Root microbiome
| Part of a series on |
| Microbiomes |
|---|
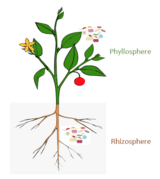 |
The root microbiome (also called rhizosphere microbiome) is the dynamic
Different microorganisms, both beneficial and harmful, affect the development and physiology of plants. Beneficial microorganisms include bacteria that fix nitrogen, various microbes that promote plant growth, mycorrhizal fungi, mycoparasitic fungi, protozoa, and certain biocontrol microorganisms.
Root microbiota affect plant host fitness and productivity in a variety of ways. Members of the root microbiome benefit from plant sugars or other carbon rich molecules. Individual members of the root microbiome may behave differently in association with different plant hosts,[4] or may change the nature of their interaction (along the mutualist-parasite continuum) within a single host as environmental conditions or host health change.[5]
Despite the potential importance of the root microbiome for
Function

Types of symbioses
Root associated microbes include fungi, bacteria, and archaea. In addition, other organisms such as viruses, algae, protozoa, nematodes, and arthropods are part of root microbiota.[1] Symbionts associated with plant roots subsist off of photosynthetic products (carbon rich molecules) from the plant host and can exist anywhere on the mutualist/parasite continuum.
Root symbionts may improve their host's access to nutrients,[14][15][16] produce plant-growth regulators,[17] improve environmental stress tolerance of their host,[18][19][20] induce host defenses and systemic resistance against pests or pathogens,[21][22][23] or be pathogenic.[24] Parasites consume carbon from the plant without providing any benefit or providing insufficient benefit relative to their carbon consumption, thereby compromising host fitness. Symbionts may be biotrophic (subsisting off of living tissue) or necrotrophic (subsisting off of dead tissue).
Mutualist-parasite continuum
While some microbes may be purely
Composition
Roots are colonized by
Fungi
Mycorrhizae
Mycorrhizal (from Greek) literally means "fungus roots" and defines symbiotic interaction between plants and fungi. Fungi are important for decomposing and recycling organic material. However, the boundaries between the pathogenic and symbiotic lifestyles of fungi are not always clear-cut. Most of the time, the association is symbiotic, with the fungus improving nutrient and water acquisition or increasing stress tolerance for the plant and benefiting from the carbohydrates produced by the plant in return.
Endophytes
Endophytes grow inside plant tissue—roots, stems, leaves—mostly symptomless. However, when plants age, they can become slightly pathogenic.
Bacteria
The zone of soil surrounding the roots is rich in nutrients released by plants and is, therefore, an attractive growth medium for both beneficial and pathogenic bacteria. Root associated beneficial bacteria promote plant growth and provide protection from pathogens. They are mostly rhizobacteria that belong to Pseudomonadota and Bacillota, with many examples from Pseudomonas and Bacillus genera.[1] Rhizobium species colonize legume roots forming nodule structures. In response to root exudates, rhizobia produce Nod signalling factors that are recognized by legumes and induce the formation of nodules on plant roots.[28] Within these structures, Rhizobium fix atmospheric nitrogen into ammonia that is then used by the plant. In turn, plants provide the bacteria with a carbon source to energize the nitrogen fixation.[29][30] In addition to nitrogen fixation, Azospirillum species promote plant growth through the production of growth phytohormones (auxins, cytokinins, gibberellins). Due to these phytohormones, root hairs expand to occupy a larger area and better acquire water and nutrients.[29][31] Pathogenic bacteria that infect plants infect plant roots are most commonly from Pectobacterium, Ralstonia, Dickeya and Agrobacterium genera. Among the most notorious are Pectobacterium carotovorum, Pectobacterium atrosepticum, Ralstonia solanacearum, Dickeya dadanthi, Dickeya solani, and Agrobacterium tumefaciens.
