Drug design
Drug design, often referred to as rational drug design or simply
Definition
The phrase "drug design" is similar to ligand design (i.e., design of a molecule that will bind tightly to its target).[6] Although design techniques for prediction of binding affinity are reasonably successful, there are many other properties, such as bioavailability, metabolic half-life, and side effects, that first must be optimized before a ligand can become a safe and effictive drug. These other characteristics are often difficult to predict with rational design techniques.
Due to high attrition rates, especially during
Drug targets
A
Most commonly, drugs are
Drug discovery
Phenotypic drug discovery
Phenotypic drug discovery is a traditional drug discovery method, also known as forward pharmacology or classical pharmacology. It uses the process of phenotypic screening on collections of synthetic small molecules, natural products, or extracts within chemical libraries to pinpoint substances exhibiting beneficial therapeutic effects. This method is to first discover the in vivo or in vitro functional activity of drugs (such as extract drugs or natural products), and then perform target identification. Phenotypic discovery uses a practical and target-independent approach to generate initial leads, aiming to discover pharmacologically active compounds and therapeutics that operate through novel drug mechanisms.[17] This method allows the exploration of disease phenotypes to find potential treatments for conditions with unknown, complex, or multifactorial origins, where the understanding of molecular targets is insufficient for effective intervention.[18]
Rational drug discovery
Rational drug design (also called reverse pharmacology) begins with a hypothesis that modulation of a specific biological target may have therapeutic value. In order for a biomolecule to be selected as a drug target, two essential pieces of information are required. The first is evidence that modulation of the target will be disease modifying. This knowledge may come from, for example, disease linkage studies that show an association between mutations in the biological target and certain disease states.[19] The second is that the target is capable of binding to a small molecule and that its activity can be modulated by the small molecule.[20]
Once a suitable target has been identified, the target is normally cloned and produced and purified. The purified protein is then used to establish a screening assay. In addition, the three-dimensional structure of the target may be determined.
The search for small molecules that bind to the target is begun by screening libraries of potential drug compounds. This may be done by using the screening assay (a "wet screen"). In addition, if the structure of the target is available, a
Due to the large number of drug properties that must be simultaneously optimized during the design process, multi-objective optimization techniques are sometimes employed.[24] Finally because of the limitations in the current methods for prediction of activity, drug design is still very much reliant on serendipity[25] and bounded rationality.[26]
Computer-aided drug design
The most fundamental goal in drug design is to predict whether a given molecule will bind to a target and if so how strongly.
Molecular mechanics methods may also be used to provide semi-quantitative prediction of the binding affinity. Also, knowledge-based
Ideally, the computational method will be able to predict affinity before a compound is synthesized and hence in theory only one compound needs to be synthesized, saving enormous time and cost. The reality is that present computational methods are imperfect and provide, at best, only qualitatively accurate estimates of affinity. In practice, it requires several iterations of design, synthesis, and testing before an optimal drug is discovered. Computational methods have accelerated discovery by reducing the number of iterations required and have often provided novel structures.[30][31]
Computer-aided drug design may be used at any of the following stages of drug discovery:
- hit identification using virtual screening (structure- or ligand-based design)
- QSAR, etc.)
- lead optimization of other pharmaceutical properties while maintaining affinity

In order to overcome the insufficient prediction of binding affinity calculated by recent scoring functions, the protein-ligand interaction and compound 3D structure information are used for analysis. For structure-based drug design, several post-screening analyses focusing on protein-ligand interaction have been developed for improving enrichment and effectively mining potential candidates:
- Consensus scoring[32][33]
- Selecting candidates by voting of multiple scoring functions
- May lose the relationship between protein-ligand structural information and scoring criterion
- Cluster analysis[34][35]
- Represent and cluster candidates according to protein-ligand 3D information
- Needs meaningful representation of protein-ligand interactions.
