RELA
Ensembl | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| UniProt | |||||||||
| RefSeq (mRNA) | |||||||||
| RefSeq (protein) | |||||||||
| Location (UCSC) | Chr 11: 65.65 – 65.66 Mb | Chr 19: 5.69 – 5.7 Mb | |||||||
| PubMed search | [3] | [4] | |||||||
| View/Edit Human | View/Edit Mouse |
Transcription factor p65 also known as nuclear factor NF-kappa-B p65 subunit is a protein that in humans is encoded by the RELA gene.[5]
RELA, also known as p65, is a REL-associated protein involved in NF-κB heterodimer formation, nuclear translocation and activation [citation needed]. NF-κB is an essential transcription factor complex involved in all types of cellular processes, including cellular metabolism, chemotaxis, etc. Phosphorylation and acetylation of RELA are crucial post-translational modifications required for NF-κB activation. RELA has also been shown to modulate immune responses, and activation of RELA is positively associated with multiple types of cancer.
Gene and expression
RELA, or v-rel avian reticuloendotheliosis viral oncogene homolog A, is also known as p65 or NFKB3.[6] It is located on chromosome 11 q13, and its nucleotide sequence is 1473 nucleotide long.[7] RELA protein has four isoforms, the longest and the predominant one being 551 amino acids. RELA is expressed alongside p50 in various cell types, including epithelial/endothelial cells and neuronal tissues.[8]
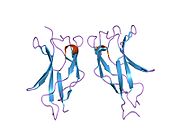
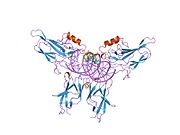
Structure
RELA is one member of the NF-κB family, one of the essential transcription factors under intensive study. Seven proteins encoded by five genes are involved in the NF-κB complex, namely p105, p100, p50, p52, RELA, c-REL and RELB.[9] Like other proteins in this complex, RELA contains a N-terminal REL-homology domain (RHD), and also a C-terminal transactivation domain (TAD). RHD is involved in DNA binding, dimerization and NF-κB/REL inhibitor interaction. On the other hand, TAD is responsible for interacting with the basal transcription complex including many coactivators of transcription such as TBP, TFIIB and CREB-CBP.[9] RELA and p50 is the mostly commonly found heterodimer complex among NF-κB homodimers and heterodimers, and is the functional component participating in nuclear translocation and activation of NF-κB.
RELA is a 65 kDa protein.[10]
Phosphorylation
Phosphorylation of RELA plays a key role in regulating NF-κB activation and function. Subsequent to NF-κB nuclear translocation, RELA undergoes site-specific post-translational modifications to further enhance the NF-κB function as a transcription factor. RELA can either be phosphorylated in the RHD region or the TAD region, attracting different interaction partners. Triggered by lipopolysaccharide (LPS), protein kinase A (PKA) specifically phosphorylates serine 276 in the RHD domain in the cytoplasm, controlling NF-κB DNA-binding and oligomerization. Two residues in the TAD region are targeted by phosphorylation. After IL-1or TNFα stimulation, serine 529 is phosphorylated by casein kinase II (
Acetylation
In vivo studies revealed that RELA is also under acetylation modification in the nucleus, which is just as important as phosphorylation as a post-translational modification of proteins. Lysines 218, 221 and 310 are acetylation targets within RELA, and response to acetylation is site-specific.[9] For instance, lysine 221 acetylation facilitates RELA dissociation from IκBα and enhances its DNA-binding affinity. Lysine 310 acetylation is indispensable for the full transcriptional activity of RELA, but does not affect its DNA-binding ability. Hypothesis about RELA acetylation suggests acetylation aids its subsequent recognition by transcriptional co-activators with bromodomains, which are specialized in recognizing acetylated lysine residues.[9] Lysine 122 and 123 acetylation are found to be negatively correlated with RELA transcriptional activation. Unknown mechanisms mediate the acetylation of RELA possibly using p300/CBP and p300/CBP factor associated coactivators under TNFα or phorbol myristate acetate (PMF) stimulation both in vivo and in vitro.[9] RELA is also under the control of deacetylation via HDAC, and HDAC3 is the mediator of this process both in vivo and in vitro.[8][9]
Methylation
Methylation of lysine 218 and 221 together or lysine 37 alone in the RHD domain of RELA can lead to increased response to cytokines such as IL-1 in mammalian cell culture.[19]
Interactions
As the prototypical heterodimer complex member of the NF-κB, together with p50, RELA/p65 interacts with various proteins in both the cytoplasm and in the nucleus during the process of classical NF-κB activation and nuclear translocation. In the inactive state, RELA/p50 complex is mainly sequestered by IκBα in the cytosol. TNFα, LPS and other factors serve as activation inducers, followed by phosphorylation at residue 32 and 36 of IκBα, leading to rapid degradation of IκBα via the ubiquitin-proteasomal system and subsequent release of RELA/p50 complex.[9] RELA nuclear localization signal used to be sequestered by IκBα is now exposed, and rapid translocation of the NF-κB occurs. In parallel, there is a non-classical NF-κB activation pathway involving the proteolytic cleavage of p100 into p52 instead of p50. This process does not require RELA, hence will not be discussed in detail here.[9] After NF-κB nuclear localization due to TNFα stimulation, p50/RELA heterodimer will function as a transcription factor and bind to a variety of genes involved in all kinds of biological processes, such as leukocyte activation/chemotaxis, negative regulation of TNFIKK pathway, cellular metabolism, antigen processing, just to name a few .[20] Phosphorylation of RELA at different residues also enables its interaction with CDKs and P-TEFb. Phosphorylation at serine 276 in RELA allows its interaction with P-TEFb containing CDK9 and cyclin T1 subunits, and phospho-ser276 RELA-P-TEFb complex is necessary for IL-8 and Gro-β activation.[20] Another mechanism is involved in the activation of genes preloaded with Pol II in a RELA serine 276 phosphorylation independent manner.
