Slippery sequence
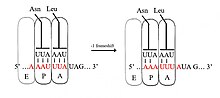
A slippery sequence is a small section of codon
polyproteins.[1]
The frameshift occurs due to wobble pairing. The Gibbs free energy of secondary structures downstream give a hint at how often frameshift happens.[7] Tension on the mRNA molecule also plays a role.[8] A list of slippery sequences found in animal viruses is available from Huang et al.[9]
Slippery sequences that cause a 2-base slip (−2 frameshift) have been constructed out of the HIV UUUUUUA sequence.[8]
See also
- Nucleic acid tertiary structure
- Open reading frame
- Ribosomal frameshifting
- Translational frameshift
- Transposable element
References
- ^ S2CID 25672863.
- PMID 18021801.
- PMID 20693527.
- ^ "Dr Ian Brierley Research description". Department of Pathology, University of Cambridge. Archived from the original on 2013-10-02. Retrieved 2013-07-28.
- ^ "Molecular Biology: Frameshifting occurs at slippery sequences". Molecularstudy.blogspot.com. 2012-10-16. Retrieved 2013-07-28.
- PMID 10075915.
- PMID 18367782.
- ^ PMID 22743270.
- PMID 24298557.
External links
- Pseudobase
- Recode
- Frameshifting,+Ribosomal at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- Wise2 - aligns a frameshifts and introns
- FastY - compare a frameshifts
- Path - tool that compares two translationprinciple)
- Recode2 - Database of recoded genes, including those that require programmed Translational frameshift.
- Page for Coronavirus frameshifting stimulation element at Rfam
