RNA-Seq

RNA-Seq (named as an abbreviation of RNA sequencing) is a technique that uses
Specifically, RNA-Seq facilitates the ability to look at
Prior to RNA-Seq, gene expression studies were done with hybridization-based

Methods
Library preparation

The general steps to prepare a complementary DNA (cDNA) library for sequencing are described below, but often vary between platforms.[10][3][11]
- RNA Isolation: RNA is isolated from tissue and mixed with Deoxyribonuclease (DNase). DNase reduces the amount of genomic DNA. The amount of RNA degradation is checked with gel and capillary electrophoresis and is used to assign an RNA integrity number to the sample. This RNA quality and the total amount of starting RNA are taken into consideration during the subsequent library preparation, sequencing, and analysis steps.
- RNA selection/depletion: To analyze signals of interest, the isolated RNA can either be kept as is, enriched for RNA with miRNA, can be further isolated through size selection with exclusion gels, magnetic beads, or commercial kits.
- cDNA synthesis: RNA is DNA polymerases) and leverage more mature DNA sequencing technology. Amplification subsequent to reverse transcription results in loss of strandedness, which can be avoided with chemical labeling or single molecule sequencing. Fragmentation and size selection are performed to purify sequences that are the appropriate length for the sequencing machine. The RNA, cDNA, or both are fragmented with enzymes, sonication, or nebulizers. Fragmentation of the RNA reduces 5' bias of randomly primed-reverse transcription and the influence of primer binding sites,[13] with the downside that the 5' and 3' ends are converted to DNA less efficiently. Fragmentation is followed by size selection, where either small sequences are removed or a tight range of sequence lengths are selected. Because small RNAs like miRNAsare lost, these are analyzed independently. The cDNA for each experiment can be indexed with a hexamer or octamer barcode, so that these experiments can be pooled into a single lane for multiplexed sequencing.
| Strategy | Predominant type of RNA | Ribosomal RNA content | Unprocessed RNA content | Isolation method |
|---|---|---|---|---|
| Total RNA | All | High | High | None |
| PolyA selection | Coding | Low | Low | Hybridization with poly(dT) oligomers |
| rRNA depletion | Coding, noncoding | Low | High | Removal of oligomers complementary to rRNA |
| RNA capture | Targeted | Low | Moderate | Hybridization with probes complementary to desired transcripts |
Complementary DNA sequencing (cDNA-Seq)
The cDNA library derived from RNA biotypes is then sequenced into a computer-readable format. There are many high-throughput sequencing technologies for cDNA sequencing including platforms developed by Illumina, Thermo Fisher, BGI/MGI, PacBio, and Oxford Nanopore Technologies.[18] For Illumina short-read sequencing, a common technology for cDNA sequencing, adapters are ligated to the cDNA, DNA is attached to a flow cell, clusters are generated through cycles of bridge amplification and denaturing, and sequence-by-synthesis is performed in cycles of complementary strand synthesis and laser excitation of bases with reversible terminators. Sequencing platform choice and parameters are guided by experimental design and cost. Common experimental design considerations include deciding on the sequencing length, sequencing depth, use of single versus paired-end sequencing, number of replicates, multiplexing, randomization, and spike-ins.[19]
Small RNA/non-coding RNA sequencing
When sequencing RNA other than mRNA, the library preparation is modified. The cellular RNA is selected based on the desired size range. For small RNA targets, such as
Direct RNA sequencing

Because converting RNA into cDNA, ligation, amplification, and other sample manipulations have been shown to introduce biases and artifacts that may interfere with both the proper characterization and quantification of transcripts,[20] single molecule direct RNA sequencing has been explored by companies including Helicos (bankrupt), Oxford Nanopore Technologies,[21] and others. This technology sequences RNA molecules directly in a massively-parallel manner.
