Serpin
| Serpin (serine protease inhibitor) | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| CDD | cd00172 | ||||||||||
| |||||||||||
Serpins are a
Protease inhibition by serpins controls an array of biological processes, including
Although most serpins control
History
Protease inhibitory activity in blood plasma was first reported in the late 1800s,
The critical role of the active centre residue in determining the specificity of inhibition of serpins was unequivocally confirmed by the finding that a natural mutation of the active centre methionine in alpha1-antitrypsin to an arginine, as in antithrombin, resulted in a severe bleeding disorder.[28] This active-centre specificity of inhibition was also evident in the many other families of protease inhibitors[7] but the serpins differed from them in being much larger proteins and also in possessing what was soon apparent as an inherent ability to undergo a change in shape. The nature of this conformational change was revealed with the determination in 1984 of the first crystal structure of a serpin, that of post-cleavage alpha1-antitrypsin.[29] This together with the subsequent solving of the structure of native (uncleaved) ovalbumin[30] indicated that the inhibitory mechanism of the serpins involved a remarkable conformational shift, with the movement of the exposed peptide loop containing the reactive site and its incorporation as a middle strand in the main beta-pleated sheet that characterises the serpin molecule.[31][32] Early evidence of the essential role of this loop movement in the inhibitory mechanism came from the finding that even minor aberrations in the amino acid residues that form the hinge of the movement in antithrombin resulted in thrombotic disease.[31][33] Ultimate confirmation of the linked displacement of the target protease by this loop movement was provided in 2000 by the structure of the post-inhibitory complex of alpha1-antitrypsin with trypsin,[6] showing how the displacement results in the deformation and inactivation of the attached protease. Subsequent structural studies have revealed an additional advantage of the conformational mechanism[34] in allowing the subtle modulation of inhibitory activity, as notably seen at tissue level[35] with the functionally diverse serpins in human plasma.
Over 1000 serpins have now been identified, including 36 human proteins, as well as molecules in all
Activity
Most serpins are
Some serpins inhibit other protease classes, typically
Biological function and localization
Protease inhibition
Approximately two-thirds of human serpins perform extracellular roles, inhibiting proteases in the bloodstream in order to modulate their activities. For example, extracellular serpins regulate the proteolytic cascades central to
The table of human serpins (below) provides examples of the range of functions performed by human serpin, as well as some of the diseases that result from serpin deficiency.The protease targets of intracellular inhibitory serpins have been difficult to identify, since many of these molecules appear to perform overlapping roles. Further, many human serpins lack precise functional equivalents in model organisms such as the mouse. Nevertheless, an important function of intracellular serpins may be to protect against the inappropriate activity of proteases inside the cell.
Some
Non-inhibitory roles
Non-inhibitory extracellular serpins also perform a wide array of important roles.
Some serpins are both protease inhibitors and perform additional roles. For example, the nuclear cysteine protease inhibitor
Structure
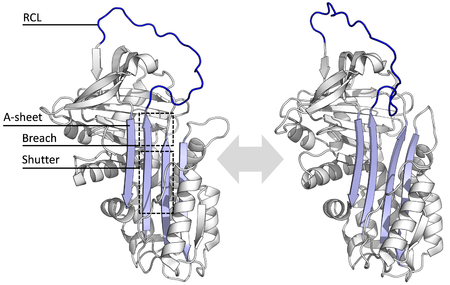
All serpins share a common
The serpin structures that have been determined cover several different conformations, which has been necessary for the understanding of their multiple-step mechanism of action. Structural biology has therefore played a central role in the understanding of serpin function and biology.[8]
Conformational change and inhibitory mechanism
Inhibitory serpins do not inhibit their target proteases by the typical
Since the RCL is still covalently attached to the protease via the ester bond, the S to R transition pulls protease from the top to the bottom of the serpin and distorts the catalytic triad. The distorted protease can only hydrolyse the acyl enzyme intermediate extremely slowly and so the protease remains covalently attached for days to weeks.
