RNA polymerase II
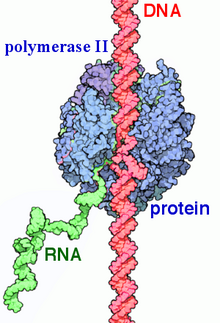
RNA polymerase II (RNAP II and Pol II) is a
Discovery
Early studies suggested a minimum of two RNAPs: one which synthesized rRNA in the
RNA polymerase II (RNAP2) undergoes regulated transcriptional pausing during early elongation. Various studies has shown that disruption of transcription elongation is implicated in
Subunits
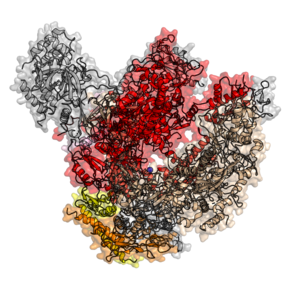
The eukaryotic core RNA polymerase II was first purified using transcription assays.[10] The purified enzyme has typically 10–12 subunits (12 in humans and yeast) and is incapable of specific promoter recognition.[11] Many subunit-subunit interactions are known.[12]
- DNA-directed RNA polymerase II subunit RPB1 – an
- RPB2 (POLR2B) – the second-largest subunit that in combination with at least two other polymerase subunits forms a structure within the polymerase that maintains contact in the active site of the enzyme between the DNA template and the newly synthesized RNA.[15]
- RPB3 (POLR2C) – the third-largest subunit. Exists as a heterodimer with another polymerase subunit, POLR2J forming a core subassembly. RPB3 strongly interacts with RPB1-5, 7, 10–12.[12]
- RNA polymerase II subunit B4 (RPB4) – encoded by the POLR2D gene[16] is the fourth-largest subunit and may have a stress protective role.
- RPB5 – In humans is encoded by the POLR2E gene. Two molecules of this subunit are present in each RNA polymerase II.[17] RPB5 strongly interacts with RPB1, RPB3, and RPB6.[12]
- RPB6 (POLR2F) – forms a structure with at least two other subunits that stabilizes the transcribing polymerase on the DNA template.[18]
- RPB7 – encoded by POLR2G and may play a role in regulating polymerase function.[19] RPB7 interacts strongly with RPB1 and RPB5.[12]
- RPB8 (POLR2H) – interacts with subunits RPB1-3, 5, and 7.[12]
- RPB9 – The groove in which the DNA template is transcribed into RNA is composed of RPB9 (POLR2I) and RPB1.
- RPB10 – the product of gene POLR2L. It interacts with RPB1-3 and 5, and strongly with RPB3.[12]
- RPB11 – the RPB11 subunit is itself composed of three subunits in humans: POLR2J (RPB11-a), POLR2J2 (RPB11-b), and POLR2J3[20] (RPB11-c).
- RPB12 – Also interacts with RPB3 is RPB12 (POLR2K).[12]
Assembly
RPB3 is involved in RNA polymerase II assembly.[21] A subcomplex of RPB2 and RPB3 appears soon after subunit synthesis.[21] This complex subsequently interacts with RPB1.[21] RPB3, RPB5, and RPB7 interact with themselves to form homodimers, and RPB3 and RPB5 together are able to contact all of the other RPB subunits, except RPB9.[12] Only RPB1 strongly binds to RPB5.[12] The RPB1 subunit also contacts RPB7, RPB10, and more weakly but most efficiently with RPB8.[12] Once RPB1 enters the complex, other subunits such as RPB5 and RPB7 can enter, where RPB5 binds to RPB6 and RPB8 and RPB3 brings in RPB10, RPB 11, and RPB12.[12] RPB4 and RPB9 may enter once most of the complex is assembled. RPB4 forms a complex with RPB7.[12]
Kinetics
Bacterial RNA polymerase, a relative of RNA Polymerase II, switches between inactivated and activated states by translocating back and forth along the DNA.[23] Concentrations of [NTP]eq = 10 μM GTP, 10 μM UTP, 5 μM ATP and 2.5 μM CTP, produce a mean elongation rate, turnover number, of ~1 bp (NTP)−1 for bacterial RNAP, a relative of RNA polymerase II.[23]

RNA polymerase II undergoes extensive co-transcriptional pausing during transcription elongation.[24][25] This pausing is especially pronounced at nucleosomes, and arises in part through the polymerase entering a transcriptionally incompetent backtracked state.[24] The duration of these pauses ranges from seconds to minutes or longer, and exit from long-lived pauses can be promoted by elongation factors such as TFIIS.[26] In turn, the transcription rate influences whether the histones of transcribed nucleosomes are evicted from chromatin, or reinserted behind the transcribing polymerase.[27]
Alpha-Amanitin
RNA polymerase II is inhibited by α-Amanitin[28] and other amatoxins. α-Amanitin is a highly poisonous substance found in many mushrooms.[5] The mushroom poison has different effects on each of the RNA Polymerases: I, II, III. RNAP I is completely unresponsive to the substance and will function normally while RNAP III has a moderate sensitivity. RNAP II, however, is completely inhibited by the toxin. Alpha-Amanitin inhibits RNAP II by strong interactions in the enzyme's "funnel", "cleft", and the key "bridge α-helix" regions of the RPB-1 subunit.[29]
Holoenzyme
RNA polymerase II holoenzyme is a form of eukaryotic RNA polymerase II that is recruited to the promoters of protein-coding genes in living cells.[11] It consists of RNA polymerase II, a subset of general transcription factors, and regulatory proteins known as SRB proteins.
