Retinoblastoma protein
Ensembl | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| UniProt | |||||||||
| RefSeq (mRNA) | |||||||||
| RefSeq (protein) | |||||||||
| Location (UCSC) | Chr 13: 48.3 – 48.6 Mb | Chr 14: 73.42 – 73.56 Mb | |||||||
| PubMed search | [3] | [4] | |||||||
| View/Edit Human | View/Edit Mouse |
The retinoblastoma protein (protein name abbreviated Rb; gene name abbreviated Rb, RB or RB1) is a
pRb belongs to the
Name and genetics
In humans, the protein is encoded by the RB1 gene located on
Two forms of retinoblastoma were noticed: a bilateral, familial form and a unilateral, sporadic form. Sufferers of the former were over six times more likely to develop other types of cancer later in life, compared to individuals with sporadic retinoblastoma.
However, in the sporadic form, both alleles would need to sustain a mutation before the cell can become cancerous. This explains why sufferers of sporadic retinoblastoma are not at increased risk of cancers later in life, as both alleles are functional in all their other cells. Future cancer incidence in sporadic pRb cases is observed with polynomial kinetics, not exactly quadratic as expected because the first mutation must arise through normal mechanisms, and then can be duplicated by LOH to result in a tumour progenitor.
RB1
RB/E2F-family proteins repress transcription.[13]
Structure denotes function
pRb is a multifunctional protein with many binding and phosphorylation sites. Although its common function is seen as binding and repressing E2F targets, pRb is likely a multifunctional protein as it binds to at least 100 other proteins.[14]
pRb has three major structural components: a carboxy-terminus, a "pocket" subunit, and an amino-terminus. Within each domain, there are a variety of protein binding sites, as well as a total of 15 possible phosphorylation sites. Generally, phosphorylation causes interdomain locking, which changes pRb's conformation and prevents binding to target proteins. Different sites may be phosphorylated at different times, giving rise to many possible conformations and likely many functions/activity levels.[15]
Cell cycle suppression

pRb restricts the cell's ability to
pRb attenuates protein levels of known E2F Targets
pRb has the ability to reversibly inhibit DNA replication through transcriptional repression of DNA replication factors. pRb is able to bind to transcription factors in the E2F family and thereby inhibit their function. When pRb is chronically activated, it leads to the downregulation of the necessary DNA replication factors. Within 72–96 hours of active pRb induction in A2-4 cells, the target DNA replication factor proteins—MCMs, RPA34, DBF4, RFCp37, and RFCp140—all showed decreased levels. Along with decreased levels, there was a simultaneous and expected inhibition of DNA replication in these cells. This process, however, is reversible. Following induced knockout of pRb, cells treated with cisplatin, a DNA-damaging agent, were able to continue proliferating, without cell cycle arrest, suggesting pRb plays an important role in triggering chronic S-phase arrest in response to genotoxic stress.
One such example of E2F-regulated genes repressed by pRb are
Repression mechanisms of E2Fs
There are at least three distinct mechanisms in which pRb can repress transcription of E2F-regulated promoters. Though these mechanisms are known, it is unclear which are the most important for the control of the cell cycle.
E2Fs are a family of proteins whose binding sites are often found in the promoter regions of genes for cell proliferation or progression of the cell cycle. E2F1 to E2F5 are known to associate with proteins in the pRb-family of proteins while E2F6 and E2F7 are independent of pRb. Broadly, the E2Fs are split into activator E2Fs and repressor E2Fs though their role is more flexible than that on occasion. The activator E2Fs are E2F1, E2F2 and E2F3 while the repressor E2Fs are E2F4, E2F5 and E2F6. Activator E2Fs along with E2F4 bind exclusively to pRb. pRb is able to bind to the activation domain of the activator E2Fs which blocks their activity, repressing transcription of the genes controlled by that E2F-promoter.
Blocking of pre-initiation complex assembly
The
The presence of pRb may change the conformation of the TFIIA/IID complex into a less active version with a decreased binding affinity. pRb can also directly interfere with their association as proteins, preventing TFIIA/IID from forming an active complex.