Allosteric activation

The
The archetypal example of this situation is antithrombin, which circulates in plasma in a partially inserted relatively inactive state. The primary specificity determining residue (the P1 arginine) points toward the body of the serpin and is unavailable to the protease. Upon binding a high-affinity pentasaccharide sequence within long-chain
Latent conformation
Certain serpins spontaneously undergo the S to R transition without having been cleaved by a protease, to form a conformation termed the latent state. Latent serpins are unable to interact with proteases and so are no longer protease inhibitors. The conformational change to latency is not exactly the same as the S to R transition of a cleaved serpin. Since the RCL is still intact, the first strand of the C-sheet has to peel off to allow full RCL insertion.[68]
Regulation of the latency transition can act as a control mechanism in some serpins, such as
Conformational change in non-inhibitory functions
Certain non-inhibitory serpins also use the serpin conformational change as part of their function. For example, the native (S) form of thyroxine-binding globulin has high affinity for thyroxine, whereas the cleaved (R) form has low affinity. Similarly, transcortin has higher affinity for cortisol when in its native (S) state, than its cleaved (R) state. Thus, in these serpins, RCL cleavage and the S to R transition has been commandeered to allow for ligand release, rather than protease inhibition.[54][55][72]
In some serpins, the S to R transition can activate cell signalling events. In these cases, a serpin that has formed a complex with its target protease, is then recognised by a receptor. The binding event then leads to downstream signalling by the receptor.[73] The S to R transition is therefore used to alert cells to the presence of protease activity.[73] This differs from the usual mechanism whereby serpins affect signalling simply by inhibiting proteases involved in a signalling cascade.[47][48]
Degradation
When a serpin inhibits a target protease, it forms a permanent complex, which needs to be disposed of. For extracellular serpins, the final serpin-enzyme complexes are rapidly cleared from circulation. One mechanism by which this occurs in mammals is via the low-density lipoprotein receptor-related protein (LRP), which binds to inhibitory complexes made by antithrombin, PA1-1, and neuroserpin, causing cellular uptake.[73][74] Similarly, the Drosophila serpin, necrotic, is degraded in the lysosome after being trafficked into the cell by the Lipophorin Receptor-1 (homologous to the mammalian LDL receptor family).[75]
Disease and serpinopathies
Serpins are involved in a wide array of physiological functions, and so mutations in genes encoding them can cause a range of diseases. Mutations that change the activity, specificity or aggregation properties of serpins all affect how they function. The majority of serpin-related diseases are the result of serpin polymerisation into aggregates, though several other types of disease-linked mutations also occur.
Inactivity or absence
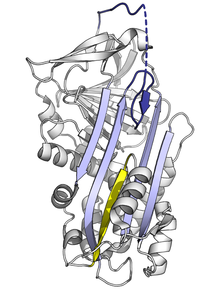
Since the stressed serpin fold is high-energy, mutations can cause them to incorrectly change into their lower-energy conformations (e.g. relaxed or latent) before they have correctly performed their inhibitory role.[10]
Mutations that affect the rate or the extent of RCL insertion into the A-sheet can cause the serpin to undergo its S to R conformational change before having engaged a protease. Since a serpin can only make this conformational change once, the resulting misfired serpin is inactive and unable to properly control its target protease.[10][78] Similarly, mutations that promote inappropriate transition to the monomeric latent state cause disease by reducing the amount of active inhibitory serpin. For example, the disease-linked antithrombin variants wibble and wobble,[79] both promote formation of the latent state.
The structure of the disease-linked mutant of antichymotrypsin (L55P) revealed another, inactive "δ-conformation". In the δ-conformation, four residues of the RCL are inserted into the top of β-sheet A. The bottom half of the sheet is filled as a result of one of the α-helices (the F-helix) partially switching to a β-strand conformation, completing the β-sheet hydrogen bonding.[80] It is unclear whether other serpins can adopt this conformer, and whether this conformation has a functional role, but it is speculated that the δ-conformation may be adopted by Thyroxine-binding globulin during thyroxine release.[55] The non-inhibitory proteins related to serpins can also cause diseases when mutated. For example, mutations in SERPINF1 cause osteogenesis imperfecta type VI in humans.[81]
In the absence of a required serpin, the protease that it normally would regulate is over-active, leading to pathologies.
