Repeated sequence (DNA)
Repeated sequences (also known as repetitive elements, repeating units or repeats) are short or long patterns of nucleic acids (DNA or RNA) that occur in multiple copies throughout the genome. In many organisms, a significant fraction of the genomic DNA is repetitive, with over two-thirds of the sequence consisting of repetitive elements in humans.[1] Some of these repeated sequences are necessary for maintaining important genome structures such as telomeres or centromeres.[2]
Repeated sequences are categorized into different classes depending on features such as structure, length, location, origin, and mode of multiplication. The disposition of repetitive elements throughout the genome can consist either in directly adjacent arrays called tandem repeats or in repeats dispersed throughout the genome called interspersed repeats.[3] Tandem repeats and interspersed repeats are further categorized into subclasses based on the length of the repeated sequence and/or the mode of multiplication.
While some repeated DNA sequences are important for cellular functioning and genome maintenance, other repetitive sequences can be harmful. Many repetitive DNA sequences have been linked to human diseases such as Huntington's disease and Friedreich's ataxia. Some repetitive elements are neutral and occur when there is an absence of selection for specific sequences depending on how transposition or crossing over occurs.[2] However, an abundance of neutral repeats can still influence genome evolution as they accumulate over time. Overall, repeated sequences are an important area of focus because they can provide insight into human diseases and genome evolution.[2]
History
In the 1950s,
Types and functions
Many repeat sequences are likely to be non-functional, decaying remnants of Transposable elements, these have been labelled "junk" or "selfish" DNA.[7][8][9] Nevertheless, occasionally some repeats may be exapted for other functions.[10]
Tandem repeats
Tandem repeats are repeated sequences which are directly adjacent to each other in the genome.[11] Tandem repeats may vary in the number of nucleotides comprising the repeated sequence, as well as the number of times the sequence repeats. When the repeating sequence is only 2–10 nucleotides long, the repeat is referred to as a short tandem repeat (STR) or microsatellite.[12] When the repeating sequence is 10–60 nucleotides long, the repeat is referred to as a minisatellite.[13] For minisatellites and microsatellites, the number of times the sequence repeats at a single locus can range from twice to hundreds of times.
Tandem repeats have a wide variety of biological functions in the genome. For example, minisatellites are often hotspots of meiotic homologous recombination in eukaryotic organisms.[14] Recombination is when two homologous chromosomes align, break, and rejoin to swap pieces. Recombination is important as a source of genetic diversity, as a mechanism for repairing damaged DNA, and a necessary step in the appropriate segregation of chromosomes in meiosis.[14] The presence of repeated sequence DNA makes it easier for areas of homology to align, thereby controlling when and where recombination occurs.
In addition to playing an important role in recombination, tandem repeats also play important structural roles in the genome. For example, telomeres are composed mainly of tandem TTAGGG repeats.[15] These repeats fold into highly organized G quadruplex structures which protect the ends of chromosomal DNA from degradation.[16] Repetitive elements are enriched in the middle of chromosomes as well. Centromeres are the highly compact regions of chromosomes which join sister chromatids together and also allow the mitotic spindle to attach and separate sister chromatids during cell division.[17] Centromeres are composed of a 177 base pair tandem repeat named the α-satellite repeat.[16] Pericentromeric heterochromatin, the DNA which surrounds the centromere and is important for structural maintenance, is composed of a mixture of different satellite subfamilies including the α-, β- and γ-satellites as well as HSATII, HSATIII, and sn5 repeats.[18][19]

Some repetitive sequences, such as those with structural roles discussed above, play roles necessary for proper biological functioning. Other tandem repeats have deleterious roles which drive diseases. Many other tandem repeats, however, have unknown or poorly understood functions.[20]
Interspersed repeats
Interspersed repeats are identical or similar DNA sequences which are found in different locations throughout the genome.[21] Interspersed repeats are distinguished from tandem repeats in that the repeated sequences are not directly adjacent to each other but instead may be scattered among different chromosomes or far apart on the same chromosome. Most interspersed repeats are transposable elements (TEs), mobile sequences which can be "cut and pasted" or "copied and pasted" into different places in the genome.[22] TEs were originally called "jumping genes" for their ability to move, yet this term is somewhat misleading as not all TEs are discrete genes.[23]