Types
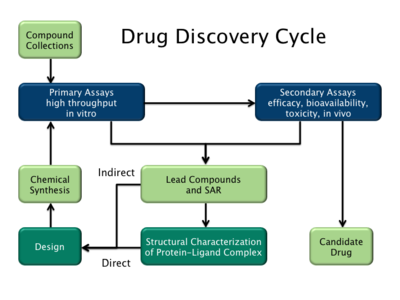
There are two major types of drug design. The first is referred to as ligand-based drug design and the second, structure-based drug design.[2]
Ligand-based
Ligand-based drug design (or indirect drug design) relies on knowledge of other molecules that bind to the biological target of interest. These other molecules may be used to derive a
Structure-based
Structure-based drug design (or direct drug design) relies on knowledge of the
Current methods for structure-based drug design can be divided roughly into three main categories.[40] The first method is identification of new ligands for a given receptor by searching large databases of 3D structures of small molecules to find those fitting the binding pocket of the receptor using fast approximate docking programs. This method is known as virtual screening.
A second category is de novo design of new ligands. In this method, ligand molecules are built up within the constraints of the binding pocket by assembling small pieces in a stepwise manner. These pieces can be either individual atoms or molecular fragments. The key advantage of such a method is that novel structures, not contained in any database, can be suggested.[41][42][43] A third method is the optimization of known ligands by evaluating proposed analogs within the binding cavity.[40]
Binding site identification
Scoring functions
Structure-based drug design attempts to use the structure of proteins as a basis for designing new ligands by applying the principles of
One early general-purposed empirical scoring function to describe the binding energy of ligands to receptors was developed by Böhm.[46][47] This empirical scoring function took the form:
where:
- ΔG0 – empirically derived offset that in part corresponds to the overall loss of translational and rotational entropy of the ligand upon binding.
- ΔGhb – contribution from hydrogen bonding
- ΔGionic – contribution from ionic interactions
- ΔGlip – contribution from lipophilic interactions where |Alipo| is surface area of lipophilic contact between the ligand and receptor
- ΔGrot – entropy penalty due to freezing a rotatable in the ligand bond upon binding
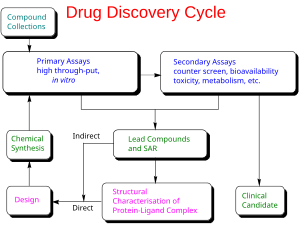
A more general thermodynamic "master" equation is as follows:[48]
where:
- desolvation – enthalpic penalty for removing the ligand from solvent
- motion – entropic penalty for reducing the degrees of freedom when a ligand binds to its receptor
- configuration – conformational strain energy required to put the ligand in its "active" conformation
- interaction – enthalpic gain for "resolvating" the ligand with its receptor
The basic idea is that the overall binding free energy can be decomposed into independent components that are known to be important for the binding process. Each component reflects a certain kind of free energy alteration during the binding process between a ligand and its target receptor. The Master Equation is the linear combination of these components. According to Gibbs free energy equation, the relation between dissociation equilibrium constant, Kd, and the components of free energy was built.
Various computational methods are used to estimate each of the components of the master equation. For example, the change in polar surface area upon ligand binding can be used to estimate the desolvation energy. The number of rotatable bonds frozen upon ligand binding is proportional to the motion term. The configurational or strain energy can be estimated using molecular mechanics calculations. Finally the interaction energy can be estimated using methods such as the change in non polar surface, statistically derived potentials of mean force, the number of hydrogen bonds formed, etc. In practice, the components of the master equation are fit to experimental data using multiple linear regression. This can be done with a diverse training set including many types of ligands and receptors to produce a less accurate but more general "global" model or a more restricted set of ligands and receptors to produce a more accurate but less general "local" model.[49]
