Structure and genome of HIV
The
Structure
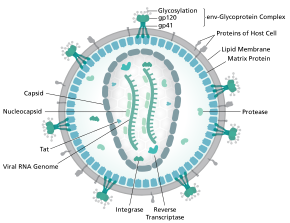


The complete sequence of the HIV-1 genome, extracted from infectious virions, has been solved to single-nucleotide resolution.[6] The HIV genome encodes a small number of
HIV-1 is composed of two copies of
The single-strand RNA is tightly bound to p7
As the only proteins on the surface of the virus, the envelope glycoproteins (gp120 and gp41) are the major targets for
The molecular structure of the
Genome organization

HIV has several major genes coding for structural proteins that are found in all retroviruses as well as several nonstructural ("accessory") genes unique to HIV.[27] The HIV genome contains nine genes that encode fifteen viral proteins.[28] These are synthesized as polyproteins which produce proteins for virion interior, called Gag, group specific antigen; the viral enzymes (Pol, polymerase) or the glycoproteins of the virion env (envelope).[29] In addition to these, HIV encodes for proteins which have certain regulatory and auxiliary functions as well.[29] HIV-1 has two important regulatory elements: Tat and Rev and few important accessory proteins such as Nef, Vpr, Vif and Vpu which are not essential for replication in certain tissues.[29] The gag gene provides the basic physical infrastructure of the virus, and pol provides the basic mechanism by which retroviruses reproduce, while the others help HIV to enter the host cell and enhance its reproduction. Though they may be altered by mutation, all of these genes except tev exist in all known variants of HIV; see Genetic variability of HIV.[citation needed]
HIV employs a sophisticated system of differential RNA splicing to obtain nine different gene products from a less than 10kb genome.[30] HIV has a 9.2kb unspliced genomic transcript which encodes for gag and pol precursors; a singly spliced, 4.5 kb encoding for env, Vif, Vpr and Vpu and a multiply spliced, 2 kb mRNA encoding for Tat, Rev and Nef.[30]
| Class | Gene name | Primary protein products | Processed protein products |
|---|---|---|---|
| Viral structural proteins | gag | Gag polyprotein | MA, CA, SP1, NC, SP2, P6 |
| pol | Pol polyprotein | RT, RNase H, IN, PR | |
| env | gp160 | gp120, gp41 | |
| Essential regulatory elements | tat | Tat | |
| rev | Rev | ||
| Accessory regulatory proteins | nef | Nef | |
| vpr | Vpr | ||
| vif | Vif | ||
| vpu | Vpu |
Viral structural proteins

- nucleocapsid protein, p7); SP2 (spacer peptide 2, p1) and P6 protein.[31]
Essential regulatory elements
- nuclear localization signals), required for the transport of Rev to the nucleus from cytosol during viral replication.[29] Rev recognizes a complex stem-loop structure of the mRNA env located in the intron separating coding exon of Tat and Rev, known as the HIV Rev response element (RRE).[9][29] Rev is important for the synthesis of major viral proteins and is hence essential for viral replication.[citation needed]
Accessory regulatory proteins
- cell apoptosis and increase virus infectivity.[29]
- oligomeric integral membrane phosphoprotein with numerous biological functions. Vpu is involved in CD4 degradation involving the ubiquitin proteasome pathway as well as in the successful release of virions from infected cells.[9][29]
- tev: This gene is only present in a few HIV-1 isolates. It is a fusion of parts of the tat, env, and rev genes, and codes for a protein with some of the properties of rev.[32]
RNA secondary structure
| HIV pol-1 stem loop | |
|---|---|
Cis-reg | |
| PDB structures | PDBe |
Several conserved
An RNA secondary structure determined by
V3 loop
The third variable loop or V3 loop is a part or region of the
See also
References
- S2CID 390173.
- PMID 6601823.
- OCLC 949701914.
- PMID 6265753.
- PMID 6789108. Archived from the original on October 22, 2012. Retrieved September 15, 2017.)
{{cite journal}}: CS1 maint: unfit URL (link - PMID 19661910.
- ^ PMID 27357278.
- OCLC 71223221.
- ^ a b c d e f g Montagnier L (1999). "Human Immunodeficiency Viruses (Retroviridae)". Encyclopedia of Virology (2nd ed.). pp. 763–774.
- ^ PMID 21762803.
- ^ PMID 19529749.
- PMID 11593039.
- S2CID 33055050.
- S2CID 4316242.
- ^ a b Castelli JC, Levy A (2002). "HIV (Human Immunodeficiency Virus)". Encyclopedia of Cancer. Vol. 2 (2nd ed.). pp. 407–415.
- PMID 21762802.
- ^ National Institute of Health (June 17, 1998). "Crystal structure of key HIV protein reveals new prevention, treatment targets" (Press release). Archived from the original on February 19, 2006. Retrieved September 14, 2006.
- PMID 26972002.
- PMID 26105115.
- PMID 26085151.
- PMID 25747313.
- PMID 24179159.
- PMID 24179160.
- PMID 24068931.
- PMID 26051934.
- PMID 26687358.
- ^ ISBN 9780444520739.
- PMID 25808207.
- ^ a b c d e f g h i j k l m Votteler J, Schubert U (2008). "Human Immunodeficiency Viruses: Molecular Biology". Encyclopedia of Virology (3rd ed.). pp. 517–525.
- ^ PMID 1356348.
- ^ PMID 7915889.
- PMID 2186172.
- S2CID 221636459.
- PMID 1738599.
- PMID 11744696.
- PMID 14757051.
- PMID 14517075.
- PMID 18974280.
- ^ "The interactions of the gp120 V3 loop of different HIV-1 strains with the potent anti-HIV human monoclonal antibody 447-52D". Weizmann Institute of Science: Department of Structural Biology. Archived from the original on 2007-07-18. Retrieved 2017-04-18.
- PMID 27178216.
External links
- Rfam entry for HIV pol-1 stem loop
- 3D model of the complete HIV1 virion
- Liu J, Wright ER, Winkler H (2010). "3D Visualization of HIV Virions by Cryoelectron Tomography". Cryo-EM, Part C: Analyses, Interpretation, and Case studies. Methods in Enzymology. Vol. 483. pp. 267–90. PMID 20888479.