Specificity change
In some rare cases, a single amino acid change in a serpin's RCL alters its specificity to target the wrong protease. For example, the Antitrypsin-Pittsburgh mutation (M358R) causes the
Polymerisation and aggregation
The majority of serpin diseases are due to
Each monomer of the serpin aggregate exists in the inactive, relaxed conformation (with the RCL inserted into the A-sheet). The polymers are therefore hyperstable to temperature and unable to inhibit proteases. Serpinopathies therefore cause pathologies similarly to other
Physiological serpin polymers are thought to form via domain swapping events, where a segment of one serpin protein inserts into another.[86] Domain-swaps occur when mutations or environmental factors interfere with the final stages of serpin folding to the native state, causing high-energy intermediates to misfold.[87] Both dimer and trimer domain-swap structures have been solved. In the dimer (of antithrombin), the RCL and part of the A-sheet incorporates into the A-sheet of another serpin molecule.[86] The domain-swapped trimer (of antitrypsin) forms via the exchange of an entirely different region of the structure, the B-sheet (with each molecule's RCL inserted into its own A-sheet).[88] It has also been proposed that serpins may form domain-swaps by inserting the RCL of one protein into the A-sheet of another (A-sheet polymerisation).[84][89] These domain-swapped dimer and trimer structures are thought to be the building blocks of the disease-causing polymer aggregates, but the exact mechanism is still unclear.[86][87][88][90]
Therapeutic strategies
Several therapeutic approaches are in use or under investigation to treat the most common serpinopathy: antitrypsin deficiency.
Evolution
Serpins are the most widely distributed and largest superfamily of protease inhibitors.
Protease-inhibition is thought to be the ancestral function, with non-inhibitory members the results of evolutionary
Distribution
Animal
Human
The human genome encodes 16 serpin clades, termed serpinA through serpinP, including 29 inhibitory and 7 non-inhibitory serpin proteins.
| Gene name | Common Name | Localisation | Function / Activity[9][83] | Effect of deficiency[9][83] | Human disease | Chromosomal location | Protein structure |
|---|---|---|---|---|---|---|---|
SERPINA1
|
α1-antitrypsin | Extracellular | Inhibitor of human neutrophil elastase.[99] The C-terminal fragment of cleaved SERPINA1 may inhibit HIV-1 infection.[100] | 14q32.1 | 1QLP, 7API, 1D5S | ||
| SERPINA2 | Antitrypsin-related protein
|
Extracellular | Possible pseudogene.[102] | 14q32.1 | |||
SERPINA3
|
α1-antichymotrypsin | Extracellular | Inhibitor of cathepsin G.[103] Additional roles in chromatin condensation in hepatic cells.[104] | Mis-regulation results in Alzheimer's disease (serpinopathy).[105] | 14q32.1 | 1YXA, 2ACH | |
SERPINA4
|
Kallistatin | Extracellular | Inhibitor of kallikrein, regulator of vascular function.[106][107] | Depletion in hypertensive rats exacerbates renal and cardiovascular injury.[108] | 14q32.1 | ||
SERPINA5
|
Protein C inhibitor | Extracellular | Inhibitor of active protein C.[109] Intracellular role in preventing phagocytosis of bacteria.[110] | Knockout in male mice causes infertility.[111] Accumulation occurs in chronic active plaques in multiple sclerosis.[112] | 14q32.1 | 2OL2, 3B9F | |
SERPINA6
|
Transcortin | Extracellular | Non-inhibitory. Cortisol binding.[54] | Deficiency associated with chronic fatigue.[113] | 14q32.1 | 2V6D, 2VDX, 2VDY | |
SERPINA7
|
Thyroxine-binding globulin | Extracellular | Non-inhibitory. Thyroxine binding.[55] | Deficiency causes hypothyroidism.[114][115] | Xq22.2 | 2CEO, 2RIV, 2RIW | |
SERPINA8
|
Angiotensinogen
|
Extracellular | Non-inhibitory, cleavage by renin results in release of angiotensin I.[116] | Knockout in mice causes hypotension.[117] | Variants linked to hypertension.[118][119][120] | 1q42-q43 | 2X0B, 2WXW, 2WXX, 2WXY, 2WXZ, 2WY0, 2WY1 |
