Ribosome
Animal cell diagram | |
|---|---|
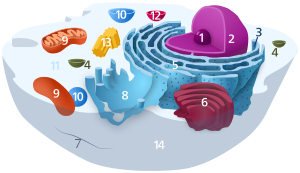 Components of a typical animal cell:
|
Ribosomes (
Overview
The sequence of
A ribosome is made from
- (30S) has mainly a decoding function and is also bound to the mRNA
- (50S) has mainly a catalytic function and is also bound to the aminoacylated tRNAs.
The synthesis of proteins from their building blocks takes place in four phases: initiation, elongation, termination, and recycling. The start codon in all mRNA molecules has the sequence AUG. The stop codon is one of UAA, UAG, or UGA; since there are no tRNA molecules that recognize these codons, the ribosome recognizes that translation is complete.[4] When a ribosome finishes reading an mRNA molecule, the two subunits separate and are usually broken up but can be reused. Ribosomes are ribozymes because the catalytic peptidyl transferase activity that links amino acids together is performed by the ribosomal RNA.[5]
Ribosomes are often associated with the intracellular membranes that make up the rough endoplasmic reticulum.
Ribosomes from bacteria, archaea, and eukaryotes in the three-domain system resemble each other to a remarkable degree, evidence of a common origin. They differ in their size, sequence, structure, and the ratio of protein to RNA. The differences in structure allow some antibiotics to kill bacteria by inhibiting their ribosomes while leaving human ribosomes unaffected. In all species, more than one ribosome may move along a single mRNA chain at one time (as a polysome), each "reading" a specific sequence and producing a corresponding protein molecule.
The mitochondrial ribosomes of eukaryotic cells functionally resemble many features of those in bacteria, reflecting the likely evolutionary origin of mitochondria.[6][7]
Discovery
Ribosomes were first observed in the mid-1950s by
During the course of the symposium a semantic difficulty became apparent. To some of the participants, "microsomes" mean the ribonucleoprotein particles of the microsome fraction contaminated by other protein and lipid material; to others, the microsomes consist of protein and lipid contaminated by particles. The phrase "microsomal particles" does not seem adequate, and "ribonucleoprotein particles of the microsome fraction" is much too awkward. During the meeting, the word "ribosome" was suggested, which has a very satisfactory name and a pleasant sound. The present confusion would be eliminated if "ribosome" were adopted to designate ribonucleoprotein particles in sizes ranging from 35 to 100S.
— Albert Claude, Microsomal Particles and Protein Synthesis[10]
Structure


The ribosome is a complex cellular machine. It is largely made up of specialized RNA known as ribosomal RNA (rRNA) as well as dozens of distinct proteins (the exact number varies slightly between species). The ribosomal proteins and rRNAs are arranged into two distinct ribosomal pieces of different sizes, known generally as the large and small subunits of the ribosome. Ribosomes consist of two subunits that fit together and work as one to translate the mRNA into a polypeptide chain during protein synthesis. Because they are formed from two subunits of non-equal size, they are slightly longer on the axis than in diameter.
Prokaryotic ribosomes
Prokaryotic ribosomes are around 20
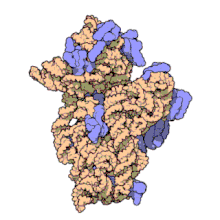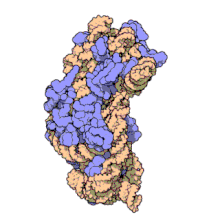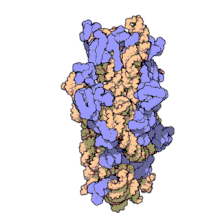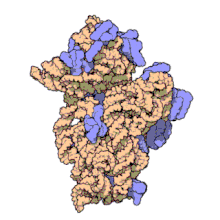
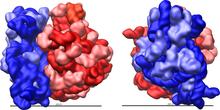
The ribosomal subunits of prokaryotes and eukaryotes are quite similar.[17]
The unit of measurement used to describe the ribosomal subunits and the rRNA fragments is the Svedberg unit, a measure of the rate of sedimentation in centrifugation rather than size. This accounts for why fragment names do not add up: for example, bacterial 70S ribosomes are made of 50S and 30S subunits.