Modification of chromatin structure
pRb acts as a recruiter that allows for the binding of proteins that alter chromatin structure onto the site E2F-regulated promoters. Access to these E2F-regulated promoters by transcriptional factors is blocked by the formation of
pRb interacts with the histone deacetylases HDAC1 and HDAC3. pRb binds to HDAC1 in its pocket domain in a region that is independent to its E2F-binding site. pRb recruitment of histone deacetylases leads to the repression of genes at E2F-regulated promoters due to nucleosome formation. Some genes activated during the G1/S transition such as cyclin E are repressed by HDAC during early to mid-G1 phase. This suggests that HDAC-assisted repression of cell cycle progression genes is crucial for the ability of pRb to arrest cells in G1. To further add to this point, the HDAC-pRb complex is shown to be disrupted by cyclin D/Cdk4 which levels increase and peak during the late G1 phase.
Senescence induced by pRb
Senescence in cells is a state in which cells are metabolically active but are no longer able to replicate. pRb is an important regulator of senescence in cells and since this prevents proliferation, senescence is an important antitumor mechanism. pRb may occupy E2F-regulated promoters during senescence. For example, pRb was detected on the cyclin A and
S-phase arrest
Cells respond to stress in the form of DNA damage, activated oncogenes, or sub-par growing conditions, and can enter a senescence-like state called "premature senescence". This allows the cell to prevent further replication during periods of damaged DNA or general unfavorable conditions. DNA damage in a cell can induce pRb activation. pRb's role in repressing the transcription of cell cycle progression genes leads to the S phase arrest that prevents replication of damaged DNA.
Activation and inactivation
When it is time for a cell to enter S phase, complexes of
Inactivation
Since the 1990s, pRb was known to be inactivated via phosphorylation. Until, the prevailing model was that Cyclin D- Cdk 4/6 progressively phosphorylated it from its unphosphorylated to its hyperphosphorylated state (14+ phosphorylations). However, it was recently shown that pRb only exists in three states: un-phosphorylated, mono-phosphorylated, and hyper-phosphorylated. Each has a unique cellular function.[25]
Before the development of 2D IEF, only hyper-phosphorylated pRb was distinguishable from all other forms, i.e. un-phosphorylated pRb resembled mono-phosphorylated pRb on immunoblots. As pRb was either in its active “hypo-phosphorylated” state or inactive “hyperphosphorylated” state. However, with 2D IEF, it is now known that pRb is un-phosphorylated in G0 cells and mono-phosphorylated in early G1 cells, prior to hyper-phosphorylation after the restriction point in late G1.[25]
pRb mono phosphorylation
When a cell enters G1, Cyclin D- Cdk4/6 phosphorylates pRb at a single phosphorylation site. No progressive phosphorylation occurs because when HFF cells were exposed to sustained cyclin D- Cdk4/6 activity (and even deregulated activity) in early G1, only mono-phosphorylated pRb was detected. Furthermore, triple knockout, p16 addition, and Cdk 4/6 inhibitor addition experiments confirmed that Cyclin D- Cdk 4/6 is the sole phosphorylator of pRb.[25]
Throughout early G1, mono-phosphorylated pRb exists as 14 different isoforms (the 15th phosphorylation site is not conserved in primates in which the experiments were performed). Together, these isoforms represent the “hypo-phosphorylated” active pRb state that was thought to exist. Each isoform has distinct preferences to associate with different exogenous expressed E2Fs.[25]
A recent report showed that mono-phosphorylation controls pRb's association with other proteins and generates functional distinct forms of pRb.[26] All different mono-phosphorylated pRb isoforms inhibit E2F transcriptional program and are able to arrest cells in G1-phase. Importantly, different mono-phosphorylated forms of pRb have distinct transcriptional outputs that are extended beyond E2F regulation.[26]
Hyper-phosphorylation
After a cell passes the restriction point, Cyclin E - Cdk 2 hyper-phosphorylates all mono-phosphorylated isoforms. While the exact mechanism is unknown, one hypothesis is that binding to the C-terminus tail opens the pocket subunit, allowing access to all phosphorylation sites. This process is hysteretic and irreversible, and it is thought accumulation of mono-phosphorylated pRb induces the process. The bistable, switch like behavior of pRb can thus be modeled as a bifurcation point:[25]

Control of pRb function by phosphorylation
Presence of un-phosphorylated pRb drives cell cycle exit and maintains senescence. At the end of mitosis, PP1 dephosphorylates hyper-phosphorylated pRb directly to its un-phosphorylated state. Furthermore, when cycling C2C12 myoblast cells differentiated (by being placed into a differentiation medium), only un-phosphorylated pRb was present. Additionally, these cells had a markedly decreased growth rate and concentration of DNA replication factors (suggesting G0 arrest).[25]
