Rhodopsin
Ensembl | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| UniProt | |||||||||
| RefSeq (mRNA) | |||||||||
| RefSeq (protein) | |||||||||
| Location (UCSC) | Chr 3: 129.53 – 129.54 Mb | Chr 6: 115.91 – 115.92 Mb | |||||||
| PubMed search | [3] | [4] | |||||||
| View/Edit Human | View/Edit Mouse |
Rhodopsin, also known as visual purple, is a
Names
Rhodopsin was discovered by
When
General
Rhodopsin is a protein found in the outer segment discs of rod cells. It mediates scotopic vision, which is monochromatic vision in dim light.[7][19] Rhodopsin most strongly absorbs green-blue light (~500 nm)[20][21] and appears therefore reddish-purple, hence the archaic term "visual purple".
Several closely related opsins differ only in a few amino acids and in the wavelengths of light that they absorb most strongly. Humans have, including rhodopsin, nine opsins,[15] as well as cryptochrome (light-sensitive, but not an opsin).[22]
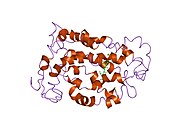
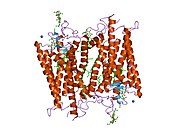
Structure

Rhodopsin, like other opsins, is a
The retinal binding lysine is conserved in almost all opsins, only a few opsins having lost it during
The rhodopsin of cattle has 348 amino acids, the retinal binding lysine being Lys296. It was the first opsin whose amino acid sequence[50] and 3D-structure were determined.[32] Its structure has been studied in detail by x-ray crystallography on rhodopsin crystals.[51] Several models (e.g., the bicycle-pedal mechanism, hula-twist mechanism) attempt to explain how the retinal group can change its conformation without clashing with the enveloping rhodopsin protein pocket.[52][53][54] Recent data support that rhodopsin is a functional monomer, instead of a dimer, which was the paradigm of G-protein-coupled receptors for many years.[55]
Within its native membrane, rhodopsin is found at a high density facilitating its ability to capture photons. Due to its dense packing within the membrane, there is a higher chance of rhodopsin capturing proteins. However, the high density also provides a disadvantage when it comes to G protein signaling because the diffusion becomes more difficult in a crowded membrane that is packed with the receptor, rhodopsin.[56]
Phototransduction

Rhodopsin is an essential G-protein coupled receptor in
Activation
In rhodopsin, the aldehyde group of retinal is covalently linked to the amino group of a lysine residue on the protein in a protonated
Phototransduction cascade
The product of light activation, Metarhodopsin II, initiates the
Deactivation
Meta II (metarhodopsin II) is deactivated rapidly after activating transducin by rhodopsin kinase and arrestin.[61] Rhodopsin pigment must be regenerated for further phototransduction to occur. This means replacing all-trans-retinal with 11-cis-retinal and the decay of Meta II is crucial in this process. During the decay of Meta II, the Schiff base link that normally holds all-trans-retinal and the apoprotein opsin (aporhodopsin) is hydrolyzed and becomes Meta III. In the rod outer segment, Meta III decays into separate all-trans-retinal and opsin.[61] A second product of Meta II decay is an all-trans-retinal opsin complex in which the all-trans-retinal has been translocated to second binding sites. Whether the Meta II decay runs into Meta III or the all-trans-retinal opsin complex seems to depend on the pH of the reaction. Higher pH tends to drive the decay reaction towards Meta III.[61]
Diseases of the retina
Mutations in the rhodopsin gene contribute majorly to various diseases of the retina such as
See also
- halobacteria as a light-driven proton pump.
Explanatory notes
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000163914 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000030324 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "RHO rhodopsin [Homo sapiens (human)]". NCBI. Retrieved 16 November 2017.
- ISBN 978-1-55938-659-3.
- ^ ISBN 978-1-55938-659-3.
- ISBN 978-0-12-385158-1.
- ISBN 978-1-4832-6227-7. Retrieved 23 September 2015.
- ^ Boll F (1877). "Zur Anatomie und Physiologie der Retina" [On the anatomy and physiology of the retina]. Archiv für Anatomie und Physiologie, Physiologische Abtheilung (in German): 4–35.
- ^ "Rhodopsin: History and Etymology for rhodopsin". Merriam-Webster on-line dictionary.