Part of the assembly of the holoenzyme is referred to as the
Control by chromatin structure
This is an outline of an example mechanism of yeast cells by which
This pathway gives examples of regulation at these points of transcription:
- Pre-initiation (promotion by Bre1, histone modification)
- Initiation (promotion by TFIIH, Pol II modification and promotion by COMPASS, histone modification)
- Elongation (promotion by Set2, Histone Modification)
This refers to various stages of the process as regulatory steps. It has not been proven that they are used for regulation, but is very likely they are.
RNA Pol II elongation promoters can be summarised in 3 classes.
- Drug/sequence-dependent arrest-affected factors (Various interfering proteins)
- Chromatin structure-oriented factors (Histone posttranscriptional modifiers, e.g., Histone Methyltransferases)
- RNA Pol II catalysis-improving factors (Various interfering proteins and Pol II cofactors; see RNA polymerase II).
Transcription mechanisms
- Chromatin structure oriented factors:
(HMTs (Histone MethylTransferases)):
COMPASS§† – (COMplex of Proteins ASsociated with Set1) – Methylates lysine 4 of histone H3: Is responsible of repression/silencing of transcription. A normal part of cell growth and transcription regulation within RNAP II.[30] - Set2 – Methylates lysine 36 of histone H3: Set2 is involved in regulation transcription elongation through its direct contact with the CTD.[31]
(interesting irrelevant example: Dot1*‡ – Methylates lysine 79 of histone H3.) - Bre1 – Ubiquinates (adds ubiquitin to) lysine 123 of histone H2B. Associated with pre-initiation and allowing RNA Pol II binding.
C-terminal Domain
The
Phosphorylation of the CTD
RNA Polymerase II exists in two forms unphosphorylated and phosphorylated, IIA and IIO respectively.
Transcription coupled recombinational repair
Oxidative DNA damage may block RNA polymerase II transcription and cause strand breaks. An RNA templated transcription-associated recombination process has been described that can protect against DNA damage.[34] During the G1/G0 stages of the cell cycle, cells exhibit assembly of homologous recombination factors at double-strand breaks within actively transcribed regions. It appears that transcription is coupled to repair of DNA double-strand breaks by RNA templated homologous recombination. This repair process efficiently and accurately rejoins double-strand breaks in genes being actively transcribed by RNA polymerase II.
See also
- Eukaryotic transcription
- Post-transcriptional modification
- RNA polymerase I
- RNA polymerase II holoenzyme
- RNA polymerase III
- Transcription (genetics)
References
- PMID 10611681.
- PMID 15145350.
- ^ PMID 1883205.
- PMID 16765890.
- ^ OCLC 789601172.
- S2CID 4283528.
- S2CID 4283528.
- S2CID 244660781.
- PMID 15591044.
- PMID 2197989.
- ^ PMID 9774381.
- ^ PMID 9201987.
- ^ PMID 7498740.
- ^ "Entrez Gene: POLR2A polymerase (RNA) II (DNA directed) polypeptide A, 220kDa".
- ^ "Entrez Gene: POLR2B polymerase (RNA) II (DNA directed) polypeptide B, 140kDa".
- PMID 9528765.
- ^ "Entrez Gene: POLR2E polymerase (RNA) II (DNA directed) polypeptide E, 25kDa".
- ^ "Entrez Gene: POLR2F polymerase (RNA) II (DNA directed) polypeptide F".
- ^ "Entrez Gene: POLR2G polymerase (RNA) II (DNA directed) polypeptide G".
- ^ "POLR2J3 polymerase (RNA) II (DNA directed) polypeptide J3".
- ^ PMID 1715023.
- PMID 22511879.
- ^ PMID 16284617.
- ^ PMID 19644123.
- PMID 21248844.
- S2CID 4310108.
- PMID 22081017.
- PMID 18538653.
- PMID 15096519.
- PMID 11751634.
- PMID 12511561.
- S2CID 4418258.
- ^ PMID 18457900.
- PMID 27233112.
External links
(Wayback Machine copy)
- RNA+Polymerase+II at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