Transposable elements that are transcribed into RNA, reverse-transcribed into DNA, then reintegrated into the genome are called retrotransposons.[22] Just as tandem repeats are further subcategorized based on the length of the repeating sequence, there are many different types of retrotransposons. Long interspersed nuclear elements (LINEs) are typically 3-7 kilobases in length.[24] Short interspersed nuclear elements (SINEs) are typically 100-300 base pairs and no longer than 600 base pairs.[24] Long-terminal repeat retrotransposons (LTRs) are a third major class of retrotransposons and are characterized by highly repetitive sequences as the ends of the repeat.[22] When a transposable element does not proceed through RNA as an intermediate, it is called a DNA transposon.[22] Other classification systems refer to retrotransposons as "Class I" and DNA transposons as "Class II" transposable elements.[23]
Transposable elements are estimated to constitute 45% of the human genome.[25] Since uncontrolled propagation of TEs could wreak havoc on the genome, many regulatory mechanisms have evolved to silence their spread, including DNA methylation, histone modifications, non-coding RNAs (ncRNAs) including small interfering RNA (siRNA), chromatin remodelers, histone variants, and other epigenetic factors.[23] However, TEs play a wide variety of important biological functions. When TEs are introduced into a new host, such as from a virus, they increase genetic diversity.[23] In some cases, host organisms find new functions for the proteins which arise from expressing TEs in an evolutionary process called TE exaptation.[23] Recent research also suggests that TEs serve to maintain higher-order chromatin structure and 3D genome organization.[26] Furthermore, TEs contribute to regulating the expression of other genes by serving as distal enhancers and transcription factor binding sites.[27]
The prevalence of interspersed elements in the genome has garnered attention for more research on their origins and functions. Some specific interspersed elements have been characterized, such as the Alu repeat and LINE1.
Intrachromosomal recombination
Homologous recombination between chromosomal repeated sequences in somatic cells of Nicotiana tabacum was found to be increased by exposure to mitomycin C, a bifunctional alkylating agent that crosslinks DNA strands.[28] This increase in recombination was attributed to increased intrachromosomal recombinational repair.[28] By this process, mitomycin C damaged DNA in one sequence is repaired using intact information from the other repeated sequence.
Direct and inverted repeats
While tandem and interspersed repeats are distinguished based on their location in the genome, direct and inverted repeats are distinguished based on the ordering of the nucleotide bases. Direct repeats occur when a nucleotide sequence is repeated with the same directionality. Inverted repeats occur when a nucleotide sequence is repeated in the inverse direction. For example, a direct repeat of "CATCAT" would be another repetition of "CATCAT". In contrast, the inverted repeated would be "ATGATG". When there are no nucleotides separating the inverted repeat, such as "CATCATATGATG", the sequence is called a palindromic repeat. Inverted repeats can play structural roles in DNA and RNA by forming stem loops and cruciforms.[29]
Repeated sequences in human disease
For humans, some repeated DNA sequences are associated with diseases. Specifically, tandem repeat sequences, underlie several
Huntington's disease
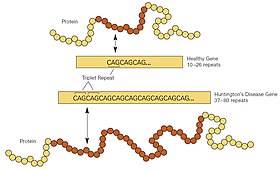
Huntington's disease is a neurodegenerative disorder which is due to the expansion of repeated trinucleotide sequence CAG in exon 1 of the huntingtin gene (HTT). This gene is responsible for encoding the protein huntingtin which plays a role in preventing apoptosis,[32] otherwise known as cell death, and repair of oxidative DNA damage.[33] In Huntington's disease the expansion of the trinucleotide sequence CAG encodes for a mutant huntingtin protein with an expanded polyglutamine domain.[34] This domain causes the protein to form aggregates in nerve cells preventing normal cellular function and resulting in neurodegeneration.

Fragile X syndrome
Fragile X syndrome is caused by the expansion of the DNA sequence CCG in the FMR1 gene on the X chromosome.[35] This gene produces the RNA-binding protein FMRP. In the case of Fragile X syndrome the repeated sequence makes the gene unstable and therefore silences the gene FMR1.[36] Because the gene resides on the X chromosome, females who have two X chromosomes are less effected than males who only have on X chromosome and one Y chromosome because the second X chromosome can compensate for the silencing of the gene on the other X chromosome.