- ^ See:
- Merriam-Webster Online Dictionary: Rhodopsin: History and Etymology for rhodopsin
- Ewald A, Kühne W (1878). "Untersuchungen über den Sehpurpur" [Investigations into rhodopsin]. Untersuchungen aus dem Physiologischen Institute der Universität Heidelberg (in German). 1: 139–218. From p. 181: "Was den Sehpurpur im Dunkel ändert, pflegt es z. Th. [= zum Theil] in derselben Weise zu thun, wie das Licht, d.h. erst eine gelbe Materie, dann farblose Substanz hervorzubringen. Der Kürze wegen und um dem Auslande unsere Bezeichnungen zugänglich zu machen, kann man sagen, Rhodopsin werde erst in Xanthopsin, dieses in Leukopsin zersetzt." (That which alters visual purple in the dark usually acts to some extent in the same way as light, that is, first producing a yellow material, then a colorless substance. For the sake of brevity, and in order to make our designations more accessible to foreigners, we can say that rhodopsin is first degraded into xanthopsin [- visual yellow], and [then] this is degraded into leucopsin [- visual white].)
- S2CID 19145558.
- PMID 14908734.
- ^ PMID 15774036.
- ^ PMID 35954284..
 Material was copied and adapted from this source, which is available under a Creative Commons Attribution 4.0 International License
Material was copied and adapted from this source, which is available under a Creative Commons Attribution 4.0 International License - ^ S2CID 253041556.
- PMID 14367777.
- ^ Rogers K. "Rhodopsin". Encyclopædia Britannica. Britannica.com. Retrieved 30 January 2016.
- S2CID 45459123.
- PMID 7359434.
- PMID 21694704.
- S2CID 38970721.
- PMID 8170923.
- S2CID 4324074.
- S2CID 4352920.
- S2CID 4022911.
- PMID 16589696.
- PMID 13367054.
- .
- PMID 16589909.
- ^ PMID 10926528.
- ^ S2CID 1657759.
- S2CID 20407577.
- S2CID 4152360.
- PMID 14351151.
- PMID 16590155.
- S2CID 45830716.
- ^ S2CID 4302421.
- PMID 4877437.
- ^ doi:10.1002/wmts.6.
- S2CID 201420079.
- PMID 32243853.
- S2CID 23060331.
- S2CID 13172583.
- PMID 9391065.
- ^ PMID 24931191.
- PMID 9414230.
- PMID 23904486.
- S2CID 85819100.
- PMID 28289214.
- PMID 16586416.
- PMID 16729349.
- PMID 17691730.
- PMID 15996094.
- PMID 31286171.
- ^ The Nobel Foundation. "The Nobel Prize in Physiology or Medicine 1967". Nobelprize.org. Nobel Media AB 2014. Retrieved 12 December 2015.
- S2CID 4263392.
- PMID 14080814.
- ISBN 978-1-55938-659-3.
- ^ PMID 12427735.
- PMID 12082151.
- ^ PMID 15823756.
Further reading
- Humphries P, Kenna P, Farrar GJ (May 1992). "On the molecular genetics of retinitis pigmentosa". Science. 256 (5058): 804–808. PMID 1589761.
- Edwards SC (July 1995). "Involvement of cGMP and calcium in the photoresponse in vertebrate photoreceptor cells". The Journal of the Florida Medical Association. 82 (7): 485–488. PMID 7673885.
- al-Maghtheh M, Gregory C, Inglehearn C, Hardcastle A, Bhattacharya S (1993). "Rhodopsin mutations in autosomal dominant retinitis pigmentosa". Human Mutation. 2 (4): 249–255. S2CID 28459589.
- Garriga P, Manyosa J (September 2002). "The eye photoreceptor protein rhodopsin. Structural implications for retinal disease". FEBS Letters. 528 (1–3): 17–22. S2CID 41860711.
- Inglehearn CF, Keen TJ, Bashir R, Jay M, Fitzke F, Bird AC, et al. (April 1992). "A completed screen for mutations of the rhodopsin gene in a panel of patients with autosomal dominant retinitis pigmentosa". Human Molecular Genetics. 1 (1): 41–45. PMID 1301135.