RELA has been shown to
- APBA2,[21]
- AHR,[22][23]
- ASCC3,[24]
- BRCA1,[25]
- BTRC,[26]
- c-Fos,[27]
- c-Jun,[27]
- C22orf25,[28]
- CDK9,[29]
- CEBPB,[30][31]
- CEBPE,[32]
- CSNK2A1,[39]
- CSNK2A2,[39]
- DHX9,[40]
- EP300,[37][41]
- ETHE1,[42]
- FUS,[43]
- GCN5,[44]
- HDAC1,[34][41][45]
- HDAC2,[41][46]
- HDAC3,[47]
- ING4,[48]
- IκBα,[26][41][47][49][50][51][52]
- KLF5,[53]
- MDM2,[54]
- MEN1,[55]
- MSK1,[12]
- MTPN,[56]
- NCF1,[57]
- NFKB1,[58][59]
- NFKB2,[58][60]
- NFKBIB,[61][62]
- NFKBIE,[63]
- NR3C1,[64][65][66]
- NCOR2,[67][68]
- PARP1,[69]
- PDLIM2,[70]
- PIAS3,[33]
- PIM1,[18]
- PIN1,[16]
- PKA,[71]
- POU2F1,[72]
- PPARG,[73]
- PPP1R13L,[74][75]
- PRKCZ,[76]
- REL,[50][58][77]
- RFC1,[78]
- RNF25,[79]
- SIRT1,[80]
- SOCS1,[16][81][82]
- STAT3,[85][86]
- TAF4B,[87]
- TBP,[88][89]
- TP53,[86]and
- TRIB3.[90]
Role in immune system
Gene knockout of NF-κB genes via homologous recombination in mice showed the role of these components in innate and adaptive immune responses. RELA knockout mice is embryonic lethal due to liver apoptosis.[8] Lymphocyte activation failure is also observed, suggesting that RELA is indispensable in the proper development of the immune system. In comparison, deletion of other REL-related genes will not cause embryonic development failure, though different levels of defects are also noted.[8] The fact that cytokines such as TNFα and IL-1 can stimulate the activation of RELA also supports its participation in immune response. In general, RELA participates in adaptive immunity and responses to invading pathogens via NF-κB activation. Mice without individual NF-κB proteins are deficient in B- and T-cell activation and proliferation, cytokine production and isotype switching.[8] Mutations in RELA is found responsible for inflammatory bowel disease as well.[8]
Cancer
NF-κB/RELA activation has been found to be correlated with cancer development,[91] suggesting the potential of RELA as a cancer biomarker.[92] Specific modification patterns of RELA have also been observed in many cancer types.[93][94]
Prostate
RELA may have a potential role as biomarker for prostate cancer progression and metastases, as suggested by the association found between RELA nuclear localization and prostate cancer aggressiveness and biochemical recurrence.[95]
Thyroid
Strong correlation between nuclear localization of RELA and clinicopathological parameters for papillary thyroid carcinoma (PTC), suggesting the role of NF-κB activation in tumor growth and aggressiveness in PTC.[96] Other than usage as an biomarker, serine 536 phosphorylation in RELA is also correlated with nuclear translocation and the expression of some transactivating genes such as
Leukemia
Mutations in the transactivation domain of RELA can lead to decrease in transactivating ability and this mutation can be found in lymphoid neoplasia.[98]
Head and neck
Nuclear localization of NF-κB/RELA is positively correlated with tumor micrometastases into lymph and blood and negatively correlated with patient survival outcome in patients with head and neck squamous cell carcinoma (HNSCC).[99] This suggests a role of NF-κB/RELA as a possible target for targeted-therapy.