Prokaryotes have 70
Ribosome of E. coli (a bacterium)[18]: 962 ribosome subunit rRNAs r-proteins 70S 50S 23S (2904 nt) 31 5S (120 nt) 30S 16S (1542 nt) 21
Affinity label for the tRNA binding sites on the E. coli ribosome allowed the identification of A and P site proteins most likely associated with the peptidyltransferase activity;[5] labelled proteins are L27, L14, L15, L16, L2; at least L27 is located at the donor site, as shown by E. Collatz and A.P. Czernilofsky.[19][20] Additional research has demonstrated that the S1 and S21 proteins, in association with the 3′-end of 16S ribosomal RNA, are involved in the initiation of translation.[21]
Archaeal ribosomes
Archaeal ribosomes share the same general dimensions of bacteria ones, being a 70S ribosome made up from a 50S large subunit, a 30S small subunit, and containing three rRNA chains. However, on the sequence level, they are much closer to eukaryotic ones than to bacterial ones. Every extra ribosomal protein archaea have compared to bacteria has a eukaryotic counterpart, while no such relation applies between archaea and bacteria.[22][23][24]
Eukaryotic ribosomes
Eukaryotes have 80S ribosomes located in their cytosol, each consisting of a small (40S) and large (60S) subunit. Their 40S subunit has an 18S RNA (1900 nucleotides) and 33 proteins.[25][26] The large subunit is composed of a 5S RNA (120 nucleotides), 28S RNA (4700 nucleotides), a 5.8S RNA (160 nucleotides) subunits and 49 proteins.[17][25][27]
eukaryotic cytosolic ribosomes (R. norvegicus)[18]: 65 ribosome subunit rRNAs r-proteins 80S 60S 28S (4718 nt) 49 5.8S (160 nt) 5S (120 nt) 40S 18S (1874 nt) 33
During 1977, Czernilofsky published research that used affinity labeling to identify tRNA-binding sites on rat liver ribosomes. Several proteins, including L32/33, L36, L21, L23, L28/29 and L13 were implicated as being at or near the peptidyl transferase center.[28]
Plastoribosomes and mitoribosomes
In eukaryotes, ribosomes are present in
The cryptomonad and chlorarachniophyte algae may contain a nucleomorph that resembles a vestigial eukaryotic nucleus.[32] Eukaryotic 80S ribosomes may be present in the compartment containing the nucleomorph.[33]
Making use of the differences
The differences between the bacterial and eukaryotic ribosomes are exploited by
Common properties
The various ribosomes share a core structure, which is quite similar despite the large differences in size. Much of the RNA is highly organized into various
High-resolution structure

The general molecular structure of the ribosome has been known since the early 1970s. In the early 2000s, the structure has been achieved at high resolutions, of the order of a few
The first papers giving the structure of the ribosome at atomic resolution were published almost simultaneously in late 2000. The 50S (large prokaryotic) subunit was determined from the
Two papers were published in November 2005 with structures of the
The first atomic structures of the ribosome complexed with
In 2011, the first complete atomic structure of the eukaryotic 80S ribosome from the yeast
Function
Ribosomes are minute particles consisting of RNA and associated proteins that function to synthesize proteins. Proteins are needed for many cellular functions, such as repairing damage or directing chemical processes. Ribosomes can be found floating within the cytoplasm or attached to the endoplasmic reticulum. Their main function is to convert genetic code into an amino acid sequence and to build protein polymers from amino acid monomers.
Ribosomes act as catalysts in two extremely important biological processes called peptidyl transfer and peptidyl hydrolysis.[5][47] The "PT center is responsible for producing protein bonds during protein elongation".[47]
In summary, ribosomes have two main functions: Decoding the message, and the formation of peptide bonds. These two functions reside in the ribosomal subunits. Each subunit is made of one or more rRNAs and many r-proteins. The small subunit (30S in bacteria and archaea, 40S in eukaryotes) has the decoding function, whereas the large subunit (50S in bacteria and archaea, 60S in eukaryotes) catalyzes the formation of peptide bonds, referred to as the peptidyl-transferase activity. The bacterial (and archaeal) small subunit contains the 16S rRNA and 21 r-proteins (Escherichia coli), whereas the eukaryotic small subunit contains the 18S rRNA and 32 r-proteins (Saccharomyces cerevisiae, although the numbers vary between species). The bacterial large subunit contains the 5S and 23S rRNAs and 34 r-proteins (E. coli), with the eukaryotic large subunit containing the 5S, 5.8S, and 25S/28S rRNAs and 46 r-proteins (S. cerevisiae; again, the exact numbers vary between species).[48]
Translation
Ribosomes are the workplaces of
Although catalysis of the

In Figure 5, both ribosomal subunits (small and large) assemble at the start codon (towards the 5' end of the mRNA). The ribosome uses tRNA that matches the current codon (triplet) on the mRNA to append an amino acid to the polypeptide chain. This is done for each triplet on the mRNA, while the ribosome moves towards the 3' end of the mRNA. Usually in bacterial cells, several ribosomes are working parallel on a single mRNA, forming what is called a polyribosome or polysome.
