Tandem mass spectrometry

Tandem mass spectrometry, also known as MS/MS or MS2, is a technique in
The
Structure
Typical tandem mass spectrometry instrumentation setups include triple quadrupole mass spectrometers (QqQ), multi-sector mass spectrometer, quadrupole–time of flight (Q-TOF), Fourier transform ion cyclotron resonance mass spectrometers, and hybrid mass spectrometers.
Triple quadrupole mass spectrometer
Triple quadrupole mass spectrometers use the first and third quadrupoles as mass filters. When analytes pass the second quadrupole, the fragmentation proceeds through collision with gas.
Quadrupole–time of flight (Q-TOF)
Q-TOF mass spectrometer combines TOF and quadrupole instruments, which cause high mass accuracy for product ions, accurate quantitation capability, and fragmentation experiment applicability. This is a method of mass spectrometry that ion fragmentation (m/z) ratio determined through a time of flight measurement.
Hybrid mass spectrometer
Hybrid mass spectrometer consists of more than two mass analyzers.
Instrumentation
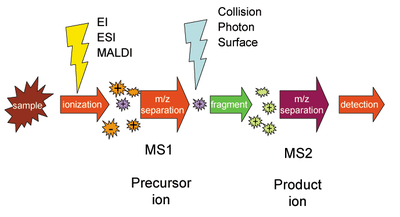
Multiple stages of mass analysis separation can be accomplished with individual mass spectrometer elements separated in space or using a single mass spectrometer with the MS steps separated in time. For tandem mass spectrometry in space, the different elements are often noted in a shorthand, giving the type of mass selector used.
Tandem in space

In tandem mass spectrometry in space, the separation elements are physically separated and distinct, although there is a physical connection between the elements to maintain
Common notation for mass analyzers is Q – quadrupole mass analyzer; q – radio frequency collision quadrupole; TOF – time-of-flight mass analyzer; B – magnetic sector, and E – electric sector. The notation can be combined to indicate various hybrid instrument, for example QqQ' – triple quadrupole mass spectrometer; QTOF – quadrupole time-of-flight mass spectrometer (also QqTOF); and BEBE – four-sector (reverse geometry) mass spectrometer.
Tandem in time