Breast
There is both a physical and a functional association between RELA and aryl hydrocarbon receptor (AhR), and the subsequent activation of c-myc gene transcription in breast cancer cells.[22] Another paper reported interactions between estrogen receptor (ER) and NF-κB members, including p50 and RELA. It is shown that ERα interacts with both p50 and RELA in vitro and in vivo, and RELA antibody can reduce ERα:ERE complex formation. The paper claims a mutual repression between ER and NF-κB.[100]
Monogenic Behçet’s Disease-like conditions
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000173039 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000024927 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- S2CID 54363279.
- ^ "Homo sapiens p65 gene for p65 subunit of transcription factor NF-kappa". Nucleotide. National Center for Biotechnology Information (NCBI), U.S. National Library of Medicine. 2006-11-14.
- ^ S2CID 6962119.
- ^ S2CID 37637033.
- PMID 10224130.
- PMID 9660950.
- ^ PMID 12628924.
- PMID 12881425.
- PMID 10938077.
- PMID 15073170.
- ^ PMID 14690596.
- PMID 19270718.
- ^ PMID 19911008.
- PMID 23904479.
- ^ PMID 18362169.
- PMID 10777610.
- ^ S2CID 26074149.
- S2CID 2376236.
- PMID 12077347.
- PMID 12700228.
- ^ PMID 9990853.
- ^ PMID 10488148.
- ^ "Molecular Interaction Database". Archived from the original on 2006-05-06.
- S2CID 7611946.
- PMID 12707271.
- PMID 9570146.
- PMID 17255362.
- ^ PMID 15140884.
- ^ PMID 11931769.
- S2CID 6315763.
- PMID 9482849.
- ^ PMID 9096323.
- PMID 22261743.
- ^ PMID 10938077.
- PMID 15355351.
- ^ PMID 12419806.
- PMID 12398897.
- PMID 11278855.
- PMID 19339690.
- PMID 11564889.
- PMID 12138131.
- ^ S2CID 45796404.
- S2CID 4427531.
- PMID 11313474.
- ^ PMID 8139561.
- PMID 9738011.
- S2CID 4327300.
- PMID 25197166.
- PMID 23839035.
- S2CID 44195141.
- PMID 11971907.
- PMID 12618429.
- ^ S2CID 11683986.
- PMID 8798655.
- PMID 8413211.
- PMID 12672800.
- PMID 8816457.
- PMID 9315679.
- PMID 8290595.
- PMID 10995388.
- S2CID 28680611.
- PMID 12589049.
- PMID 10777532.
- PMID 11590148.
- S2CID 13357628.
- PMID 20562110.
- PMID 12019209.
- PMID 23250430.
- PMID 10336463.
- PMID 12134007.
- PMID 11684013.
- PMID 11967310.
- PMID 12509469.
- PMID 12748188.
- PMID 15152190.
- S2CID 25195631.
- PMID 17183367.
- PMID 12055073.
- PMID 7933095.
- PMID 12057007.
- ^ PMID 20546595.
- PMID 9724652.
- PMID 9584164.
- PMID 7706261.
- PMID 12736262.
- PMID 30240635.
In cells from lung tumors containing mutated KRAS, nuclei stained positive for RelA. RelA is implicated in tumor-stroma interactions and correlates strongly with the severity of tumor infiltration by inflammatory cells in NSCLC patients. (pg. 6) — This links RelA activation, and promoter methylation to cancer. (pg. 14) — Thus PARP/RelA and OGG1/RelA/BRD4 are two complementary targets for cancer, as they induce alternative mechanisms of RelA-mediated inflammation. (pg. 17)
- S2CID 38404099.
- PMID 30723172.
- PMID 28681591.
- PMID 23541563.
- PMID 23528368.
- PMID 22558476.
- PMID 9047386.
- PMID 23664323.
- S2CID 24756428.
- S2CID 221888042.
Further reading
- Baldwin AS (1996). "The NF-kappa B and I kappa B proteins: new discoveries and insights". Annual Review of Immunology. 14: 649–83. PMID 8717528.
- Bottex-Gauthier C, Pollet S, Favier A, Vidal DR (Apr 2002). "[The Rel/NF-kappa-B transcription factors: complex role in cell regulation]". Pathologie-Biologie. 50 (3): 204–11. PMID 11980335.
- Garg A, Aggarwal BB (Jun 2002). "Nuclear transcription factor-kappaB as a target for cancer drug development". Leukemia. 16 (6): 1053–68. S2CID 9560168.
- Clarke R, Liu MC, Bouker KB, Gu Z, Lee RY, Zhu Y, Skaar TC, Gomez B, O'Brien K, Wang Y, Hilakivi-Clarke LA (Oct 2003). "Antiestrogen resistance in breast cancer and the role of estrogen receptor signaling". Oncogene. 22 (47): 7316–39. PMID 14576841.
- Bhatt D, Ghosh S (Feb 2014). "Regulation of the NF-κB-Mediated Transcription of Inflammatory Genes". Frontiers in Immunology. 5 (71): 71. PMID 24611065.
External links
- RELA+protein,+human at the U.S. National Library of Medicine Medical Subject Headings (MeSH)