Cotranslational folding
The ribosome is known to actively participate in the protein folding.[54][55] The structures obtained in this way are usually identical to the ones obtained during protein chemical refolding; however, the pathways leading to the final product may be different.[56][57] In some cases, the ribosome is crucial in obtaining the functional protein form. For example, one of the possible mechanisms of folding of the deeply knotted proteins relies on the ribosome pushing the chain through the attached loop.[58]
Addition of translation-independent amino acids
Presence of a ribosome quality control protein Rqc2 is associated with mRNA-independent protein elongation.[59][60] This elongation is a result of ribosomal addition (via tRNAs brought by Rqc2) of CAT tails: ribosomes extend the C-terminus of a stalled protein with random, translation-independent sequences of alanines and threonines.[61][62]
Ribosome locations
Ribosomes are classified as being either "free" or "membrane-bound".

Free and membrane-bound ribosomes differ only in their spatial distribution; they are identical in structure. Whether the ribosome exists in a free or membrane-bound state depends on the presence of an ER-targeting signal sequence on the protein being synthesized, so an individual ribosome might be membrane-bound when it is making one protein, but free in the cytosol when it makes another protein.
Ribosomes are sometimes referred to as organelles, but the use of the term organelle is often restricted to describing sub-cellular components that include a phospholipid membrane, which ribosomes, being entirely particulate, do not. For this reason, ribosomes may sometimes be described as "non-membranous organelles".
Free ribosomes
Free ribosomes can move about anywhere in the
Membrane-bound ribosomes
When a ribosome begins to synthesize proteins that are needed in some organelles, the ribosome making this protein can become "membrane-bound". In eukaryotic cells this happens in a region of the endoplasmic reticulum (ER) called the "rough ER". The newly produced polypeptide chains are inserted directly into the ER by the ribosome undertaking
Biogenesis
In bacterial cells, ribosomes are synthesized in the cytoplasm through the
Origin
The ribosome may have first originated as a protoribosome,
As amino acids gradually appeared in the RNA world under prebiotic conditions,[75][76] their interactions with catalytic RNA would increase both the range and efficiency of function of catalytic RNA molecules.[66] Thus, the driving force for the evolution of the ribosome from an ancient self-replicating machine into its current form as a translational machine may have been the selective pressure to incorporate proteins into the ribosome's self-replicating mechanisms, so as to increase its capacity for self-replication.[72][77][78]
Heterogeneous ribosomes
Ribosomes are compositionally heterogeneous between species and even within the same cell, as evidenced by the existence of cytoplasmic and mitochondria ribosomes within the same eukaryotic cells. Certain researchers have suggested that heterogeneity in the composition of ribosomal proteins in mammals is important for gene regulation, i.e., the specialized ribosome hypothesis.[79][80] However, this hypothesis is controversial and the topic of ongoing research.[81][82]
Heterogeneity in ribosome composition was first proposed to be involved in translational control of protein synthesis by Vince Mauro and Gerald Edelman.[83] They proposed the ribosome filter hypothesis to explain the regulatory functions of ribosomes. Evidence has suggested that specialized ribosomes specific to different cell populations may affect how genes are translated.[84] Some ribosomal proteins exchange from the assembled complex with cytosolic copies[85] suggesting that the structure of the in vivo ribosome can be modified without synthesizing an entire new ribosome.
Certain ribosomal proteins are absolutely critical for cellular life while others are not. In
Heterogeneity of ribosomal RNA modifications plays a significant role in structural maintenance and/or function and most mRNA modifications are found in highly conserved regions.[93][94] The most common rRNA modifications are pseudouridylation and 2'-O-methylation of ribose.[95]
See also
- Aminoglycosides
- Biological machines
- Posttranslational modification
- Protein dynamics
- RNA tertiary structure
- Translation (genetics)
- Wobble base pair
- Ada Yonath—Israeli crystallographer known for her pioneering work on the structure of the ribosome, for which she won the Nobel Prize.
References
- ^ Konikkat S (February 2016). Dynamic Remodeling Events Drive the Removal of the ITS2 Spacer Sequence During Assembly of 60S Ribosomal Subunits in S. cerevisiae (Ph.D. thesis). Carnegie Mellon University. Archived from the original on 3 August 2017.
- ISBN 9783131527912.
- PMID 25706898.
- ^ "Scitable by nature translation / RNA translation".
- ^ PMID 34756086.
