Sphingomyelin phosphodiesterase
| Sphingomyelin phosphodiesterase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
ExPASy NiceZyme view | | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
Sphingomyelin phosphodiesterase (EC 3.1.4.12, also known as neutral sphingomyelinase, sphingomyelinase, or SMase; systematic name sphingomyelin cholinephosphohydrolase) is a
Sphingomyelinase family
Five types of SMase have been identified. These are classified according to their cation dependence and pH optima of action and are:
- Lysosomal acid SMase
- Secreted zinc-dependent acid SMase
- Magnesium-dependent neutral SMase
- Magnesium-independent neutral SMase
- Alkaline SMase
Of these, the lysosomal acidic SMase and the magnesium-dependent neutral SMase are considered major candidates for the production of ceramide in the cellular response to stress.
Neutral sphingomyelinase
Neutral sphingomyelinase (N-SMase) activity was first described in fibroblasts from patients with
A major breakthrough came in the mid-1980s with the cloning of the first N-SMases from Bacillus cereus and Staphylococcus aureus.[6][7] Using the sequences of these bacterial sphingomyelinases in homology searches ultimately led to the identification of the yeast N-SMases ISC1 in the budding yeast Saccharomyces cerevisiae[8] and the mammalian N-SMase enzymes, nSMase1 and nSMase2.[9][10] The identity between mammalian, yeast and bacterial SMases is very low - being approximately 20% between nSMase2 and the B. cereus SMase. However, an alignment of the sequences (see figure) indicate a number of conserved residues throughout the family, particularly in the catalytic region of the enzymes.[11] This has led to the suggestion of a common catalytic mechanism for the N-SMase family.
A third N-SMase protein – termed nSMase3 – was cloned and characterized in 2006.[12] nSMase3 bears little sequence similarity to either nSMase1 or nSMase2. However, there appears to be a high degree of evolutionary conservation from lower to higher organisms, suggesting that it may comprise a unique and distinct N-SMase. The high expression of nSMase3 in heart and skeletal muscle also suggests potential roles in heart function.[13]
Active site

The solving of the crystal structure of the neutral sphingomyelinase from
Mechanism
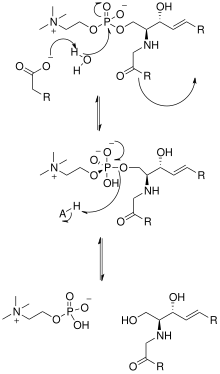
The solving of the crystal structure of the neutral sphingomyelinase from
References
Further reading
External links
- Sphingomyelin+Phosphodiesterase at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