This function of un-phosphorylated pRb gives rise to a hypothesis for the lack of cell cycle control in cancerous cells: Deregulation of Cyclin D - Cdk 4/6 phosphorylates un-phosphorylated pRb in senescent cells to mono-phosphorylated pRb, causing them to enter G1. The mechanism of the switch for Cyclin E activation is not known, but one hypothesis is that it is a metabolic sensor. Mono-phosphorylated pRb induces an increase in metabolism, so the accumulation of mono-phosphorylated pRb in previously G0 cells then causes hyper-phosphorylation and mitotic entry. Since any un-phosphorylated pRb is immediately phosphorylated, the cell is then unable to exit the cell cycle, resulting in continuous division.[25]
DNA damage to G0 cells activates Cyclin D - Cdk 4/6, resulting in mono-phosphorylation of un-phosphorylated pRb. Then, active mono-phosphorylated pRb causes repression of E2F-targeted genes specifically. Therefore, mono-phosphorylated pRb is thought to play an active role in DNA damage response, so that E2F gene repression occurs until the damage is fixed and the cell can pass the restriction point. As a side note, the discovery that damages causes Cyclin D - Cdk 4/6 activation even in G0 cells should be kept in mind when patients are treated with both DNA damaging chemotherapy and Cyclin D - Cdk 4/6 inhibitors.[25]
Activation
During the M-to-G1 transition, pRb is then progressively dephosphorylated by
pRb family proteins are components of the DREAM complex composed of DP, E2F4/5, RB-like (p130/p107) And MuvB (Lin9:Lin37:Lin52:RbAbP4:Lin54). The DREAM complex is assembled in Go/G1 and maintains quiescence by assembling at the promoters of > 800 cell-cycle genes and mediating transcriptional repression. Assembly of DREAM requires DYRK1A (Ser/Thr kinase) dependant phosphorylation of the MuvB core component, Lin52 at Serine28. This mechanism is crucial for recruitment of p130/p107 to the MuvB core and thus DREAM assembly.
Consequences of pRb loss
Consequences of loss of pRb function is dependent on cell type and cell cycle status, as pRb's tumor suppressive role changes depending on the state and current identity of the cell.
In G0 quiescent stem cells, pRb is proposed to maintain G0 arrest although the mechanism remains largely unknown. Loss of pRb leads to exit from quiescence and an increase in the number of cells without loss of cell renewal capacity. In cycling progenitor cells, pRb plays a role at the G1, S, and G2 checkpoints and promotes differentiation. In differentiated cells, which make up the majority of cells in the body and are assumed to be in irreversible G0, pRb maintains both arrest and differentiation.[28]
Loss of pRb therefore exhibits multiple different responses within different cells that ultimately all could result in cancer phenotypes. For cancer initiation, loss of pRb may induce cell cycle re-entry in both quiescent and post-mitotic differentiated cells through dedifferentiation. In cancer progression, loss of pRb decreases the differentiating potential of cycling cells, increases chromosomal instability, prevents induction of cellular senescence, promotes angiogenesis, and increases metastatic potential.[28]
Although most cancers rely on glycolysis for energy production (Warburg effect),[29] cancers due to pRb loss tend to upregulate oxidative phosphorylation.[30] The increased oxidative phosphorylation can increase stemness, metastasis, and (when enough oxygen is available) cellular energy for anabolism.[30]
In vivo, it is still not entirely clear how and which cell types cancer initiation occurs with solely loss of pRb, but it is clear that the pRb pathway is altered in large number of human cancers.[110] In mice, loss of pRb is sufficient to initiate tumors of the pituitary and thyroid glands, and mechanisms of initiation for these hyperplasia are currently being investigated.[31]
Non-canonical roles
The classic view of pRb's role as a tumor suppressor and cell cycle regulator developed through research investigating mechanisms of interactions with E2F family member proteins. Yet, more data generated from biochemical experiments and clinical trials reveal other functions of pRb within the cell unrelated (or indirectly related) to tumor suppression.[32]
Functional hyperphosphorylated pRb
In proliferating cells, certain pRb conformations (when RxL motif if bound by protein phosphatase 1 or when it is acetylated or methylated) are resistant to CDK phosphorylation and retain other function throughout cell cycle progression, suggesting not all pRb in the cell are devoted to guarding the G1/S transition.[32]
Studies have also demonstrated that hyperphosphorylated pRb can specifically bind E2F1 and form stable complexes throughout the cell cycle to carry out unique unexplored functions, a surprising contrast from the classical view of pRb releasing E2F factors upon phosphorylation.[32]
In summary, many new findings about pRb's resistance to CDK phosphorylation are emerging in pRb research and shedding light on novel roles of pRb beyond cell cycle regulation.