| SERPINA9 | Centerin / GCET1
|
Extracellular | Inhibitory, maintenance of naive B cells.[121][122] | Strongly expressed in most B-cell lymphomas.[123][124] | 14q32.1 | ||
SERPINA10
|
Protein Z-related protease inhibitor | Extracellular | Binds factor Xa and factor XIa.[125]
|
14q32.1 | 3F1S, 3H5C | ||
| SERPINA11 | – | Probably extracellular | Unknown | 14q32.13 | |||
| SERPINA12 | Vaspin | Extracellular | Inhibitor of Kallikrein-7. Insulin-sensitizing adipocytokine.[126] | High plasma levels associated with type II diabetes.[127] | 14q32.1 | 4IF8 | |
| SERPINA13 | – | Probably extracellular | Unknown | 14q32 | |||
| SERPINB1 | Monocyte neutrophil elastase inhibitor | Intracellular | Inhibitor of neutrophil elastase.[128] | Knockout in mice causes neutrophil survival defect and immune deficiency.[129] | 6p25 | 1HLE | |
SERPINB2
|
Plasminogen activator inhibitor-2 | Intracellular/extracellular | Inhibitor of extracellular uPA. Intracellular function unclear, but may protect against viral infection.[130] | Deficiency in mice reduces immune response to nematode infection.[131] Knockout in mice causes no obvious phenotype.[132] | 18q21.3 | 1BY7 | |
| SERPINB3 | Squamous cell carcinoma antigen-1 (SCCA-1)
|
Intracellular | Inhibitor of papain-like cysteine proteases[44] and cathepsins K, L and S.[133][134] | Knockout in mice of Serpinb3a (the murine | 18q21.3 | 2ZV6 | |
| SERPINB4 | Squamous cell carcinoma antigen-2 (SCCA-2)
|
Intracellular | Inhibitor of chymotrypsin-like serine proteases, cathepsin G and chymase.[134][136] | Knockout in mice of Serpinb3a (the murine | 18q21.3 | ||
SERPINB5
|
Maspin | Intracellular | Non-inhibitory, function unclear[137][138][139] (see also maspin) | Knockout in mice originally reported as lethal,[140] but subsequently shown to have no obvious phenotype.[139] Expression may be a prognostic indicator that reflects expression of a neighbouring tumour suppressor gene (the phosphatase PHLPP1).[139] | 18q21.3 | 1WZ9 | |
| SERPINB6 | PI-6 | Intracellular | Inhibitor of cathepsin G.[141] | Knockout in mice causes hearing loss[142] and mild neutropenia.[143] | Deficiency associated with hearing loss.[144] | 6p25 | |
| SERPINB7 | Megsin
|
Intracellular | Involved in megakaryocyte maturation.[145] | Over-expression in mice causes kidney disease.[146] Knockout in mice does not cause histological abnormalities.[146] | Mutations associated with Nagashima-type Palmoplantar Keratosis.[147] | 18q21.3 | |
| SERPINB8 | PI-8 | Intracellular | Possible inhibitor of furin.[148] | 18q21.3 | |||
| SERPINB9 | PI-9 | Intracellular | Inhibitor of the cytotoxic granule protease granzyme B.[149] | Knockout in mice causes immune dysfunction.[150][151] | 6p25 | ||
| SERPINB10 | Bomapin | Intracellular | Unknown[152] | Knockout in mice causes no obvious phenotype (C57/BL6; lab strain BC069938). | 18q21.3 | ||
| SERPINB11 | Intracellular | Unknown[153] | Murine Serpinb11 is an active inhibitor whereas the human orthalogue is inactive.[153] Deficiency in ponies is associated with hoof wall separation disease.[154] | 18q21.3 | |||
| SERPINB12 | Yukopin | Intracellular | Unknown[155] | 18q21.3 | |||
| SERPINB13 | Hurpin/Headpin | Intracellular | Inhibitor of papain-like cysteine proteases.[156] | 18q21.3 | |||
SERPINC1
|
Antithrombin | Extracellular | Inhibitor of coagulation, specifically factor X, factor IX and thrombin.[34] | Knockouts in mice are lethal.[157] | Deficiency results in thrombosis and other clotting disorders (serpinopathy).[158][159] | 1q23-q21 | 2ANT, 2ZNH, 1AZX, 1TB6, 2GD4, 1T1F |
SERPIND1
|
Heparin cofactor II | Extracellular | Inhibitor of thrombin.[160] | Knockouts in mice are lethal.[161] | 22q11 | 1JMJ, 1JMO | |
SERPINE1
|
Plasminogen activator inhibitor 1
|
