Signal transduction
When signaling pathways interact with one another they form networks, which allow cellular responses to be coordinated, often by combinatorial signaling events.
Each component (or node) of a signaling pathway is classified according to the role it plays with respect to the initial stimulus.

Stimuli

The basis for signal transduction is the transformation of a certain stimulus into a biochemical signal. The nature of such stimuli can vary widely, ranging from extracellular cues, such as the presence of
Ligands
The majority of signal transduction pathways involve the binding of signaling molecules, known as ligands, to receptors that trigger events inside the cell. The binding of a signaling molecule with a receptor causes a change in the conformation of the receptor, known as receptor activation. Most ligands are soluble molecules from the extracellular medium which bind to
Not all classifications of signaling molecules take into account the molecular nature of each class member. For example,
Some receptors such as HER2 are capable of ligand-independent activation when overexpressed or mutated. This leads to constitutive activation of the pathway, which may or may not be overturned by compensation mechanisms. In the case of HER2, which acts as a dimerization partner of other EGFRs, constitutive activation leads to hyperproliferation and cancer.[14]
Mechanical forces
The prevalence of
Osmolarity
Cellular and systemic control of
Temperature
The sensing of temperature in cells is known as thermoception and is primarily mediated by
Light
In mammals,
Receptors
Receptors can be roughly divided into two major classes:
Extracellular receptors
Extracellular receptors are
In
Many adaptor proteins and enzymes activated as part of signal transduction possess specialized
G protein–coupled receptors
G protein–coupled receptors (GPCRs) are a family of integral transmembrane proteins that possess seven transmembrane domains and are linked to a heterotrimeric
Signal transduction by a GPCR begins with an inactive G protein coupled to the receptor; the G protein exists as a heterotrimer consisting of Gα, Gβ, and Gγ subunits.
A study was conducted where a point mutation was inserted into the gene encoding the chemokine receptor CXCR2; mutated cells underwent a malignant transformation due to the expression of CXCR2 in an active conformation despite the absence of chemokine-binding. This meant that chemokine receptors can contribute to cancer development.[30]
Tyrosine, Ser/Thr and Histidine-specific protein kinases
As is the case with GPCRs, proteins that bind GTP play a major role in signal transduction from the activated RTK into the cell. In this case, the G proteins are members of the
Histidine-specific protein kinases are structurally distinct from other protein kinases and are found in prokaryotes, fungi, and plants as part of a two-component signal transduction mechanism: a phosphate group from ATP is first added to a histidine residue within the kinase, then transferred to an aspartate residue on a receiver domain on a different protein or the kinase itself, thus activating the aspartate residue.[36]
Integrins

Integrins are produced by a wide variety of cells; they play a role in cell attachment to other cells and the extracellular matrix and in the transduction of signals from extracellular matrix components such as fibronectin and collagen. Ligand binding to the extracellular domain of integrins changes the protein's conformation, clustering it at the cell membrane to initiate signal transduction. Integrins lack kinase activity; hence, integrin-mediated signal transduction is achieved through a variety of intracellular protein kinases and adaptor molecules, the main coordinator being integrin-linked kinase.[37] As shown in the adjacent picture, cooperative integrin-RTK signaling determines the timing of cellular survival, apoptosis, proliferation, and differentiation.
Important differences exist between integrin-signaling in circulating blood cells and non-circulating cells such as
In plants, there are no bona fide integrin receptors identified to date; nevertheless, several integrin-like proteins were proposed based on structural homology with the metazoan receptors.
Toll-like receptors
When activated, toll-like receptors (TLRs) take adapter molecules within the cytoplasm of cells in order to propagate a signal. Four adaptor molecules are known to be involved in signaling, which are
Ligand-gated ion channels
A ligand-gated ion channel, upon binding with a ligand, changes conformation to open a channel in the cell membrane through which ions relaying signals can pass. An example of this mechanism is found in the receiving cell of a neural
An example of an ion allowed into the cell during a ligand-gated ion channel opening is Ca2+; it acts as a second messenger initiating signal transduction cascades and altering the physiology of the responding cell. This results in amplification of the synapse response between synaptic cells by remodelling the
Intracellular receptors
Intracellular receptors, such as
Activated nuclear receptors attach to the DNA at receptor-specific
Nucleic receptors have DNA-binding domains containing zinc fingers and a ligand-binding domain; the zinc fingers stabilize DNA binding by holding its phosphate backbone. DNA sequences that match the receptor are usually hexameric repeats of any kind; the sequences are similar but their orientation and distance differentiate them. The ligand-binding domain is additionally responsible for dimerization of nucleic receptors prior to binding and providing structures for transactivation used for communication with the translational apparatus.
