RNA-binding protein
RNA-binding proteins (often abbreviated as RBPs) are
Structure
Many RBPs have modular structures and are composed of multiple repeats of just a few specific basic domains that often have limited sequences. Different RBPs contain these sequences arranged in varying combinations. A specific protein's recognition of a specific RNA has evolved through the rearrangement of these few basic domains. Each basic domain recognizes RNA, but many of these proteins require multiple copies of one of the many common domains to function.[2]
Diversity
As nuclear
Function
RNA processing and modification
Alternative splicing
RNA editing
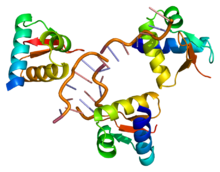
The most extensively studied form of RNA editing involves the ADAR protein. This protein functions through post-transcriptional modification of mRNA transcripts by changing the nucleotide content of the RNA. This is done through the conversion of adenosine to inosine in an enzymatic reaction catalyzed by ADAR. This process effectively changes the RNA sequence from that encoded by the genome and extends the diversity of the gene products. The majority of RNA editing occurs on non-coding regions of RNA; however, some protein-encoding RNA transcripts have been shown to be subject to editing resulting in a difference in their protein's amino acid sequence. An example of this is the glutamate receptor mRNA where glutamine is converted to arginine leading to a change in the functionality of the protein.[5]
Polyadenylation
Export
After processing is complete, mRNA needs to be transported from the
mRNA localization
mRNA localization is critical for regulation of gene expression by allowing spatially regulated protein production. Through mRNA localization proteins are translated in their intended target site of the cell. This is especially important during early development when rapid cell cleavages give different cells various combinations of mRNA which can then lead to drastically different cell fates. RBPs are critical in the localization of this mRNA that insures proteins are only translated in their intended regions. One of these proteins is ZBP1. ZBP1 binds to beta-actin mRNA at the site of transcription and moves with mRNA into the cytoplasm. It then localizes this mRNA to the lamella region of several asymmetric cell types where it can then be translated.[5] In 2008 it was proposed that FMRP was involved in the stimulus-induced localization of several dendritic mRNAs in the neuronal dendrites of cultured hippocampal neurons.[9] More recent studies of FMRP-bound RNAs present in microdissected dendrites of CA1 hippocampal neurons revealed no changes in localization in wild type versus FMRP-null mouse brains.[10]
Translation
Translational regulation provides a rapid mechanism to control gene expression. Rather than controlling gene expression at the transcriptional level, mRNA is already transcribed but the recruitment of ribosomes is controlled. This allows rapid generation of proteins when a signal activates translation. ZBP1 in addition to its role in the localization of B-actin mRNA is also involved in the translational repression of beta-actin mRNA by blocking translation initiation. ZBP1 must be removed from the mRNA to allow the ribosome to properly bind and translation to begin.[5]
Protein–RNA interactions

RNA-binding proteins exhibit highly specific recognition of their RNA targets by recognizing their sequences, structures, motifs and RNA modifications.[11] Specific binding of the RNA-binding proteins allow them to distinguish their targets and regulate a variety of cellular functions via control of the generation, maturation, and lifespan of the RNA transcript. This interaction begins during transcription as some RBPs remain bound to RNA until degradation whereas others only transiently bind to RNA to regulate RNA splicing, processing, transport, and localization.[12] Cross-linking immunoprecipitation (CLIP) methods are used to stringently identify direct RNA binding sites of RNA-binding proteins in a variety of tissues and organisms. In this section, three classes of the most widely studied RNA-binding domains (RNA-recognition motif, double-stranded RNA-binding motif, zinc-finger motif) will be discussed.
RNA-recognition motif (RRM)
The
Double-stranded RNA-binding motif
| Double-stranded RNA-binding motif | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| |||||||||
| Use the Pfam clan for the homologous superfamily. | |||||||||
The double-stranded RNA-binding motif (dsRM, dsRBD), a 70–75 amino-acid domain, plays a critical role in
Zinc fingers