Bacteria attach to roots in a biphasic mechanism with two steps—first weak, non-specific binding, then a strong irreversible residence phase. Both beneficial and pathogenic bacteria attach in this fashion. Bacteria can stay attached to the outer surface or colonize the inner root.[29] Primary attachment is governed by chemical forces or extracellular structures such as pili or flagella. Secondary attachment is mainly characterized by the synthesis of cellulose, extracellular fibrils, and specific attachment factors such as surface proteins that help bacteria aggregate and form colonies.[29]
Archaea
Though archaea are often thought of as extremophiles, microbes belonging to extreme environments, advances in metagenomics and gene sequencing have revealed that archaea are found in nearly any environment, including the root microbiome.[8][32][33][34][35][36] For example, root-colonizing archaea have been discovered in maize,[33] rice,[37] wheat,[34] and mangroves.[38] Methanogen and ammonium-oxidizing archaea are prevalent members of the root microbiome, especially in anaerobic soils and wetlands.[32][39][40][41] Archaeal phyla found in the root microbiome include Euryarchaeota,[32][40][42] Nitrososphaerota (formerly Thaumarchaeota),[32][42] and Thermoproteota (formerly Crenarchaeota).[40]
The presence and relative abundance of archaea in various environments suggest that they likely play an important role in the root microbiome.[32] Archaea have been found to promote plant growth and development, provide stress tolerance, improve nutrient uptake, and protect against pathogens.[32][36][43] For example, Arabidopsis thaliana colonized with an ammonia-oxidizing soil archaea, Nitrosocosmicus oleophilius, exhibited increased shoot weight, photosynthetic activity, and immune response.[43]
Examination of microbial communities in soil and roots has identified archaeal organisms and genes with functions similar to that of bacteria and fungi, such as auxin synthesis, protection against abiotic stress, and nitrogen fixation.[36][44] In some cases, key genes for plant growth and development, such as metabolism and cell wall synthesis, are more prevalent in archaea than bacteria.[36]
Archaeal presence in the root microbiome can also be affected by plant hosts, which can change the diversity, presence, and health of archaeal communities.[8][38][45]
Viruses
Viruses also infect plants via the roots; however, to penetrate the root tissues, they typically use vectors such as nematodes or fungi.[1]
Assembly mechanisms
There is an ongoing debate regarding what mechanisms are responsible for assembling individual microbes into
Microbial dispersal mechanisms include wind, water, and hitchhiking on more mobile macrobes. Microbial dispersion is difficult to study, and little is known about its effect on microbial community assembly relative to the effect of abiotic and biotic assembly mechanisms,[7] particularly in roots. For this reason, only assembly mechanisms that fit within the niche hypothesis are discussed below.
The taxa within root microbial communities seem to be drawn from the surrounding soil, though the relative abundance of various taxa may differ greatly from those found in bulk soil due to unique
Biotic assembly mechanisms
Different parts of the root are associated with different microbial communities. For example, fine roots, root tips, and the main root are all associated with different communities,[8][46] and the rhizosphere, root surface, and root tissue are all associated with different communities,[2][3] likely due to the unique chemistry and nutrient status of each of these regions. Additionally, different plant species, and even different cultivars, harbor different microbial communities,[9][10][46] probably due to host specific immune responses[4] and differences in carbon root exudates.[47] Host age affects root microbial community composition, likely for similar reasons as host identity.[8] The identity of neighboring vegetation has also been shown to impact a host plant's root microbial community composition.[9][10][48][49]
Abiotic assembly mechanisms
Abiotic mechanisms also affect root microbial community assembly[9][10][11][12][13] because individual taxa have different optima along various environmental gradients, such as nutrient concentrations, pH, moisture, temperature, etc. In addition to chemical and climatic factors, soil structure and disturbance impact root biotic assembly.[8]
Succession
The root microbiome is dynamic and fluid within the constraints imposed by the biotic and abiotic environment. As in macroecological systems, the historical trajectory of the microbiotic community may partially determine the present and future community. Due to antagonistic and mutualistic interactions between microbial taxa, the taxa colonizing a root at any given moment could be expected to influence which new taxa are acquired, and therefore how the community responds to changes in the host or environment.[7] While the effect of initial community on microbial succession has been studied in various environmental samples, human microbiome, and laboratory settings, it has yet to be studied in roots.
See also
- Mangrove root microbiome
References
- ^ PMID 23790204.
- ^ PMID 21764952.
- ^ ISBN 978-90-481-2665-1
- ^ PMID 16713330.
- ^ ISBN 978-0-08-055934-6.[page needed]
- S2CID 14961515.
- ^ PMID 24006468.
- ^ S2CID 45768977.
- ^ S2CID 15540300.
- ^ PMID 25870878.
- ^ PMID 21426364.
- ^ JSTOR 27646105.
- ^ PMID 22568722.
- ^ PMID 18047587.
- .
- JSTOR 20143553.
- PMID 19575558.
- .
- PMID 18267941.
- PMID 21658184.
- S2CID 12770904.
- S2CID 26283073.
- PMID 25288940.
- S2CID 23292138.
- ISBN 978-0-08-053719-1.[page needed]
- ^ PMID 26591004.
- PMID 33862835.
- PMID 20070373.
- ^ PMID 29672765.
- PMID 23451778.
- PMID 10978548.
- ^ PMID 28826642.
- ^ S2CID 20069511.
- ^ ISSN 2516-8290.
- S2CID 4380804.
- ^ PMID 29743201.
- PMID 25367104.
- ^ S2CID 84753901.
- PMID 18344332.
- ^ S2CID 19039050.
- PMID 18430011.
- ^ PMID 23909555.
- ^ S2CID 53944018.
- PMID 30258074.
- S2CID 22134029.
- ^ .
- PMID 18083870.
- PMID 23078526.
- PMID 22611852.

