation may depend on its dynamic transitions of conformational states.
Unfolding and translocation
After a protein has been ubiquitinated, it is recognized by the 19S regulatory particle in an ATP-dependent binding step.
The gate formed by the α subunits prevents peptides longer than about four residues from entering the interior of the 20S particle. The ATP molecules bound before the initial recognition step are hydrolyzed before translocation. While energy is needed for substrate unfolding, it is not required for translocation.[27][28] The assembled 26S proteasome can degrade unfolded proteins in the presence of a non-hydrolyzable ATP analog, but cannot degrade folded proteins, indicating that energy from ATP hydrolysis is used for substrate unfolding.[27] Passage of the unfolded substrate through the opened gate occurs via facilitated diffusion if the 19S cap is in the ATP-bound state.[62]
The mechanism for unfolding of
Proteolysis
The proteasome functions as an
Although the proteasome normally produces very short peptide fragments, in some cases these products are themselves biologically active and functional molecules. Certain
Ubiquitin-independent degradation
Although most proteasomal substrates must be ubiquitinated before being degraded, there are some exceptions to this general rule, especially when the proteasome plays a normal role in the post-
Evolution
The 20S proteasome is both ubiquitous and essential in eukaryotes and archaea. The
Sequence analysis suggests that the catalytic β subunits diverged earlier in evolution than the predominantly structural α subunits. In bacteria that express a 20S proteasome, the β subunits have high
Cell cycle control
Cell cycle progression is controlled by ordered action of
Earlier cell cycle checkpoints such as post-
Individual components of the 19S particle have their own regulatory roles.
Like eukaryotes, some archaea also use the proteasome to control cell cycle, specifically by controlling ESCRT-III-mediated cell division.[83]
Regulation of plant growth
In
Apoptosis
Both internal and external
Proteasome inhibition has different effects on apoptosis induction in different cell types. In general, the proteasome is not required for apoptosis, although inhibiting it is pro-apoptotic in most cell types that have been studied. Apoptosis is mediated through disrupting the regulated degradation of pro-growth cell cycle proteins.
Response to cellular stress
In response to cellular stresses – such as
Similar mechanisms exist to promote the degradation of
Dysregulation of the ubiquitin proteasome system may contribute to several neural diseases. It may lead to brain tumors such as
Role in the immune system
The proteasome plays a straightforward but critical role in the function of the
The strength of MHC class I ligand binding is dependent on the composition of the ligand C-terminus, as peptides bind by hydrogen bonding and by close contacts with a region called the "B pocket" on the MHC surface. Many MHC class I alleles prefer hydrophobic C-terminal residues, and the immunoproteasome complex is more likely to generate hydrophobic C-termini.[101]
Due to its role in generating the activated form of
The proteasome is also involved in
Proteasome inhibitors
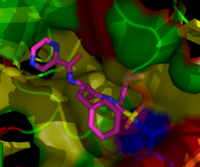
The molecule ritonavir, marketed as Norvir, was developed as a protease inhibitor and used to target HIV infection. However, it has been shown to inhibit proteasomes as well as free proteases; to be specific, the chymotrypsin-like activity of the proteasome is inhibited by ritonavir, while the trypsin-like activity is somewhat enhanced.[113] Studies in animal models suggest that ritonavir may have inhibitory effects on the growth of glioma cells.[114]
Proteasome inhibitors have also shown promise in treating autoimmune diseases in animal models. For example, studies in mice bearing human
Labeling and inhibition of the proteasome is also of interest in laboratory settings for both in vitro and in vivo study of proteasomal activity in cells. The most commonly used laboratory inhibitors are
Clinical significance
The proteasome and its subunits are of clinical significance for at least two reasons: (1) a compromised complex assembly or a dysfunctional proteasome can be associated with the underlying pathophysiology of specific diseases, and (2) they can be exploited as drug targets for therapeutic interventions. More recently, more effort has been made to consider the proteasome for the development of novel diagnostic markers and strategies. An improved and comprehensive understanding of the pathophysiology of the proteasome should lead to clinical applications in the future.
The proteasomes form a pivotal component for the ubiquitin–proteasome system (UPS)
Research has implicated UPS defects in the pathogenesis of neurodegenerative and myodegenerative disorders, including
See also
- The Proteolysis Map
- DSS1/SEM1 protein family
- Exosome complex
- Endoplasmic-reticulum-associated protein degradation
- JUNQ and IPOD
References
- ^ ISBN 978-0-7167-4366-8.
- PMID 8125997.
- S2CID 29860298.
- ^ a b Nobel Prize Committee (2004). "Nobel Prize Awardees in Chemistry, 2004". Retrieved 11 December 2006.
- PMID 264694.
- PMID 666810.
- PMID 265581.
- PMID 16094393.
- S2CID 9028201.
- PMID 6304111.
- PMID 3298229.
- PMID 16094391.
- PMID 3524688.