Retinoic acid receptors are another subset of nuclear receptors. They can be activated by an endocrine-synthesized ligand that entered the cell by diffusion, a ligand synthesised from a precursor like retinol brought to the cell through the bloodstream or a completely intracellularly synthesised ligand like prostaglandin. These receptors are located in the nucleus and are not accompanied by HSPs. They repress their gene by binding to their specific DNA sequence when no ligand binds to them, and vice versa.
Certain intracellular receptors of the immune system are cytoplasmic receptors; recently identified
Second messengers
First messengers are the signaling molecules (hormones, neurotransmitters, and paracrine/autocrine agents) that reach the cell from the extracellular fluid and bind to their specific receptors. Second messengers are the substances that enter the cytoplasm and act within the cell to trigger a response. In essence, second messengers serve as chemical relays from the plasma membrane to the cytoplasm, thus carrying out intracellular signal transduction.
Calcium
The release of calcium ions from the
Calcium is used in many processes including muscle contraction, neurotransmitter release from nerve endings, and cell migration. The three main pathways that lead to its activation are GPCR pathways, RTK pathways, and gated ion channels; it regulates proteins either directly or by binding to an enzyme.
Lipid messengers
Lipophilic second messenger molecules are derived from lipids residing in cellular membranes; enzymes stimulated by activated receptors activate the lipids by modifying them. Examples include diacylglycerol and ceramide, the former required for the activation of protein kinase C.
Nitric oxide
Nitric oxide (NO) acts as a second messenger because it is a
Redox signaling
In addition to nitric oxide, other electronically activated species are also signal-transducing agents in a process called
Cellular responses
Gene activations
Mammalian cells require stimulation for cell division and survival; in the absence of growth factor, apoptosis ensues. Such requirements for extracellular stimulation are necessary for controlling cell behavior in unicellular and multicellular organisms; signal transduction pathways are perceived to be so central to biological processes that a large number of diseases are attributed to their dysregulation. Three basic signals determine cellular growth:
- Stimulatory (growth factors)
- Transcription dependent response
For example, steroids act directly as transcription factor (gives slow response, as transcription factor must bind DNA, which needs to be transcribed. Produced mRNA needs to be translated, and the produced protein/peptide can undergoposttranslational modification(PTM)) - Transcription independent response
For example, epidermal growth factor (EGF) binds the epidermal growth factor receptor (EGFR), which causes dimerization and autophosphorylation of the EGFR, which in turn activates the intracellular signaling pathway .[52]
- Transcription dependent response
- Inhibitory (cell-cell contact)
- Permissive (cell-matrix interactions)
The combination of these signals is integrated into altered cytoplasmic machinery which leads to altered cell behaviour.
Major pathways
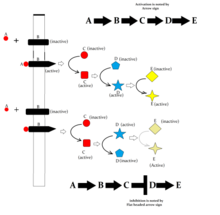
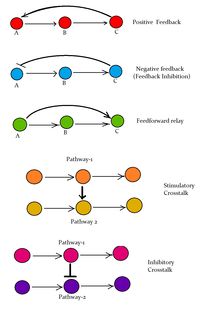
Following are some major signaling pathways, demonstrating how ligands binding to their receptors can affect second messengers and eventually result in altered cellular responses.
- MAPK/ERK pathway: A pathway that couples intracellular responses to the binding of growth factors to cell surface receptors. This pathway is very complex and includes many protein components.[53] In many cell types, activation of this pathway promotes cell division, and many forms of cancer are associated with aberrations in it.[54]
- cAMP-dependent protein kinase, which vary based on the type of cell.
- IP3 receptors, particular calcium channels in the endoplasmic reticulum (ER). These channels are specific to calcium and allow the passage of only calcium to move through. This causes the cytosolic concentration of Calcium to increase, causing a cascade of intracellular changes and activity.[55] In addition, calcium and DAG together works to activate PKC, which goes on to phosphorylate other molecules, leading to altered cellular activity. End-effects include taste, manic depression, tumor promotion, etc.[55]
History

The earliest notion of signal transduction can be traced back to 1855, when
The discovery of
In 1970,
Signal transduction in Immunology
The purpose of this section is to briefly describe some developments in immunology in the 1960s and 1970s, relevant to the initial stages of transmembrane signal transduction, and how they impacted our understanding of immunology, and ultimately of other areas of cell biology.