- Farrar GJ, Findlay JB, Kumar-Singh R, Kenna P, Humphries MM, Sharpe E, et al. (December 1992). "Autosomal dominant retinitis pigmentosa: a novel mutation in the rhodopsin gene in the original 3q linked family". Human Molecular Genetics. 1 (9): 769–771. PMID 1302614.
- Robinson PR, Cohen GB, Zhukovsky EA, Oprian DD (October 1992). "Constitutively active mutants of rhodopsin". Neuron. 9 (4): 719–725. S2CID 13172583.
- Fujiki K, Hotta Y, Hayakawa M, Sakuma H, Shiono T, Noro M, et al. (June 1992). "Point mutations of rhodopsin gene found in Japanese families with autosomal dominant retinitis pigmentosa (ADRP)". The Japanese Journal of Human Genetics. 37 (2): 125–132. PMID 1391967.
- Olsson JE, Gordon JW, Pawlyk BS, Roof D, Hayes A, Molday RS, et al. (November 1992). "Transgenic mice with a rhodopsin mutation (Pro23His): a mouse model of autosomal dominant retinitis pigmentosa". Neuron. 9 (5): 815–830. S2CID 37524461.
- Andréasson S, Ehinger B, Abrahamson M, Fex G (September 1992). "A six-generation family with autosomal dominant retinitis pigmentosa and a rhodopsin gene mutation (arginine-135-leucine)". Ophthalmic Paediatrics and Genetics. 13 (3): 145–153. PMID 1484692.
- Inglehearn CF, Lester DH, Bashir R, Atif U, Keen TJ, Sertedaki A, et al. (March 1992). "Recombination between rhodopsin and locus D3S47 (C17) in rhodopsin retinitis pigmentosa families". American Journal of Human Genetics. 50 (3): 590–597. PMID 1539595.
- Fishman GA, Stone EM, Gilbert LD, Sheffield VC (May 1992). "Ocular findings associated with a rhodopsin gene codon 106 mutation. Glycine-to-arginine change in autosomal dominant retinitis pigmentosa". Archives of Ophthalmology. 110 (5): 646–653. PMID 1580841.
- Keen TJ, Inglehearn CF, Lester DH, Bashir R, Jay M, Bird AC, et al. (September 1991). "Autosomal dominant retinitis pigmentosa: four new mutations in rhodopsin, one of them in the retinal attachment site". Genomics. 11 (1): 199–205. PMID 1765377.
- Dryja TP, Hahn LB, Cowley GS, McGee TL, Berson EL (October 1991). "Mutation spectrum of the rhodopsin gene among patients with autosomal dominant retinitis pigmentosa". Proceedings of the National Academy of Sciences of the United States of America. 88 (20): 9370–9374. PMID 1833777.
- Gal A, Artlich A, Ludwig M, Niemeyer G, Olek K, Schwinger E, et al. (October 1991). "Pro-347-Arg mutation of the rhodopsin gene in autosomal dominant retinitis pigmentosa". Genomics. 11 (2): 468–470. PMID 1840561.
- Sung CH, Davenport CM, Hennessey JC, Maumenee IH, Jacobson SG, Heckenlively JR, et al. (August 1991). "Rhodopsin mutations in autosomal dominant retinitis pigmentosa". Proceedings of the National Academy of Sciences of the United States of America. 88 (15): 6481–6485. PMID 1862076.
- Jacobson SG, Kemp CM, Sung CH, Nathans J (September 1991). "Retinal function and rhodopsin levels in autosomal dominant retinitis pigmentosa with rhodopsin mutations". American Journal of Ophthalmology. 112 (3): 256–271. PMID 1882937.
- Sheffield VC, Fishman GA, Beck JS, Kimura AE, Stone EM (October 1991). "Identification of novel rhodopsin mutations associated with retinitis pigmentosa by GC-clamped denaturing gradient gel electrophoresis". American Journal of Human Genetics. 49 (4): 699–706. PMID 1897520.
External links
- Rhodopsin at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- Kolb H, Fernandez E, Nelson R, Jones BW (1 March 2010). "Webvision Home Page: The organization of the retina and visual system". University of Utah.
- The Rhodopsin Protein
- Photoisomerization of rhodopsin, animation.
- Rhodopsin and the eye, summary with pictures.