Single-molecule real-time RNA sequencing
Massively parallel single molecule direct RNA-Seq has been explored as an alternative to traditional RNA-Seq, in which RNA-to-cDNA conversion, ligation, amplification, and other sample manipulation steps may introduce biases and artifacts.[22] Technology platforms that perform single-molecule real-time RNA-Seq include Oxford Nanopore Technologies (ONT) Nanopore sequencing,[21] PacBio IsoSeq, and Helicos (bankrupt). Sequencing RNA in its native form preserves modifications like methylation, allowing them to be investigated directly and simultaneously.[21] Another benefit of single-molecule RNA-Seq is that transcripts can be covered in full length, allowing for higher confidence isoform detection and quantification compared to short-read sequencing. Traditionally, single-molecule RNA-Seq methods have higher error rates compared to short-read sequencing, but newer methods like ONT direct RNA-Seq limit errors by avoiding fragmentation and cDNA conversion. Recent uses of ONT direct RNA-Seq for differential expression in human cell populations have demonstrated that this technology can overcome many limitations of short and long cDNA sequencing.[23]
Single-cell RNA sequencing (scRNA-Seq)
Standard methods such as microarrays and standard bulk RNA-Seq analysis analyze the expression of RNAs from large populations of cells. In mixed cell populations, these measurements may obscure critical differences between individual cells within these populations.[24][25]
Single-cell RNA sequencing (scRNA-Seq) provides the expression profiles of individual cells. Although it is not possible to obtain complete information on every RNA expressed by each cell, due to the small amount of material available, patterns of gene expression can be identified through gene clustering analyses. This can uncover the existence of rare cell types within a cell population that may never have been seen before. For example, rare specialized cells in the lung called pulmonary ionocytes that express the Cystic fibrosis transmembrane conductance regulator were identified in 2018 by two groups performing scRNA-Seq on lung airway epithelia.[26][27]
Experimental procedures

Current scRNA-Seq protocols involve the following steps: isolation of single cell and RNA,
Challenges for scRNA-Seq include preserving the initial relative abundance of mRNA in a cell and identifying rare transcripts.[32] The reverse transcription step is critical as the efficiency of the RT reaction determines how much of the cell's RNA population will be eventually analyzed by the sequencer. The processivity of reverse transcriptases and the priming strategies used may affect full-length cDNA production and the generation of libraries biased toward the 3’ or 5' end of genes.
In the amplification step, either PCR or in vitro transcription (IVT) is currently used to amplify cDNA. One of the advantages of PCR-based methods is the ability to generate full-length cDNA. However, different PCR efficiency on particular sequences (for instance, GC content and snapback structure) may also be exponentially amplified, producing libraries with uneven coverage. On the other hand, while libraries generated by IVT can avoid PCR-induced sequence bias, specific sequences may be transcribed inefficiently, thus causing sequence drop-out or generating incomplete sequences.[33][24] Several scRNA-Seq protocols have been published: Tang et al.,[34] STRT,[35] SMART-seq,[36] CEL-seq,[37] RAGE-seq,[38] Quartz-seq[39] and C1-CAGE.[40] These protocols differ in terms of strategies for reverse transcription, cDNA synthesis and amplification, and the possibility to accommodate sequence-specific barcodes (i.e. UMIs) or the ability to process pooled samples.[41]
In 2017, two approaches were introduced to simultaneously measure single-cell mRNA and protein expression through oligonucleotide-labeled antibodies known as REAP-seq,[42] and CITE-seq.[43]
Applications
scRNA-Seq is becoming widely used across biological disciplines including Development,
scRNA-Seq has provided considerable insight into the development of embryos and organisms, including the worm
Experimental considerations
A variety of
- Tissue specificity: Gene expression varies within and between tissues, and RNA-Seq measures this mix of cell types. This may make it difficult to isolate the biological mechanism of interest. Single cell sequencingcan be used to study each cell individually, mitigating this issue.
- Time dependence: Gene expression changes over time, and RNA-Seq only takes a snapshot. Time course experiments can be performed to observe changes in the transcriptome.