Genome stability
pRb is able to be localize to sites of DNA breaks during the repair process and assist in non-homologous end joining and homologous recombination through complexing with E2F1. Once at the breaks, pRb is able to recruit regulators of chromatin structure such as the DNA helicase transcription activator BRG1. pRb has been shown to also be able to recruit protein complexes such as condensin and cohesin to assist in the structural maintenance of chromatin.[32]
Such findings suggest that in addition to its tumor suppressive role with E2F, pRb is also distributed throughout the genome to aid in important processes of genome maintenance such as DNA break-repair, DNA replication, chromosome condensation, and heterochromatin formation.[32]
Regulation of metabolism
pRb has also been implicated in regulating metabolism through interactions with components of cellular metabolic pathways. RB1 mutations can cause alterations in metabolism, including reduced mitochondrial respiration, reduced activity in the electron transport chain, and changes in flux of glucose and/or glutamine. Particular forms of pRb have been found to localize to the outer mitochondrial membrane and directly interacts with Bax to promote apoptosis.[33]
As a drug target
pRb Reactivation
While the frequency of alterations of the RB gene is substantial for many human cancer types including as lung, esophageal, and liver, alterations in up-steam regulatory components of pRb such as CDK4 and CDK6 have been the main targets for potential therapeutics to treat cancers with dysregulation in the RB pathway.[34] This focus has resulted in the recent development and FDA clinical approval of three small molecule CDK4/6 inhibitors (Palbociclib (IBRANCE, Pfizer Inc. 2015), Ribociclib (KISQUALI, Novartis. 2017), & Abemaciclib (VERZENIO, Eli Lilly. 2017)) for the treatment of specific breast cancer subtypes. However, recent clinical studies finding limited efficacy, high toxicity, and acquired resistance[35][36] of these inhibitors suggests the need to further elucidate mechanisms that influence CDK4/6 activity as well as explore other potential targets downstream in the pRb pathway to reactivate pRb's tumor suppressive functions. Treatment of cancers by CDK4/6 inhibitors depends on the presence of pRb within the cell for therapeutic effect, limiting their usage only to cancers where RB is not mutated and pRb protein levels are not significantly depleted.[34]
Direct pRb reactivation in humans has not been achieved. However, in murine models, novel genetic methods have allowed for in vivo pRb reactivation experiments. pRb loss induced in mice with oncogenic KRAS-driven tumors of lung adenocarcinoma negates the requirement of MAPK signal amplification for progression to carcinoma and promotes loss of lineage commitment as well as accelerate the acquisition of metastatic competency. Reactivation of pRb in these mice rescues the tumors towards a less metastatic state, but does not completely stop tumor growth due to a proposed rewiring of MAPK pathway signaling, which suppresses pRb through a CDK-dependent mechanism.[37]
Pro-apoptotic effects of pRb loss
Besides trying to re-activate the tumor suppressive function of pRb, one other distinct approach to treat dysregulated pRb pathway cancers is to take advantage of certain cellular consequences induced by pRb loss. It has been shown that E2F stimulates expression of pro-apoptotic genes in addition to G1/S transition genes, however, cancer cells have developed defensive signaling pathways that protect themselves from death by deregulated E2F activity. Development of inhibitors of these protective pathways could thus be a synthetically lethal method to kill cancer cells with overactive E2F.[34]
In addition, it has been shown that the pro-apoptotic activity of p53 is restrained by the pRb pathway, such that pRb deficient tumor cells become sensitive to p53 mediated cell death. This opens the door to research of compounds that could activate p53 activity in these cancer cells and induce apoptosis and reduce cell proliferation.[34]
Regeneration
While the loss of a tumor suppressor such as pRb leading to uncontrolled cell proliferation is detrimental in the context of cancer, it may be beneficial to deplete or inhibit suppressive functions of pRb in the context of cellular regeneration.[38] Harvesting the proliferative abilities of cells induced to a controlled “cancer like” state could aid in repairing damaged tissues and delay aging phenotypes. This idea remains to be thoroughly explored as a potential cellular injury and anti-aging treatment.