Extracellular | Inhibitor of thrombin, uPA and TPa.[162] | 7q21.3-q22 | 1DVN, 1OC0 | ||
| SERPINE2 | Glia derived nexin / Protease nexin I | Extracellular | Inhibitor of uPA and tPA.[163] | Abnormal expression leads to male infertility.[164] Knockout in mice causes epilepsy.[165] | 2q33-q35 | 4DY0 | |
SERPINF1
|
Pigment epithelium derived factor
|
Extracellular | Non-inhibitory, potent anti-angiogenic molecule.[166] PEDF has been reported to bind the glycosaminoglycan hyaluronan.[167] | Knockout in mice affects the vasculature and mass of the pancreas and the prostate.[166] Promotes Notch–dependent renewal of adult periventricular neural stem cells.[168] Mutations in humans cause osteogenesis imperfecta type VI.[81] | 17p13.3 | 1IMV | |
SERPINF2
|
α2-antiplasmin | Extracellular | Inhibitor of plasmin, inhibitor of fibrinolysis.[169] | Knockouts in mice show increased mice show increased fibrinolysis but no bleeding disorder.[170] | Deficiency causes a rare bleeding disorder.[171][172] | 17pter-p12 | 2R9Y |
SERPING1
|
Complement 1-inhibitor | Extracellular | Inhibitor of C1 esterase.[173] | Several polymorphisms associated with macular degeneration[174] and hereditary angeoedema.[175] | 11q11-q13.1 | 2OAY | |
SERPINH1
|
47 kDa Heat shock protein (HSP47)
|
Intracellular | Non-inhibitory, molecular chaperone in collagen folding.[57] | Knockouts in mice are lethal.[176] | Mutation in humans causes severe osteogenesis imperfecta.[177][178] | 11p15 | 4AXY |
SERPINI1
|
Neuroserpin
|
Extracellular | Inhibitor of tPA, uPA and plasmin.[179] | Mutation causes FENIB dementia (serpinopathy).[180][181] | 3q26 | 1JJO, 3FGQ, 3F5N, 3F02 | |
| SERPINI2 | Pancpin | Extracellular | Unknown[182] | Deficiency in mice causes pancreatic insufficiency via acinar cell loss.[183] | 3q26 |
Specialised mammalian serpins
Many
Insect
The Drosophila melanogaster genome contains 29 serpin encoding genes. Amino acid sequence analysis has placed 14 of these serpins in serpin clade Q and three in serpin clade K with the remaining twelve classified as orphan serpins not belonging to any clade.[186] The clade classification system is difficult to use for Drosophila serpins and instead a nomenclature system has been adopted that is based on the position of serpin genes on the Drosophila chromosomes. Thirteen of the Drosophila serpins occur as isolated genes in the genome (including Serpin-27A, see below), with the remaining 16 organised into five gene clusters that occur at chromosome positions 28D (2 serpins), 42D (5 serpins), 43A (4 serpins), 77B (3 serpins) and 88E (2 serpins).[186][187][188]
Studies on Drosophila serpins reveal that Serpin-27A inhibits the Easter protease (the final protease in the Nudel, Gastrulation Defective, Snake and Easter proteolytic cascade) and thus controls
Nematode
The genome of the nematode worm C. elegans contains 9 serpins, all of which lack signal sequences and so are likely intracellular.[192] However, only 5 of these serpins appear to function as protease inhibitors.[192] One, SRP-6, performs a protective function and guards against stress-induced calpain-associated lysosomal disruption. Further, SRP-6 inhibits lysosomal cysteine proteases released after lysosomal rupture. Accordingly, worms lacking SRP-6 are sensitive to stress. Most notably, SRP-6 knockout worms die when placed in water (the hypo-osmotic stress lethal phenotype or Osl). It has therefore been suggested that lysosomes play a general and controllable role in determining cell fate.[193]
Plant
Plant serpins were amongst the first members of the superfamily that were identified.[194] The serpin barley protein Z is highly abundant in barley grain, and one of the major protein components in beer. The genome of the model plant, Arabidopsis thaliana contain 18 serpin-like genes, although only 8 of these are full-length serpin sequences.