- Coverage (also known as depth): RNA harbors the same mutations observed in DNA, and detection requires deeper coverage. With high enough coverage, RNA-Seq can be used to estimate the expression of each allele. This may provide insight into phenomena such as cis-regulatory effects. The depth of sequencing required for specific applications can be extrapolated from a pilot experiment.[57]
- Data generation artifacts (also known as technical variance): The reagents (e.g., library preparation kit), personnel involved, and type of sequencer (e.g., latent variables (typically principal component analysis or factor analysis) and subsequently correcting for these variables.[58]
- Data management: A single RNA-Seq experiment in humans is usually 1-5 Gb (compressed), or more when including intermediate files.[59] This large volume of data can pose storage issues. One solution is compressing the data using multi-purpose computational schemas (e.g., gzip) or genomics-specific schemas. The latter can be based on reference sequences or de novo. Another solution is to perform microarray experiments, which may be sufficient for hypothesis-driven work or replication studies (as opposed to exploratory research).
Analysis

Transcriptome assembly
Two methods are used to assign raw sequence reads to genomic features (i.e., assemble the transcriptome):
- De novo: This approach does not require a Velvet[63]), Bridger,[64] and rnaSPAdes.[65] Paired-end and long-read sequencing of the same sample can mitigate the deficits in short read sequencing by serving as a template or skeleton. Metrics to assess the quality of a de novo assembly include median contig length, number of contigs and N50.[66]

- Genome guided: This approach relies on the same methods used for DNA alignment, with the additional complexity of aligning reads that cover non-continuous portions of the reference genome.[67] These non-continuous reads are the result of sequencing spliced transcripts (see figure). Typically, alignment algorithms have two steps: 1) align short portions of the read (i.e., seed the genome), and 2) use dynamic programming to find an optimal alignment, sometimes in combination with known annotations. Software tools that use genome-guided alignment include Bowtie,[68] TopHat (which builds on BowTie results to align splice junctions),[69][70] Subread,[71] STAR,[67] HISAT2,[72] and GMAP.[73] The output of genome guided alignment (mapping) tools can be further used by tools such as Cufflinks[70] or StringTie[74] to reconstruct contiguous transcript sequences (i.e., a FASTA file). The quality of a genome guided assembly can be measured with both 1) de novo assembly metrics (e.g., N50) and 2) comparisons to known transcript, splice junction, genome, and protein sequences using precision, recall, or their combination (e.g., F1 score).[66] In addition, in silico assessment could be performed using simulated reads.[75][76]
A note on assembly quality: The current consensus is that 1) assembly quality can vary depending on which metric is used, 2) assembly tools that scored well in one species do not necessarily perform well in the other species, and 3) combining different approaches might be the most reliable.[77][78][79]
Gene expression quantification
Expression is quantified to study cellular changes in response to external stimuli, differences between healthy and
Expression is quantified by counting the number of reads that mapped to each locus in the transcriptome assembly step. Expression can be quantified for exons or genes using contigs or reference transcript annotations.[10] These observed RNA-Seq read counts have been robustly validated against older technologies, including expression microarrays and qPCR.[57][81] Tools that quantify counts are HTSeq,[82] FeatureCounts,[83] Rcount,[84] maxcounts,[85] FIXSEQ,[86] and Cuffquant. These tools determine read counts from aligned RNA-Seq data, but alignment-free counts can also be obtained with Sailfish[87] and Kallisto.[88] The read counts are then converted into appropriate metrics for hypothesis testing, regressions, and other analyses. Parameters for this conversion are:
- Sequencing depth/coverage: Although depth is pre-specified when conducting multiple RNA-Seq experiments, it will still vary widely between experiments.[89] Therefore, the total number of reads generated in a single experiment is typically normalized by converting counts to fragments, reads, or counts per million mapped reads (FPM, RPM, or CPM). The difference between RPM and FPM was historically derived during the evolution from single-end sequencing of fragments to paired-end sequencing. In single-end sequencing, there is only one read per fragment (i.e., RPM = FPM). In paired-end sequencing, there are two reads per fragment (i.e., RPM = 2 x FPM). Sequencing depth is sometimes referred to as library size, the number of intermediary cDNA molecules in the experiment.