- PMID 7725097.
- ^ PMID 30479383.
- ^ PMID 16978533.
- ^ PMID 21211719.
- ^ PMID 17803938.
- ^ "MEROPS Family T1". EMBL-EBI. Retrieved 16 February 2019.
- S2CID 23508675.
- ^ S2CID 21603835.
- ^ PMID 9312134.
- ^ PMID 26997268.
- PMID 14739934.
- S2CID 21181637.
- PMID 10473546.
- ^ PMID 16337593.
- ^ PMID 17018291.
- S2CID 4421764.
- ^ PMID 22927375.
- ^ PMID 22237024.
- PMID 22307589.
- ^ PMID 27791164.
- S2CID 21909333.
- PMID 27791164.
- ^ PMID 29636472.
- ^ PMID 24706844.
- PMID 23589842.
- PMID 23770819.
- PMID 28689658.
- PMID 11430818.
- PMID 15916965.
- PMID 16843899.
- PMID 11295488.
- S2CID 21263837.
- PMID 21658604.
- PMID 20471955.
- PMID 6277905.
- PMID 10619848.
- PMID 12795696.
- S2CID 21069699.
- PMID 22897362.
- S2CID 7929162.
- PMID 15556404.
- PMID 19345192.
- ^ PMID 11084366.
- ^ PMID 35477760.
- PMID 15950624.
- S2CID 41599619.
- PMID 21278740.
- PMID 25220455.
- PMID 16919475.
- PMID 16601692.
- PMID 14688254.
- PMID 7725107.
- PMID 8811196.
- S2CID 4261663.
- PMID 9748229.
- PMID 10872471.
- ^ S2CID 7126477.
- PMID 15571816.
- PMID 16927316.
- PMID 12660156.
- PMID 16082197.
- ^ PMID 12401807.
- ^ PMID 12595256.
- PMID 10410804.
- PMID 16921258.
- PMID 16782009.
- PMID 16738333.
- S2CID 4401971.
- PMID 16177571.
- PMID 32764038.
- S2CID 7669386.
- PMID 15889151.
- PMID 7721865.
- S2CID 33829749.
- S2CID 10873052.
- S2CID 29256092.
- ^ PMID 10363983.
- PMID 10381632.
- PMID 17106261.
- PMID 17065559.
- PMID 16169850.
- S2CID 30385354.
- PMID 11295490.
- ^ PMID 19597829.
- PMID 16717251.
- S2CID 27762513.
- S2CID 37185716.
- PMID 11350924.
- PMID 21045130.
- S2CID 37779687.
- ^ United States Food and Drug Administration press release Archived 19 February 2007 at the Wayback Machine 13 May 2003. Access date 29 December 2006. See also FDA Velcade information page.
- PMID 17001068.
- PMID 17095627.
- S2CID 21223980.
- S2CID 38429730.
- PMID 12141956.
- PMID 15613699.
- PMID 22653976.
- S2CID 28749125.
- PMID 10585454.
- S2CID 24423806.
- PMID 11877475.
- PMID 10452747.
- PMID 17114003.
- PMID 24457024.
- PMID 9383453.
- S2CID 14103185.
- PMID 25324717.
- PMID 24380730.
- PMID 25133688.
- PMID 25651176.
- ^ PMID 10723801.
- PMID 25560147.
- PMID 10899438.
- ^ S2CID 2211658.
- ^ S2CID 22396490.
- S2CID 28190967.
- S2CID 5780576.
- PMID 12792671.
- PMID 23220331.
- PMID 20159828.
- S2CID 7073263.
- PMID 12654543.
- S2CID 26973319.
- PMID 12375310.
Further reading
- Glickman MH, Adir N (January 2004). "The proteasome and the delicate balance between destruction and rescue". PLOS Biology. 2 (1): e13. PMID 14737189.
- The Yeast 26S Proteasome with list of subunits and pictures
- Ciechanover A (September 2005). "Early work on the ubiquitin proteasome system, an interview with Aaron Ciechanover. Interview by CDD". Cell Death and Differentiation. 12 (9): 1167–77. PMID 16094393.
- Hershko A (September 2005). "Early work on the ubiquitin proteasome system, an interview with Avram Hershko. Interview by CDD". Cell Death and Differentiation. 12 (9): 1158–61. PMID 16094391.
- Rose I (September 2005). "Early work on the ubiquitin proteasome system, an interview with Irwin Rose. Interview by CDD". Cell Death and Differentiation. 12 (9): 1162–6. PMID 16094392.
- Cvek B, Dvorak Z (2007). "Targeting of nuclear factor-kappaB and proteasome by dithiocarbamate complexes with metals". Current Pharmaceutical Design. 13 (30): 3155–67. PMID 17979756. Archived from the originalon 29 July 2012.
External links
- Proteasome subunit nomenclature guide
- 3D proteasome structures in the EM Data Bank(EMDB)
- Key points of proteasome function