The relevant events begin with the sequencing of myeloma protein light chains, which are found in abundance in the urine of individuals with multiple myeloma. Biochemical experiments revealed that these so-called Bence Jones proteins consisted of 2 discrete domains –one that varied from one molecule to the next (the V domain) and one that did not (the Fc domain or the Fragment crystallizable region).[67] An analysis of multiple V region sequences by Wu and Kabat [68] identified locations within the V region that were hypervariable and which, they hypothesized, combined in the folded protein to form the antigen recognition site. Thus, within a relatively short time a plausible model was developed for the molecular basis of immunological specificity, and for mediation of biological function through the Fc domain. Crystallization of an IgG molecule soon followed [69] ) confirming the inferences based on sequencing, and providing an understanding of immunological specificity at the highest level of resolution.
The biological significance of these developments was encapsulated in the theory of clonal selection[70] which holds that a B cell has on its surface immunoglobulin receptors whose antigen-binding site is identical to that of antibodies that are secreted by the cell when it encounters an antigen, and more specifically a particular B cell clone secretes antibodies with identical sequences. The final piece of the story, the Fluid mosaic model of the plasma membrane provided all the ingredients for a new model for the initiation of signal transduction; viz, receptor dimerization.
The first hints of this were obtained by Becker et al
Such observations led to a number of theoretical (mathematical) developments. The first of these was a simple model proposed by Bell [74] which resolved an apparent paradox: clustering forms stable networks; i.e. binding is essentially irreversible, whereas the affinities of antibodies secreted by B cells increase as the immune response progresses. A theory of the dynamics of cell surface clustering on lymphocyte membranes was developed by DeLisi and Perelson [75] who found the size distribution of clusters as a function of time, and its dependence on the affinity and valence of the ligand. Subsequent theories for basophils and mast cells were developed by Goldstein and Sobotka and their collaborators,[76][77] all aimed at the analysis of dose-response patterns of immune cells and their biological correlates.[78] For a recent review of clustering in immunological systems see.[79]
Ligand binding to cell surface receptors is also critical to motility, a phenomenon that is best understood in single-celled organisms. An example is a detection and response to concentration gradients by bacteria [80]-–the classic mathematical theory appearing in.[81] A recent account can be found in [82]
See also
- Adaptor protein
- Scaffold protein
- Biosemiotics
- Cell signaling
- Gene regulatory network
- Hormonal imprinting
- Metabolic pathway
- Protein–protein interaction
- Two-component regulatory system
References
- ISBN 9780123741455.
- S2CID 3065483.
- ISBN 978-3527313976.
- ISBN 978-0-8053-6624-2.
- S2CID 35252401.
- ^ Bago R, Sommer E, Castel P, Crafter C, Bailey FP, Shpiro N, Baselga J, Cross D, Eyers PA, Alessi DR (2016) The hVps34-SGK3 pathway alleviates sustained PI3K/Akt inhibition by stimulating mTORC1 and tumour growth. EMBO Journal 35:1902-22
- PMID 12169636.
- PMID 16581495.
- S2CID 20654561.
- PMID 12606724.
- PMID 11826268.
- PMID 9457173.
- PMID 959823.
- PMID 23000897.
- S2CID 205225137.
- S2CID 21267494.
- S2CID 4374012.
- ^ PMID 21852585.
- PMID 18003769.
- PMID 12040128.
- ^ PMID 23618661.
- S2CID 4311262.
- PMID 16269358.
- PMID 17351786.
- ^ "A molecular model for receptor activation" (PDF).
- ^ S2CID 7938806.
- PMID 21873996.
- ISBN 978-0-7167-4954-7.
- PMID 17203515.
- S2CID 45743458.
- ^ PMID 16700535.
- PMID 3075366.
- S2CID 26554314.
- PMID 24818526.
- PMID 15158434.
- PMID 12372152.
- ^ PMID 17084981.
- PMID 16725254.
- PMID 21398259.
- ^ PMID 27208244.
- PMID 17360592.
- S2CID 19276476.
- S2CID 13016860.
- S2CID 16163262.
- PMID 17126540.
- ^ ISBN 978-0-07-067065-5.
- PMID 25422863.
- PMID 19735727.
- PMID 8021233.
- ^ PMID 2442814.
- PMID 16678165.
- S2CID 25515586.
- PMID 16293107.
- S2CID 205383514.
- ^ ISBN 978-0-8153-3218-3.
- ^ a b Bradshaw & Dennis (2010) p. 1.
- PMID 15940278.
- S2CID 6747529.
- ^ Bradshaw & Dennis (2010) p. 2.
- ^ S2CID 5650340.
- PMID 4621276.
- PMID 169771.
- PMID 782447.
- S2CID 62775788.
- S2CID 29355685.
- ISBN 978-0-12-289631-6.