- Gene length: Longer genes will have more fragments/reads/counts than shorter genes if transcript expression is the same. This is adjusted by dividing the FPM by the length of a feature (which can be a gene, transcript, or exon), resulting in the metric fragments per kilobase of feature per million mapped reads (FPKM).[90] When looking at groups of features across samples, FPKM is converted to transcripts per million (TPM) by dividing each FPKM by the sum of FPKMs within a sample.[91][92][93]
- Total sample RNA output: Because the same amount of RNA is extracted from each sample, samples with more total RNA will have less RNA per gene. These genes appear to have decreased expression, resulting in false positives in downstream analyses.[89] Normalization strategies including quantile, DESeq2, TMM and Median Ratio attempt to account for this difference by comparing a set of non-differentially expressed genes between samples and scaling accordingly.[94]
- Variance for each gene's expression: is modeled to account for sampling error (important for genes with low read counts), increase power, and decrease false positives. Variance can be estimated as a normal, Poisson, or negative binomial distribution[95][96][97] and is frequently decomposed into technical and biological variance.
Spike-ins for absolute quantification and detection of genome-wide effects
RNA spike-ins are samples of RNA at known concentrations that can be used as gold standards in experimental design and during downstream analyses for absolute quantification and detection of genome-wide effects.
- Absolute quantification: Absolute quantification of gene expression is not possible with most RNA-Seq experiments, which quantify expression relative to all transcripts. It is possible by performing RNA-Seq with spike-ins, samples of RNA at known concentrations. After sequencing, read counts of spike-in sequences are used to determine the relationship between each gene's read counts and absolute quantities of biological fragments.Xenopus tropicalis embryos to determine transcription kinetics.[99]
- Detection of genome-wide effects: Changes in global regulators including transcription factors (e.g., MYC), acetyltransferase complexes, and nucleosome positioning are not congruent with normalization assumptions and spike-in controls can offer precise interpretation.[100][101]
Differential expression
The simplest but often most powerful use of RNA-Seq is finding differences in gene expression between two or more conditions (e.g., treated vs not treated); this process is called differential expression. The outputs are frequently referred to as differentially expressed genes (DEGs) and these genes can either be up- or down-regulated (i.e., higher or lower in the condition of interest). There are many tools that perform differential expression. Most are run in R, Python, or the Unix command line. Commonly used tools include DESeq,[96] edgeR,[97] and voom+limma,[95][102] all of which are available through R/Bioconductor.[103][104] These are the common considerations when performing differential expression:
- Inputs: Differential expression inputs include (1) an RNA-Seq expression matrix (M genes x N samples) and (2) a unsupervised machine learning approaches including principal component, surrogate variable,[105] and PEER[58]analyses. Hidden variable analyses are often employed for human tissue RNA-Seq data, which typically have additional artifacts not captured in the metadata (e.g., ischemic time, sourcing from multiple institutions, underlying clinical traits, collecting data across many years with many personnel).
- Methods: Most tools use non-parametric statistics to identify differentially expressed genes, and are either based on read counts mapped to a reference genome (DESeq2, limma, edgeR) or based on read counts derived from alignment-free quantification (sleuth,[106] Cuffdiff,[107] Ballgown[108]).[109] Following regression, most tools employ either familywise error rate (FWER) or false discovery rate (FDR) p-value adjustments to account for multiple hypotheses(in human studies, ~20,000 protein-coding genes or ~50,000 biotypes).
- Outputs: A typical output consists of rows corresponding to the number of genes and at least three columns, each gene's log log-transform of the ratio in expression between conditions, a measure of effect size), p-value, and p-value adjusted for multiple comparisons. Genes are defined as biologically meaningful if they pass cut-offs for effect size (log fold change) and statistical significance. These cut-offs should ideally be specified a priori, but the nature of RNA-Seq experiments is often exploratory so it is difficult to predict effect sizes and pertinent cut-offs ahead of time.
- Pitfalls: The raison d'etre for these complex methods is to avoid the myriad of pitfalls that can lead to ) into dates or floating point numbers.