- PMID 2446672.
- ^ "Ribosomes". Archived from the original on 2009-03-20. Retrieved 2011-04-28.
- PMID 14381428.
- ^ Rheinberger, Hans-Jörg (September 2022). "A Brief History of Protein Biosynthesis and Ribosome Research". Lindau Nobel Laureate Meetings. Retrieved 2023-08-16.
- ^ Roberts RB, ed. (1958). "Introduction". Microsomal Particles and Protein Synthesis. New York: Pergamon Press, Inc.
- ^ "The Nobel Prize in Physiology or Medicine 1974". Nobelprize.org. The Nobel Foundation. Archived from the original on 26 January 2013. Retrieved 10 December 2012.
- ^ "2009 Nobel Prize in Chemistry". The Nobel Foundation. Archived from the original on 28 April 2012. Retrieved 10 December 2012.
- .
- PMID 22550233.
- S2CID 8370119. Archived from the original(PDF) on 2020-11-30.
- ^ S2CID 4419944.
- ^ ISBN 978-0-8153-4072-0.
- ^ ISBN 978-0-495-11464-2.
- PMID 770196.
- PMID 4589893.
- PMID 1225593.
- S2CID 243730576.
- PMID 34908470.
- PMID 19478915.
- ^ S2CID 9099683.
- ^ S2CID 24771575.
- ^ S2CID 206536444.
- S2CID 9751945.
- PMID 22959417.
- PMID 19497863.
- S2CID 58004990.
- PMID 19617523.
- ^ "Specialized Internal Structures of Prokaryotes | Boundless Microbiology". courses.lumenlearning.com. Retrieved 2021-09-24.
- PMID 10357824.
- PMID 4930061.
- PMID 5468266.
- PMID 1981764.
- PMID 32785681.
- ^ PMID 10937989.
- S2CID 1024446.
- S2CID 39505192.
- S2CID 37382005.
- PMID 16292303.
- S2CID 9737925.
- S2CID 13452915.
- S2CID 4419198.
- ^ a b "Specialized Internal Structures of Prokaryotes". courses.lumenlearning.com. Boundless Microbiology. Retrieved 2018-09-27.
- S2CID 2637106.
- PMID 23582332.
- PMID 28743745.
- PMID 14681588.
- PMID 17157507.
- S2CID 24172338.
- PMID 25341659.
- PMID 9407040.
- PMID 1094916.
- PMID 18702035.
- S2CID 52176392.
- PMID 23178123.
- PMID 23479637.
- PMID 25554787.
- ^ Keeley, J.; Gutnikoff, R. (2 January 2015). "Ribosome studies turn up new mechanism of protein synthesis" (Press release). Howard Hughes Medical Institute. Archived from the original on 12 January 2015. Retrieved 16 January 2015.
- ISBN 978-0-8153-4072-0.
- PMID 19879902.
- PMID 36854922.
- ^ PMID 20610545.
- ^ Dabbs ER (1986). Mutant studies on the prokaryotic ribosome. New York: Springer-Verlag.
- PMID 1604315.
- PMID 4909519.
- PMID 21930590.
- PMID 35137169.
- ^ PMID 25500179.
- PMID 12429689.
- ISBN 978-94-007-4898-9.
- S2CID 41725226.
- PMID 22684046.
- PMID 20534711.
- S2CID 27493054.
- PMID 28625553.
- PMID 22617470.
- PMID 30733326.
- PMID 31376929.
- PMID 12221294.
- PMID 22617470.
- PMID 27932527.
- PMID 22377630.
- PMID 11983894.
- PMID 17934214.
- S2CID 10710245.
- PMID 15883184.
- PMID 20410138.
- PMID 19952110.
- PMID 12114023.
- S2CID 4465175.
- S2CID 51609077.
External links
- Lab computer simulates ribosome in motion
- Role of the Ribosome, Gwen V. Childs, copied here
- Ribosome in Proteopedia—The free, collaborative 3D encyclopedia of proteins & other molecules
- Ribosomal proteins families in ExPASy Archived 2011-04-30 at the Wayback Machine
- Molecule of the Month Archived 2009-10-27 at the Wayback Machine © RCSB Protein Data Bank:
- Ribosome Archived 2010-11-14 at the Wayback Machine
- Elongation Factors Archived 2011-03-16 at the Wayback Machine
- Palade
- 3D electron microscopy structures of ribosomes at the EM Data Bank (EMDB)
 This article incorporates NCBI. Archived from the originalon 2009-12-08.
This article incorporates NCBI. Archived from the originalon 2009-12-08.