- ^ Steiner, L A (1996) Immunoglobulin evolution, 30 years on. Glycobiology 6 , 649-656
- ^ Wu, T T, Kabat, E A (1970) An analysis of the sequences of the variable regions of Bence Jones proteins and myeloma light chains and their implications for antibody complementarity. J. Exp. Med. 132: 211-250
- ^ Sarma, V R, Silverton, E W, Davies, D R, Terry W D (1971) The three-dimensional structure at 6 A resolution of a human gamma G1 immunoglobulin molecule, J Biol. Chem. 246 (11) 3752- 9
- ^ Burnet, F M (1976) A modification of Jerne's theory of antibody production using the concept of clonal selection. CA: A Cancer Journal for Clinicians 26 (2) 119–21
- ^ Becker, K E, Ishizaka, T, Metzger, H, Ishizaka, K and Grimley, P M (1973) Surface IgE on Human Basophils during histamine release. J Exp med, 138, 394-408
- ^ Fanger, M W, Hart, D A, Wells, J V, and Nisonoff, A J (1970) Requirement for cross-linkage in the stimulation of transformation of rabbit peripheral lymphocytes by antiglobulin reagents J. Immun., 105, 1484 - 92
- ^ Klemm J D, Schreiber S L, Crabtree G R (1998) Ann. Rev. Immunol. Dimerization as a regulatory mechanism in signal transduction 16: 569-592
- ^ Bell, G I (1974) Model for the binding of multivalent antigens to cells, Nature Lond. 248, 430
- ^ DeLisi, C and Perelson A (1976). The kinetics of aggregation phenomena, J. theor. Biol. 62, 159-210
- ^ Dembo, M and Goldstein, B (1978) Theory of equilibrium binding of symmetric bivalent haptens to cell surface antibody: application to histamine release from basophils. The Journal of Immunology 121 (1), 345-353
- ^ Sobotka, A.K. Dembo, M, Goldstein, B and Lichtenstein, L M, (1979) Antigen-specific desensitization of human basophils The Journal of Immunology, 122 (2) 511-517
- ^ Kagey-Sobotka, A, Dembo, M, Goldstein, B, Metzger, H and Lichtenstein, L M (1981) Qualitative characteristics of histamine release from human basophils by covalently cross-linked IgE. The Journal of Immunology 127 (6), 2285-2291
- Published 11 February 2019
- ^ MacNab, R., and D. E. Koshland, Jr. (1972). The gradient-sensing mechanism in bacterial chemotaxis. Proc. Natl. Acad. Sci. U.S.A. 69:2509-2512
- ^ Berg, H C and Purcell, E M (1977) Physics of chemoreception, Biophys. J 20(2):193-219
- ^ Kirsten Jung, Florian Fabiani, Elisabeth Hoyer, and Jürgen Lassak 2018 Bacterial transmembrane signaling systems and their engineering for biosensing Open Biol. Apr; 8(4): 180023
External links
- Netpath - A curated resource of signal transduction pathways in humans Archived 2012-09-20 at the Wayback Machine
- Signal Transduction - The Virtual Library of Biochemistry, Molecular Biology and Cell Biology
- TRANSPATH(R) - A database about signal transduction pathways
- Science's STKE - Signal Transduction Knowledge Environment, from the journal Science, published by AAAS.
- Signal+Transduction at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- UCSD-Nature Signaling Gateway Archived 2013-02-12 at the Wayback Machine, from Nature Publishing Group
- LitInspector Archived 2019-05-11 at the Wayback Machine - Signal transduction pathway mining in PubMed abstracts
- Huaxian Chen, et al. A Cell Based Immunocytochemical Assay For Monitoring Kinase Signaling Pathways And Drug Efficacy (PDF) Archived 2012-02-22 at the Wayback Machine Analytical Biochemistry 338 (2005) 136-142
- www.Redoxsignaling.com
- Signaling PAthway Database Archived 2012-09-17 at the Wayback Machine - Kyushu University
- Cell cycle - Homo sapiens (human) Archived 2012-10-23 at the Wayback Machine - KEGG PATHWAY [1]
- Pathway Interaction Database - NCI
- Literature-curated human signaling network, the largest human signaling network database
| |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| MAP | |||||||||||||||
| Calcium |
| ||||||||||||||
| G protein |
| ||||||||||||||
| Cyclin | |||||||||||||||
| Lipid | |||||||||||||||
| Other protein kinase |
| ||||||||||||||
| Other protein phosphatase |
| ||||||||||||||
| Apoptosis | |||||||||||||||
| GTP-binding protein regulators | |||||||||||||||
| Other |
| ||||||||||||||
| Signaling pathways | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Agents |
| ||||||||||||
| By distance |
| ||||||||||||
| Other concepts |
| ||||||||||||