- Choice of tools and benchmarking: There are numerous efforts that compare the results of these tools, with DESeq2 tending to moderately outperform other methods.[111][112][113][114][19][109][115][116] As with other methods, benchmarking consists of comparing tool outputs to each other and known gold standards.
Downstream analyses for a list of differentially expressed genes come in two flavors, validating observations and making biological inferences. Owing to the pitfalls of differential expression and RNA-Seq, important observations are replicated with (1) an orthogonal method in the same samples (like
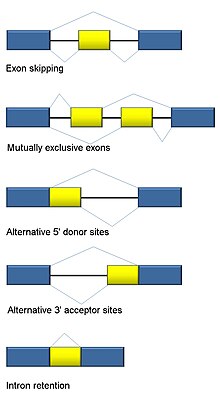
Alternative splicing
RNA splicing is integral to eukaryotes and contributes significantly to protein regulation and diversity, occurring in >90% of human genes.[118] There are multiple alternative splicing modes: exon skipping (most common splicing mode in humans and higher eukaryotes), mutually exclusive exons, alternative donor or acceptor sites, intron retention (most common splicing mode in plants, fungi, and protozoa), alternative transcription start site (promoter), and alternative polyadenylation.[118] One goal of RNA-Seq is to identify alternative splicing events and test if they differ between conditions. Long-read sequencing captures the full transcript and thus minimizes many of issues in estimating isoform abundance, like ambiguous read mapping. For short-read RNA-Seq, there are multiple methods to detect alternative splicing that can be classified into three main groups:[119][91][120]
- Count-based (also event-based, differential splicing): estimate exon retention. Examples are DEXSeq,[121] MATS,[122] and SeqGSEA.[123]
- Isoform-based (also multi-read modules, differential isoform expression): estimate isoform abundance first, and then relative abundance between conditions. Examples are Cufflinks 2[124] and DiffSplice.[125]
- Intron excision based: calculate alternative splicing using split reads. Examples are MAJIQ[126] and Leafcutter.[120]
Differential gene expression tools can also be used for differential isoform expression if isoforms are quantified ahead of time with other tools like RSEM.[127]
Coexpression networks
Coexpression networks are data-derived representations of genes behaving in a similar way across tissues and experimental conditions.
Variant discovery
RNA-Seq captures DNA variation, including
RNA editing (post-transcriptional alterations)
Having the matching genomic and transcriptomic sequences of an individual can help detect post-transcriptional edits (RNA editing).[3] A post-transcriptional modification event is identified if the gene's transcript has an allele/variant not observed in the genomic data.

Fusion gene detection
Caused by different structural modifications in the genome, fusion genes have gained attention because of their relationship with cancer.[139] The ability of RNA-Seq to analyze a sample's whole transcriptome in an unbiased fashion makes it an attractive tool to find these kinds of common events in cancer.[4]
The idea follows from the process of aligning the short transcriptomic reads to a reference genome. Most of the short reads will fall within one complete exon, and a smaller but still large set would be expected to map to known exon-exon junctions. The remaining unmapped short reads would then be further analyzed to determine whether they match an exon-exon junction where the exons come from different genes. This would be evidence of a possible fusion event, however, because of the length of the reads, this could prove to be very noisy. An alternative approach is to use paired-end reads, when a potentially large number of paired reads would map each end to a different exon, giving better coverage of these events (see figure). Nonetheless, the end result consists of multiple and potentially novel combinations of genes providing an ideal starting point for further validation.
Copy number alteration
Copy number alteration (CNA) analyses are commonly used in cancer studies. Gain and loss of the genes have signalling pathway implications and are a key biomarker of molecular dysfunction in oncology. Calling the CNA information from RNA-Seq data is not straightforward because of the differences in gene expression, which lead to the read depth variance of different magnitudes across genes. Due to these difficulties, most of these analyses are usually done using whole-genome sequencing / whole-exome sequencing (WGS/WES). But advanced bioinformatics tools can call CNA from RNA-Seq.[140]
Other emerging analysis and applications
The applications of RNA-Seq are growing day by day. Other new application of RNA-Seq includes detection of microbial contaminants,[141] determining cell type abundance (cell type deconvolution),[8] measuring the expression of TEs and Neoantigen prediction etc.[8]
History

RNA-Seq was first developed in mid 2000s with the advent of next-generation sequencing technology.