Plant serpins are potent inhibitors of mammalian chymotrypsin-like serine proteases in vitro, the best-studied example being barley serpin Zx (BSZx), which is able to inhibit trypsin and chymotrypsin as well as several blood coagulation factors.[195] However, close relatives of chymotrypsin-like serine proteases are absent in plants. The RCL of several serpins from wheat grain and rye contain poly-Q repeat sequences similar to those present in the prolamin storage proteins of the endosperm.[196][197] It has therefore been suggested that plant serpins may function to inhibit proteases from insects or microbes that would otherwise digest grain storage proteins. In support of this hypothesis, specific plant serpins have been identified in the phloem sap of pumpkin (CmPS-1)[198] and cucumber plants.[199][200] Although an inverse correlation between up-regulation of CmPS-1 expression and aphid survival was observed, in vitro feeding experiments revealed that recombinant CmPS-1 did not appear to affect insect survival.[198]
Alternative roles and protease targets for plant serpins have been proposed. The Arabidopsis serpin, AtSerpin1 (At1g47710; 3LE2), mediates set-point control over programmed cell death by targeting the 'Responsive to Desiccation-21' (RD21) papain-like cysteine protease.[53][201] AtSerpin1 also inhibits metacaspase-like proteases in vitro.[52] Two other Arabidopsis serpins, AtSRP2 (At2g14540) and AtSRP3 (At1g64030) appear to be involved in responses to DNA damage.[202]
Fungal
A single
Prokaryotic
Predicted serpin genes are sporadically distributed in
Viral
Serpins are also expressed by viruses as a way to evade the host's immune defense.[206] In particular, serpins expressed by pox viruses, including cow pox (vaccinia) and rabbit pox (myxoma), are of interest because of their potential use as novel therapeutics for immune and inflammatory disorders as well as transplant therapy.[207][208] Serp1 suppresses the TLR-mediated innate immune response and allows indefinite cardiac allograft survival in rats.[207][209] Crma and Serp2 are both cross-class inhibitors and target both serine (granzyme B; albeit weakly) and cysteine proteases (caspase 1 and caspase 8).[210][211] In comparison to their mammalian counterparts, viral serpins contain significant deletions of elements of secondary structure. Specifically, crmA lacks the D-helix as well as significant portions of the A- and E-helices.[212]
References
- ^ PMID 11435447.
- PMID 33744972.
- )
- PMID 20498369.
- PMID 20498368.
- ^ S2CID 205009937.
- ^ PMID 12475206.
- ^ PMID 17079131.
- ^ PMID 16737556.
- ^ S2CID 21223825.
- ^ PMID 21367592.
- ^ S2CID 39124185.
- S2CID 24373770.
- S2CID 95960716.
- ^ Petersen TE, Dudeck-Wojciechowska G, Sottrup-Jensen L, Magnusson S (1979). "Primary structure of antithrombin III (heparin cofactor): partial homology between alpha-1-antitrypsin and antithrombin III". In Collen D, Wiman B, Verstraete M (eds.). The Physiological Inhibitors of Coagulation and Fibrinolysis. Amsterdam: Elsevier. pp. 43–54.
- PMID 316698.
- PMID 6968211.
- PMID 7045697.
- S2CID 36366089.
- PMID 4182334.
- PMID 1082356.
- S2CID 84576569.
- S2CID 11904305.
- PMID 6966929.
- PMID 701239.
- S2CID 32540550.
- S2CID 42594050.
- ^ PMID 6604220.
- ^ PMID 6332197.
- ^ S2CID 4342263.
- ^ S2CID 4361304.
- S2CID 4365370.
- PMID 2029579.
- ^ PMID 16820297.
- PMID 28027946.
- ^ PMID 11116082.
- ^ PMID 12411597.
- ^ PMID 18539447.
- PMID 2690952.
- ^ PMID 14705960.
- PMID 7733651.
- S2CID 37306786.
- PMID 11939796.
- ^ PMID 9811823.
- ^ PMID 16810322.
- PMID 17923478.
- ^ PMID 25758225.
- ^ PMID 14667416.
- S2CID 44268106.
- PMID 9774654.
- S2CID 7398844.
- ^ PMID 17028019.
- ^ PMID 20181955.
- ^ PMID 17644521.
- ^ PMID 16938877.
- PMID 11419711.
- ^ PMID 20888348.
- PMID 10026180.
- S2CID 22976014.
- PMID 16141197.
- PMID 10669617.
- PMID 9405673.
- PMID 10966821.
- S2CID 28790576.
- PMID 16619025.
- S2CID 24796086.
- PMID 15199558.
- ^ S2CID 19433778.
- S2CID 43029244.
- PMID 17557112.
- PMID 18234218.
- S2CID 4326356.
- ^ PMID 16601674.
- PMID 19439404.
- PMID 19557185.
- S2CID 20760165.
- PMID 12426287.
- PMID 8347575.