CCHH-type
Role in embryonic development

RNA-binding proteins' transcriptional and
Germline development
In
Somatic development
In addition to RBPs' functions in germline development, post-transcriptional control also plays a significant role in somatic development. Differing from RBPs that are involved in germline and early embryo development, RBPs functioning in somatic development regulate tissue-specific alternative splicing of the mRNA targets. For instance, MEC-8 and UNC-75 containing RRM domains localize to regions of hypodermis and nervous system, respectively.[14] Furthermore, another RRM-containing RBP, EXC-7, is revealed to localize in embryonic excretory canal cells and throughout the nervous system during somatic development.
Neuronal development
ZBP1 was shown to regulate dendritogenesis (dendrite formation) in hippocampal neurons.[16] Other RNA-binding proteins involved in dendrite formation are Pumilio and Nanos,[17] FMRP, CPEB and Staufen 1[18]
Role in cancer
RBPs are emerging to play a crucial role in tumor development.[19] Hundreds of RBPs are markedly dysregulated across human cancers and showed predominant downregulation in tumors related to normal tissues.[19] Many RBPs are differentially expressed in different cancer types for example KHDRBS1(Sam68),[20][21][22] ELAVL1(HuR),[23][24] FXR1[25] and UHMK1.[26] For some RBPs, the change in expression are related with Copy Number Variations (CNV), for example CNV gains of BYSL in colorectal cancer cells[19] and ESRP1, CELF3 in breast cancer, RBM24 in liver cancer, IGF2BP2, IGF2BP3 in lung cancer or CNV losses of KHDRBS2 in lung cancer.[27] Some expression changes are cause due to protein affecting mutations on these RBPs for example NSUN6, ZC3H13, ELAC1, RBMS3, and ZGPAT, SF3B1, SRSF2, RBM10, U2AF1, SF3B1, PPRC1, RBMXL1, HNRNPCL1 etc.[19][27][28][29][30] Several studies have related this change in expression of RBPs to aberrant alternative splicing in cancer.[27][31][32]
Current research

As RNA-binding proteins exert significant control over numerous cellular functions, they have been a popular area of investigation for many researchers. Due to its importance in the biological field, numerous discoveries regarding RNA-binding proteins' potentials have been recently unveiled.[12] Recent development in experimental identification of RNA-binding proteins has extended the number of RNA-binding proteins significantly[33][34][35]
RNA-binding protein Sam68 controls the spatial and temporal compartmentalization of RNA

Neuron-specific CELF family RNA-binding protein UNC-75 specifically binds to the UUGUUGUGUUGU mRNA stretch via its three RNA recognition motifs for the exon 7a selection in C. elegans' neuronal cells. As exon 7a is skipped due to its weak splice sites in non-neuronal cells, UNC-75 was found to specifically activate splicing between exon 7a and exon 8 only in the neuronal cells.[37]
The cold inducible RNA binding protein
Serine-arginine family of RNA-binding protein Slr1 was found exert control on the polarized growth in
See also
- DNA-binding protein
- RNA-binding protein database
- Ribonucleoprotein
External links
- starBase platform: a platform for decoding binding sites of RNA binding proteins (RBPs) from large-scale CLIP-Seq(HITS-CLIP, PAR-CLIP, iCLIP, CLASH) datasets.
- RBPDB database: a database of RNA binding proteins.
- oRNAment: a database of putative RBP binding site instances in both coding and non-coding RNA in various species.
- ATtRACt database: a database of RNA binding proteins and associated motifs.
- SplicedAid-F: a database of hand -cureted human RNA binding proteins database.
- RsiteDB: RNA binding site database
- SPOT-Seq-RNA: Template-based prediction of RNA binding proteins and their complex structures.
- SPOT-Struct-RNA: RNA binding proteins prediction from 3D structures.
- ENCODE Project: A collection of genomic datasets (i.e. RNA Bind-n-seq, eCLIP, RBP targeted shRNA RNA-seq) for RBPs
- RBP Image Database: Images showing the cellular localization of RBPs in cells
- RBPSpot Software: A Deep-Learning based highly accurate software to detect RBP-RNA interaction. It also provides a module to build new RBP-RNA interaction models.
References
- ^ RNA-Binding+Proteins at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- ^ PMID 17473849.
- S2CID 219284021.
- PMID 18959479.
- ^ PMID 18342629.
- S2CID 219284021.
- ^ S2CID 30268055.
- PMID 25112293.
- PMID 18539120.
- PMID 34939924.
- S2CID 219284021.
- ^ PMID 15643449.
- ISBN 978-0-521-86598-2. Retrieved 12 May 2013.
- ^ )
- PMID 2470643.
- PMID 21471362.
- PMID 14972682.
- PMID 18922781.
- ^ PMID 29298429.
- PMID 21565971.
- PMID 23937454.
- PMID 26273626.
- PMID 21935886.
- PMID 23665903.
- PMID 25733852.
- PMID 31975428.
- ^ PMID 27197215.
- S2CID 4429386.
- PMID 22980975.
- PMID 22722193.
- PMID 21041405.
- PMID 25985083.
- PMID 27040163.
- PMID 22658674.
- PMID 22681889.
- PMID 23382180.
- PMID 23468662.
- PMID 23437386.
- PMID 23381995.