Cochlea
The retinoblastoma protein is involved in the growth and development of mammalian
Neurons
Disrupting pRb expression in vitro, either by gene deletion or knockdown of pRb short interfering RNA, causes dendrites to branch out farther. In addition, Schwann cells, which provide essential support for the survival of neurons, travel with the neurites, extending farther than normal. The inhibition of pRb supports the continued growth of nerve cells.[42]
Interactions
pRb is known to interact with more than 300 proteins, some of which are listed below:
- Androgen receptor[45][46]
- ARID4A[49]
- Aryl hydrocarbon receptor[50]
- BRCA1[51][52][53]
- BRF1[54][55]
- C-jun[56]
- C-Raf[57][58]
- CDK9[59]
- CUTL1[60]
- Cyclin A1[61]
- Cyclin D1[62][63]
- Cyclin T2[59]
- DNMT1[64]
- E2F1[65][66][67][68][69][17][70]
- E2F2,[71]
- E4F1[68]
- EID1[72][73]
- ENC1[74]
- FRK[75]
- HBP1[76]
- HDAC1[49][77][78][79][80][81][82]
- HDAC3[49][83]
- Histone deacetylase 2[49]
- Insulin[84]
- Large tumor antigen[87][88]
- LIN9[89]
- MCM7[90]
- MORF4L1[66][91]
- MRFAP1,[66][91]
- MyoD[92][93]
- NCOA6[94]
- PA2G4[95]
- Peroxisome proliferator-activated receptor gamma[83]
- PIK3R3[96]
- Plasminogen activator inhibitor-2[97]
- Polymerase (DNA directed), alpha 1[98]
- PRDM2[99]
- PRKRA[100]
- Prohibitin[58][101]
- Promyelocytic leukemia protein[102]
- RBBP4[65][103]
- RBBP7[53][103]
- RBBP8[77][104]
- RBBP9[105]
- SNAPC1[106]
- SKP2[107][108]
- SNAPC3[106]
- SNW1[109]
- SUV39H1[110][111]
- TAF1[62][112][113][114]
- THOC1[115]
- TRAP1[116]
- TRIP11[117]
- UBTF[118]
- USP4.[119]
Detection
Several methods for detecting the RB1 gene mutations have been developed[120] including a method that can detect large deletions that correlate with advanced stage retinoblastoma.[121]
See also
- p53 - involved in the DNA repair support function of pRb
- Transcription coregulator
- Retinoblastoma
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000139687 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000022105 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- PMID 6320372.
- PMID 7838522.
- ^ PMID 16081278.
- ^ PMID 12445661.
- ^ Gallego J (May 2016). "Multicellular Life Was Caused By The Same Gene That Suppresses Cancer". Kansas State University.
- PMID 15800318.
- PMID 5279523.
- ^ "OrthoMaM phylogenetic marker: RB1 coding sequence". Archived from the original on 2015-09-24. Retrieved 2009-12-02.
- PMID 15126619.
- PMID 11447760.
- PMID 23594950.
- S2CID 12990398.
- ^ PMID 7739537.
- ^ PMID 9284048.
- ^ PMID 14611951.
- PMID 15840447.
- ^ .
- PMID 16210110.
- PMID 16236452.
- PMID 12642231.
- ^ PMID 24876129.
- ^ PMID 30711375.
- PMID 16466572.
- ^ PMID 18650841.
- PMID 20181022.
- ^ PMID 29120753.
- PMID 22751497.
- ^ PMID 29692417.
- PMID 27401552.
- ^ PMID 20145169.
- PMID 30675102.
- PMID 27217383.
- PMID 31043741.
- PMID 23715544.
- PMID 16648263.
- PMID 18178626.
- PMID 23211596.
- PMID 24752312.
- PMID 9071815.
- PMID 8242749.
- PMID 9813067.
- PMID 9675141.
- PMID 12450794.
- S2CID 43175069.
- ^ PMID 10490602.
- PMID 9712901.
- PMID 10518542.
- PMID 11521194.
- ^ PMID 10220405.
- PMID 11997511.
- PMID 10330166.
- PMID 10026157.
- PMID 9819434.
- ^ PMID 10523633.
- ^ PMID 12037672.
- S2CID 2287673.
- PMID 10022926.
- ^ PMID 11126356.
- S2CID 24708871.
- S2CID 10983932.
- ^ PMID 11470869.
- ^ PMID 12397079.