- PMID 9763552.
- ^ PMID 10618372.
- ^ PMID 21826736.
- PMID 9207454.
- ^ PMID 24172014.
- ^ S2CID 4359543.
- PMID 19549782.
- ^ S2CID 205215121.
- ^ PMID 21921939.
- ^ PMID 21909074.
- PMID 9007980.
- S2CID 8188156.
- PMID 15454659.
- PMID 22634722.
- PMID 18565211.
- PMID 21993621.
- PMID 17918823.
- PMID 21103396.
- PMID 17635906.
- PMID 22085334.
- S2CID 54415934.
- PMID 17448989.
- PMID 24374162.
- PMID 17135331.
- S2CID 11230631.
- PMID 23295442.
- PMID 12214135.
- PMID 9202051.
- PMID 12384424.
- PMID 22811485.
- S2CID 4828322.
- PMID 18025309.
- PMID 11120760.
- S2CID 4421395.
- S2CID 43352358.
- PMID 1422238.
- PMID 22851492.
- S2CID 24041932.
- PMID 7989296.
- S2CID 42614761.
- PMID 12805070.
- PMID 16754793.
- PMID 11069088.
- PMID 17447896.
- PMID 18550480.
- PMID 23624075.
- PMID 11049983.
- PMID 16030142.
- PMID 25151227.
- PMID 1376927.
- PMID 17664292.
- PMID 9607921.
- PMID 23630350.
- PMID 9892694.
- S2CID 35146299.
- ^ PMID 25984243.
- ^ PMID 21126757.
- PMID 8999871.
- PMID 20123984.
- PMID 8290962.
- ^ PMID 24445777.
- PMID 14985257.
- PMID 10068683.
- PMID 23669344.
- PMID 15082799.
- PMID 20451170.
- PMID 11877466.
- ^ PMID 18645605.
- PMID 25029323.
- PMID 9442015.
- PMID 8910377.
- PMID 16618603.
- S2CID 39276036.
- PMID 9454755.
- ^ PMID 17562709.
- PMID 25875171.
- PMID 11604408.
- PMID 12809493.
- PMID 11018075.
- S2CID 20768425.
- S2CID 1020630.
- PMID 15315969.
- PMID 17549254.
- PMID 17896949.
- PMID 17409231.
- PMID 11248026.
- PMID 9169529.
- ^ S2CID 5967666.
- PMID 18805795.
- S2CID 5332822.
- PMID 158022.
- PMID 10090937.
- S2CID 205295156.
- S2CID 71010865.
- PMID 17488724.
- PMID 17768101.
- PMID 25827463.
- PMID 10995453.
- PMID 25007323.
- PMID 23145505.
- PMID 9442076.
- S2CID 22326349.
- PMID 19164889.
- S2CID 9525421.
- PMID 16184191.
- S2CID 9248169.
- PMID 20678169.
- ^ PMID 16260136.
- PMID 18854145.
- PMID 18801354.
- PMID 14711428.
- PMID 14654000.
- PMID 21862574.
- ^ PMID 14739286.
- PMID 17889653.
- S2CID 84790014.
- PMID 8843156.
- S2CID 27933086.
- PMID 10874043.
- ^ PMID 10960478.
- PMID 16246856.
- S2CID 22960858.
- PMID 23398119.
- PMID 19426562.
- ^ S2CID 23738804.
- PMID 12679017.
- PMID 15590653.
- PMID 12297326.
- ^ PMID 16146796.
- S2CID 20168458.
- PMID 12637546.
- PMID 10400732.
- PMID 17214579.
- PMID 10903953.
External links
- PDB Molecule of the Month Serpin
- Merops protease inhibitor claudication (Family I4) Archived 8 December 2016 at the Wayback Machine
- Serpins at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- James Whisstock laboratory at Monash University
- Jim Huntington laboratory Archived 30 October 2016 at the Wayback Machine at University of Cambridge
- Frank Church laboratory at University of North Carolina at Chapel Hill
- Paul Declerck laboratory at Katholieke Universiteit Leuven
- Tom Roberts laboratory at University of Sydney
- Robert Fluhr laboratory at Weizmann Institute of Science
- Peter Gettins laboratory at University of Illinois at Chicago
- Overview of all the structural information available in the PDB for UniProt: P01009 (Human Alpha-1-antitrypsin) at the PDBe-KB.