Applications to medicine
RNA-Seq has the potential to identify new disease biology, profile biomarkers for clinical indications, infer druggable pathways, and make genetic diagnoses. These results could be further personalized for subgroups or even individual patients, potentially highlighting more effective prevention, diagnostics, and therapy. The feasibility of this approach is in part dictated by costs in money and time; a related limitation is the required team of specialists (bioinformaticians, physicians/clinicians, basic researchers, technicians) to fully interpret the huge amount of data generated by this analysis.[151]
Large-scale sequencing efforts
A lot of emphasis has been given to RNA-Seq data after the Encyclopedia of DNA Elements (ENCODE) and The Cancer Genome Atlas (TCGA) projects have used this approach to characterize dozens of cell lines[152] and thousands of primary tumor samples,[153] respectively. ENCODE aimed to identify genome-wide regulatory regions in different cohort of cell lines and transcriptomic data are paramount to understand the downstream effect of those epigenetic and genetic regulatory layers. TCGA, instead, aimed to collect and analyze thousands of patient's samples from 30 different tumor types to understand the underlying mechanisms of malignant transformation and progression. In this context RNA-Seq data provide a unique snapshot of the transcriptomic status of the disease and look at an unbiased population of transcripts that allows the identification of novel transcripts, fusion transcripts and non-coding RNAs that could be undetected with different technologies.
See also
- Transcriptomics
- DNA microarray
- List of RNA-Seq bioinformatics tools
References
![]() This article was submitted to WikiJournal of Science for external academic peer review in 2019 (reviewer reports). The updated content was reintegrated into the Wikipedia page under a CC-BY-SA-3.0 license (2021). The version of record as reviewed is:
Felix Richter, et al. (17 May 2021). "A broad introduction to RNA-Seq" (PDF). WikiJournal of Science. 4 (2): 4.
This article was submitted to WikiJournal of Science for external academic peer review in 2019 (reviewer reports). The updated content was reintegrated into the Wikipedia page under a CC-BY-SA-3.0 license (2021). The version of record as reviewed is:
Felix Richter, et al. (17 May 2021). "A broad introduction to RNA-Seq" (PDF). WikiJournal of Science. 4 (2): 4. {{cite journal}}: CS1 maint: unflagged free DOI (link
- PMID 28545146.
- PMID 22830413.
- ^ PMID 19015660.
- ^ PMID 19136943.
- PMID 22836135.
- PMID 30999927.
- PMID 24578530.
- ^ PMID 34329375.
- PMID 25870306.
- ^ PMID 26248053.
- ^ "RNA-seqlopedia". rnaseq.uoregon.edu. Retrieved 8 February 2017.
- PMID 18611170.
- ^ S2CID 205418589.
- PMID 29249332.
- PMID 24632678.
- PMID 25339126.
- S2CID 83424788.
- PMID 32733532.
- ^ PMID 26813401.
- PMID 16503995.
- ^ S2CID 3589823.
- PMID 16503995.
- S2CID 220975367.
- ^ S2CID 500845."
- PMID 26000846.
- PMID 30069044.
- PMID 30069046.
- PMID 29534489.
- PMID 26000487.
- PMID 26000488.
- S2CID 6765530.
- PMID 24832513."
- S2CID 11575439.
- S2CID 16570747.
- PMID 21543516.
- PMID 22820318.
- PMID 22939981.
- PMID 31311926.
- PMID 23594475.
- PMID 30664627.
- PMID 29394315.
- S2CID 205285357.
- PMID 28759029.
- PMID 29608178.
- PMID 18695026.
- PMID 29606308.
- PMID 30388455.