Examples
A particular example of rational drug design involves the use of three-dimensional information about biomolecules obtained from such techniques as X-ray crystallography and NMR spectroscopy. Computer-aided drug design in particular becomes much more tractable when there is a high-resolution structure of a target protein bound to a potent ligand. This approach to drug discovery is sometimes referred to as structure-based drug design. The first unequivocal example of the application of
Another case study in rational drug design is
Additional examples include:
- Many of the atypical antipsychotics
- H2-receptor antagonistfrom which the later members of the class were developed
- Selective NSAIDs
- Enfuvirtide, a peptide HIV entry inhibitor
- HIV integrase inhibitor[53]
- SSRIs (selective serotonin reuptake inhibitors), a class of antidepressants
- Zanamivir, an antiviral drug
Drug screening
Types of drug screening include phenotypic screening, high-throughput screening, and virtual screening. Phenotypic screening is characterized by the process of screening drugs using cellular or animal disease models to identify compounds that alter the phenotype and produce beneficial disease-related effects.[54][55] Emerging technologies in high-throughput screening substantially enhance processing speed and decrease the required detection volume.[56] Virtual screening is completed by computer, enabling a large number of molecules can be screened with a short cycle and low cost. Virtual screening uses a range of computational methods that empower chemists to reduce extensive virtual libraries into more manageable sizes.[57]
Case studies
- 5-HT3 antagonists
- Acetylcholine receptor agonists
- Angiotensin receptor antagonists
- Bcr-Abl tyrosine-kinase inhibitors
- Cannabinoid receptor antagonists
- CCR5 receptor antagonists
- Cyclooxygenase 2 inhibitors
- Dipeptidyl peptidase-4 inhibitors
- HIV protease inhibitors
- NK1 receptor antagonists
- Non-nucleoside reverse transcriptase inhibitors
- Nucleoside and nucleotide reverse transcriptase inhibitors
- PDE5 inhibitors
- Proton pump inhibitors
- Renin inhibitors
- Triptans
- TRPV1 antagonists
- c-Met inhibitors
Criticism
It has been argued that the highly rigid and focused nature of rational drug design suppresses serendipity in drug discovery.[58]
See also
References
- ISBN 978-0-415-28288-8.
- ^ ISBN 978-0521887236.
- ^ PMID 25450771.
- ^ PMID 29733895.
- PMID 25110827.
- S2CID 21550508.
- S2CID 25292436.
- PMID 12963322.
- PMID 19740696.
- S2CID 8872470.
- PMID 14522049.
- PMID 24981487.
- ISBN 978-1-59745-532-9.
- PMID 25916823.
- PMID 37892179.
- S2CID 27454546.
- PMID 32850117.
- S2CID 6180139.
- ISBN 9780123971760.
- ^ PMID 23082974.
- PMID 12565011.
- ISBN 978-0-12-374194-3.
- ISBN 978-3-527-67301-8.
- PMID 24050140.
- PMID 17117615.
- SSRN 604123.
- ISBN 978-1849731669.
- PMID 17554857.
- PMID 19128212.
- PMID 14643325.
- PMID 16722631.
- PMID 18831053.
- PMID 16426072.
- PMID 14711306.
- PMID 16426058.
- ISBN 978-0-9636817-6-8.
- ISBN 978-0521887236.
- ISBN 978-1-4020-4406-9.
- PMID 18428090.
- ^ S2CID 21314020.
- S2CID 59482623.
- S2CID 2549851.
- S2CID 1307935.
- ^ PMID 20166931.
- ISBN 978-1849731669.
- S2CID 2491616.
- PMID 25647463.
- PMID 8544170.
- ISBN 978-1849731669.
- PMID 8164249.
- ISBN 978-3-527-29343-8.
- S2CID 2728341.
- ^ "AutoDock's role in Developing the First Clinically-Approved HIV Integrase Inhibitor". Press Release. The Scripps Research Institute. 2007-12-17.
- PMID 24902068.
- S2CID 72519717.
- PMID 10959774.
- .
- PMID 18319418.
External links
- Drug+Design at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- [Drug Design Org](https://www.drugdesign.org/chapters/drug-design/)


![{\displaystyle {\begin{array}{lll}\Delta G_{\text{bind}}=-RT\ln K_{\text{d}}\\[1.3ex]K_{\text{d}}={\dfrac {[{\text{Ligand}}][{\text{Receptor}}]}{[{\text{Complex}}]}}\\[1.3ex]\Delta G_{\text{bind}}=\Delta G_{\text{desolvation}}+\Delta G_{\text{motion}}+\Delta G_{\text{configuration}}+\Delta G_{\text{interaction}}\end{array}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/ba49ddd9dec7415d129787213744ca1afcd2d021)