- PMID 8896460.
- ^ PMID 10869426.
- PMID 8230483.
- PMID 9422723.
- PMID 12502741.
- PMID 11073989.
- PMID 11073990.
- PMID 9566959.
- PMID 7664264.
- PMID 9178770.
- ^ PMID 10779361.
- S2CID 20428600.
- PMID 11684023.
- PMID 12466959.
- S2CID 18857544.
- PMID 9724731.
- ^ PMID 12479814.
- PMID 7818556.
- PMID 11358960.
- PMID 7935440.
- PMID 22994493.
- PMID 22710026.
- PMID 15538385.
- PMID 9566894.
- ^ PMID 11500496.
- PMID 11285237.
- S2CID 21581966.
- PMID 14645241.
- S2CID 42721280.
- PMID 12588990.
- PMID 12944478.
- PMID 9395244.
- PMID 7538672.
- PMID 9010216.
- PMID 10376528.
- PMID 9448006.
- ^ PMID 7503932.
- PMID 9721205.
- S2CID 11374970.
- ^ PMID 11094070.
- PMID 15469821.
- PMID 19966802.
- PMID 12466551.
- S2CID 4378296.
- PMID 11533237.
- PMID 7724524.
- PMID 9858607.
- PMID 9242374.
- PMID 7525595.
- PMID 8756626.
- PMID 9256431.
- PMID 11042686.
- PMID 11571651.
- S2CID 10723496.
- PMID 23960888.
Further reading
- Momand J, Wu HH, Dasgupta G (January 2000). "MDM2--master regulator of the p53 tumor suppressor protein". Gene. 242 (1–2): 15–29. PMID 10721693.
- Zheng L, Lee WH (2003). "Retinoblastoma tumor suppressor and genome stability". Advances in Cancer Research Volume 85. Vol. 85. pp. 13–50. PMID 12374284.
- Classon M, Harlow E (December 2002). "The retinoblastoma tumour suppressor in development and cancer". Nature Reviews. Cancer. 2 (12): 910–7. S2CID 22937378.
- Lai H, Ma F, Lai S (January 2003). "Identification of the novel role of pRB in eye cancer". Journal of Cellular Biochemistry. 88 (1): 121–7. S2CID 34538683.
- Simin K, Wu H, Lu L, Pinkel D, Albertson D, Cardiff RD, Van Dyke T (February 2004). "pRb inactivation in mammary cells reveals common mechanisms for tumor initiation and progression in divergent epithelia". PLOS Biology. 2 (2): E22. PMID 14966529.
- Lohmann DR, Gallie BL (August 2004). "Retinoblastoma: revisiting the model prototype of inherited cancer". American Journal of Medical Genetics. Part C, Seminars in Medical Genetics. 129C (1): 23–8. S2CID 41922148.
- Clemo NK, Arhel NJ, Barnes JD, Baker J, Moorghen M, Packham GK, et al. (August 2005). "The role of the retinoblastoma protein (Rb) in the nuclear localization of BAG-1: implications for colorectal tumour cell survival". Biochemical Society Transactions. 33 (Pt 4): 676–8. PMID 16042572.
- Rodríguez-Cruz M, del Prado M, Salcedo M (2006). "[Genomic retinoblastoma perspectives: implications of tumor supressor [sic] gene RB1]". Revista de Investigacion Clinica. 57 (4): 572–81. PMID 16315642.
- Knudsen ES, Knudsen KE (July 2006). "Retinoblastoma tumor suppressor: where cancer meets the cell cycle". Experimental Biology and Medicine. 231 (7): 1271–81. S2CID 29078799.
External links
- RB1+protein,+human at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- Retinoblastoma+genes at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- GeneReviews/NIH/NCBI/UW entry on Retinoblastoma
- Retinoblastoma Genetics
- Drosophila Retinoblastoma-family protein - The Interactive Fly
- Drosophila Retinoblastoma-family protein 2 - The Interactive Fly
- Evolutionary Homologs Retinoblastoma-family proteins - The Interactive Fly
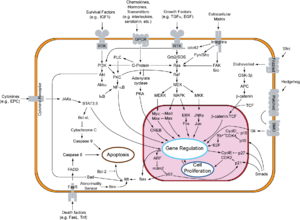
- There is a diagram of the pRb-E2F interactions here[permanent dead link].
This article incorporates text from the United States National Library of Medicine, which is in the public domain.