- PMID 29476078.
- PMID 26343579.
- PMID 28818938.
- PMID 29674432.
- PMID 29674431.
- PMID 29700229.
- PMID 29700225.
- PMID 29700227.
- ^ You J. "Science's 2018 Breakthrough of the Year: tracking development cell by cell". Science Magazine. American Association for the Advancement of Science.
- ^ PMID 19088194.
- ^ PMID 22343431.
- PMID 25649622.
- ^ PMID 21572440.
- ^ "De Novo Assembly Using Illumina Reads" (PDF). Retrieved 22 October 2016.
- ^ Oases: a transcriptome assembler for very short reads
- PMID 18349386.
- PMID 25723335.
- PMID 31494669.
- ^ PMID 25608678.
- ^ PMID 23104886.
- PMID 19261174.
- PMID 19289445.
- ^ PMID 22383036.
- PMID 23558742.
- PMID 25751142.
- PMID 15728110.
- PMID 25690850.
- PMID 27941783.
- PMID 24185836.
- PMID 23393030.
- PMID 23870653.
- PMID 31077315.
- PMID 12952525.
- PMID 25119138.
- PMID 25260700.
- PMID 24227677.
- PMID 25322836.
- PMID 24564404.
- PMID 24603409.
- PMID 24752080.
- S2CID 205282743.
- ^ PMID 20196867.
- PMID 20436464.
- ^ ].
- ^ "What the FPKM? A review of RNA-Seq expression units". The farrago. 8 May 2014. Retrieved 28 March 2018.
- S2CID 16752581.
- PMID 28334202.
- ^ PMID 24485249.
- ^ PMID 20979621.
- ^ PMID 19910308.
- PMID 23101633.
- PMID 26774488.
- PMID 26711261.
- PMID 23101621.
- PMID 25605792.
- ^ "Bioconductor - Open source software for bioinformatics".
- PMID 25633503.
- PMID 17907809.
- S2CID 15063247.
- PMID 23222703.
- PMID 25748911.
- ^ PMID 28680106.
- PMID 27552985.
- PMID 23497356.
- PMID 25268973.
- PMID 24300110.
- PMID 24020486.
- PMID 29267363.
- PMID 33184454.
- PMID 31114916.
- ^ S2CID 5184582.
- PMID 25511303.
- ^ PMID 29229983.
- PMID 22722343.
- PMID 22266656.
- PMID 24535097.
- PMID 23222703.
- PMID 23155066.
- PMID 26829591.
- S2CID 22706028.
- ^ S2CID 144447.
- ^ PMID 23376351.
- PMID 22556371.
- PMID 24244129.
- PMID 24951248.
- PMID 28298217.
- PMID 19505943.
- PMID 21478889.
- PMID 29022597.
- PMID 30903145.
- S2CID 212739959.
- S2CID 40770452.
- PMID 34329375.
- PMID 30999839.
- ^ "PubMed search: "RNA Seq" OR "RNA-Seq" OR "RNA sequencing" OR "RNASeq"". PubMed. Retrieved 20 June 2021.
- ^ "PubMed search: ("RNA Seq" OR "RNA-Seq" OR "RNA sequencing" OR "RNASeq") AND "Medicine"". PubMed. Retrieved 20 June 2021.
- PMID 26353759.
- PMID 17010196.
- PMID 17062153.
- PMID 17095711.
- PMID 17351049.
- PMID 18451266.
- .
- S2CID 27632439.
- ^ "ENCODE Data Matrix". Retrieved 28 July 2013.
- ^ "The Cancer Genome Atlas – Data Portal". Retrieved 28 July 2013.
Further reading
- Taguchi Y (2019). "Comparative Transcriptomics Analysis". Encyclopedia of Bioinformatics and Computational Biology. pp. 814–818. S2CID 65302519.
External links
- Cresko B, Voelker R, Small C (2001). Bassham S, Catchen J (eds.). "RNA-seqlopedia". University of Oregon.: a high-level guide to designing and implementing an RNA-Seq experiment.