Spinocerebellar ataxias
The disease spinocerebellar ataxias has CAG trinucleotide repeat sequences that underlie several types of spinocerebellar ataxias (SCAs-SCA1; SCA2; SCA3; SCA6; SCA7; SCA12; SCA17).[37] Similar to Huntington's disease, the polyglutamine tail created due to this trinucleotide expansion causes aggregation of proteins, preventing normal cellular function and causing neurodegeneration.[38]
Friedreich's Ataxia
Friedreich's ataxia is a type of ataxia that has an expanded repeat sequence GAA in the frataxin gene.[39] The frataxin gene is responsible for producing the frataxin protein, which is a mitochondrial protein involved in energy production and cellular respiration.[40] The expanded GAA sequence results in the silencing of the first intron resulting in loss of function in the frataxin protein. The loss of a functional FXN gene leads to issues with mitochondrial functioning as a whole and can present phenotypically in patients as difficulty walking.
Myotonic dystrophy
Myotonic dystrophy is a disorder that presents as muscle weakness and consists of two main types: DM1 and DM2.[41] Both types of myotonic dystrophy are due to expanded DNA sequences. In DM1 the DNA sequence that is expanded is CCG while in DM2 it is CCTG. These two sequences are found on different genes with the expanded sequence in DM2 being found on the ZNF9 gene and the expanded sequence in DM1 found on the DMPK gene. The two genes don't encode for proteins unlike other disorders like Huntington's disease or Fragile X syndrome. It has been shown, however, that there is a link between RNA toxicity and the repeat sequences in DM1 and DM2.
Amyotrophic lateral sclerosis and Frontotemporal dementia
Not all diseases caused by repeated DNA sequences are trinucleotide repeat diseases. The diseases
Biotechnology
Repetitive DNA is hard to
Bustos. et al. proposed one method of sequencing long stretches of repetitive DNA.[43] The method combines the use of a linear vector for stabilization and exonuclease III for deletion of continuing simple sequence repeats (SSRs) rich regions. First, SSR-rich fragments are cloned into a linear vector that can stably incorporate tandem repeats up to 30kb. Expression of repeats is prohibited by the transcriptional terminators in the vector. The second step involves the use of exonuclease III. The enzyme can delete nucleotide at the 3' end which results in the production of a unidirectional deletion of SSR fragments. Finally, this product which has deleted fragments is multiplied and analyzed with colony PCR. The sequence is then built by an ordered sequencing of a set of clones containing different deletions.
See also
References
- PMID 22144907.
- ^ PMID 31698818.
- ^ "Repeated Sequence (DNA) - an overview | ScienceDirect Topics". www.sciencedirect.com. Retrieved 2022-10-04.
- PMID 14942727.
- PMID 4874239.
- S2CID 18866824.
- PMID 5065367.
- S2CID 4370178.
- PMID 24809441.
- PMID 29525543.
- ^ "Tandem Repeat". Genome.gov. Retrieved 2022-09-30.
- PMID 31323950.
- ^ "MeSH Browser". meshb.nlm.nih.gov. Retrieved 2022-09-30.
- ^ PMID 9352183.
- S2CID 51718804.
- ^ PMID 15933211.
- ^ "Centromere". Genome.gov. Retrieved 2022-09-30.
- S2CID 15229421.
- S2CID 615040.
- PMID 25917896.
- ^ "Interspersed repetitive sequences - Latest research and news | Nature". www.nature.com. Retrieved 2022-09-30.
- ^ S2CID 32132898.
- ^ PMID 34831175.
- ^ S2CID 222199613.
- PMID 26781081.
- PMID 33990851.
- PMID 23676707.
- ^ a b Lebel EG, Masson J, Bogucki A, Paszkowski J. Stress-induced intrachromosomal recombination in plant somatic cells. Proc Natl Acad Sci U S A. 1993 Jan 15;90(2):422-6. doi: 10.1073/pnas.90.2.422. PMID: 11607349; PMCID: PMC45674
- S2CID 22204780.
- ^ PMID 25608779.
- ^ PMID 29419417.
- S2CID 10119487.
- PMID 28017939.
- PMID 22180703.
- PMID 17477822.
- S2CID 583204.
- ^ PMID 31331820.
- PMID 18568057.
- PMID 28405347.
- PMID 32826895.
- PMID 27141276.
- PMID 23160421.
- ^ PMID 27819354.
- PMID 29743957.
External links
- Function of Repetitive DNA
- DNA+Repetitious+Region at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