By doing tandem mass spectrometry in time, the separation is accomplished with ions trapped in the same place, with multiple separation steps taking place over time. A
Tandem in space MS/MS modes
When tandem MS is performed with an in space design, the instrument must operate in one of a variety of modes. There are a number of different tandem MS/MS experimental setups and each mode has its own applications and provides different information. Tandem MS in space uses the coupling of two instrument components which measure the same mass spectrum range but with a controlled fractionation between them in space, while tandem MS in time involves the use of an ion trap.
There are four main scan experiments possible using MS/MS: precursor ion scan, product ion scan, neutral loss scan, and selected reaction monitoring.
For a precursor ion scan, the product ion is selected in the second mass analyzer, and the precursor masses are scanned in the first mass analyzer. Note that precursor ion[4] is synonymous with parent ion[5] and product ion[6] with daughter ion;[7] however the use of these anthropomorphic terms is discouraged.[8][9]
In a product ion scan, a precursor ion is selected in the first stage, allowed to fragment and then all resultant masses are scanned in the second mass analyzer and detected in the detector that is positioned after the second mass analyzer. This experiment is commonly performed to identify transitions used for quantification by tandem MS.
In a neutral loss scan, the first mass analyzer scans all the masses. The second mass analyzer also scans, but at a set offset from the first mass analyzer.[10] This offset corresponds to a neutral loss that is commonly observed for the class of compounds. In a constant-neutral-loss scan, all precursors that undergo the loss of a specified common neutral are monitored. To obtain this information, both mass analyzers are scanned simultaneously, but with a mass offset that correlates with the mass of the specified neutral. Similar to the precursor-ion scan, this technique is also useful in the selective identification of closely related class of compounds in a mixture.
In selected reaction monitoring, both mass analyzers are set to a selected mass. This mode is analogous to selected ion monitoring for MS experiments. A selective analysis mode, which can increase sensitivity.[11]
Fragmentation
Fragmentation of gas-phase ions is essential to tandem mass spectrometry and occurs between different stages of mass analysis. There are many methods used to fragment the ions and these can result in different types of fragmentation and thus different information about the structure and composition of the molecule.
In-source fragmentation
Often, the
Collision-induced dissociation
Post-source fragmentation is most often what is being used in a tandem mass spectrometry experiment. Energy can also be added to the ions, which are usually already vibrationally excited, through post-source collisions with neutral atoms or molecules, the absorption of radiation, or the transfer or capture of an electron by a multiply charged ion. Collision-induced dissociation (CID), also called collisionally activated dissociation (CAD), involves the collision of an ion with a neutral atom or molecule in the gas phase and subsequent dissociation of the ion.[16][17] For example, consider
where the ion AB+ collides with the neutral species M and subsequently breaks apart. The details of this process are described by collision theory. Due to different instrumental configuration, two main different types of CID are possible: (i) beam-type (in which precursor ions are fragmented on-the-flight)[18] and (ii) ion trap-type (in which precursor ions are first trapped, and then fragmented).[19][20]
A third and more recent type of CID fragmentation is
Electron capture and transfer methods
The energy released when an electron is transferred to or captured by a multiply charged ion can induce fragmentation.
Electron-capture dissociation
If an electron is added to a multiply charged positive ion, the Coulomb energy is liberated. Adding a free electron is called electron-capture dissociation (ECD),[27] and is represented by
for a multiply protonated molecule M.
Electron-transfer dissociation
Adding an electron through an ion-ion reaction is called electron-transfer dissociation (ETD).[28][29] Similar to electron-capture dissociation, ETD induces fragmentation of cations (e.g. peptides or proteins) by transferring electrons to them. It was invented by Donald F. Hunt, Joshua Coon, John E. P. Syka and Jarrod Marto at the University of Virginia.[30]
ETD does not use free electrons but employs radical anions (e.g. anthracene or azobenzene) for this purpose:
where A is the anion.[31]
ETD cleaves randomly along the peptide backbone (c and z ions) while side chains and modifications such as phosphorylation are left intact. The technique only works well for higher charge state ions (z>2), however relative to
Electron-transfer and higher-energy collision dissociation (EThcD) is a combination ETD and HCD where the peptide precursor is initially subjected to an ion/ion reaction with fluoranthene anions in a linear ion trap, which generates c- and z-ions.[28][33] In the second step HCD all-ion fragmentation is applied to all ETD derived ions to generate b- and y- ions prior to final analysis in the orbitrap analyzer.[21] This method employs dual fragmentation to generate ion- and thus data-rich MS/MS spectra for peptide sequencing and PTM localization.[34]
Negative electron-transfer dissociation
Fragmentation can also occur with a deprotonated species, in which an electron is transferred from the species to an cationic reagent in a negative electron transfer dissociation (NETD):[35]
Following this transfer event, the electron-deficient anion undergoes internal rearrangement and
NETD is compatible with fragmenting peptide and proteins along the backbone at the Cα-C bond. The resulting fragments are usually a•- and x-type product ions.
Electron-detachment dissociation
Electron-detachment dissociation (EDD) is a method for fragmenting anionic species in mass spectrometry.
Charge-transfer dissociation
Reaction between positively charged peptides and cationic reagents,[37] also known as charge transfer dissociation (CTD),[38] has recently been demonstrated as an alternative high-energy fragmentation pathway for low-charge state (1+ or 2+) peptides. The proposed mechanism of CTD using helium cations as the reagent is:
Initial reports are that CTD causes backbone Cα-C bond cleavage of peptides and provides a•- and x-type product ions.
Photodissociation
The energy required for dissociation can be added by photon absorption, resulting in ion photodissociation and represented by
where represents the photon absorbed by the ion. Ultraviolet lasers can be used, but can lead to excessive fragmentation of biomolecules.[39]
Infrared multiphoton dissociation
Blackbody infrared radiative dissociation
Surface-induced dissociation
With surface-induced dissociation (SID), the fragmentation is a result of the collision of an ion with a surface under high vacuum.[43][44] Today, SID is used to fragment a wide range of ions. Years ago, it was only common to use SID on lower mass, singly charged species because ionization methods and mass analyzer technologies weren't advanced enough to properly form, transmit, or characterize ions of high m/z. Over time, self-assembled monolayer surfaces (SAMs) composed of CF3(CF2)10CH2CH2S on gold have been the most prominently used collision surfaces for SID in a tandem spectrometer. SAMs have acted as the most desirable collision targets due to their characteristically large effective masses for the collision of incoming ions. Additionally, these surfaces are composed of rigid fluorocarbon chains, which don't significantly dampen the energy of the projectile ions. The fluorocarbon chains are also beneficial because of their ability to resist facile electron transfer from the metal surface to the incoming ions.[45] SID's ability to produce subcomplexes that remain stable and provide valuable information on connectivity is unmatched by any other dissociation technique. Since the complexes produced from SID are stable and retain distribution of charge on the fragment, this produces a unique, spectra which the complex centers around a narrower m/z distribution. The SID products and the energy at which they form are reflective of the strengths and topology of the complex. The unique dissociation patterns help discover the Quaternary structure of the complex. The symmetric charge distribution and dissociation dependence are unique to SID and make the spectra produced distinctive from any other dissociation technique.[45]
The SID technique is also applicable to ion-mobility mass spectrometry (IM-MS). Three different methods for this technique include analyzing the characterization of topology, intersubunit connectivity, and the degree of unfolding for protein structure. Analysis of protein structure unfolding is the most commonly used application of the SID technique. For Ion-mobility mass spectrometry (IM-MS), SID is used for dissociation of the source activated precursors of three different types of protein complexes: C-reactive protein (CRP), transthyretin (TTR), and concanavalin A (Con A). This method is used to observe the unfolding degree for each of these complexes. For this observation, SID showed the precursor ions' structures that exist before the collision with the surface. IM-MS utilizes the SID as a direct measure of the conformation for each proteins' subunit.[46]
Fourier-transform ion cyclotron resonance (FTICR) are able to provide ultrahigh resolution and high mass accuracy to instruments that take mass measurements. These features make FTICR mass spectrometers a useful tool for a wide variety of applications such as several dissociation experiments[47] such as collision-induced dissociation (CID, electron transfer dissociation (ETD),[48] and others. In addition, surface-induced dissociation has been implemented with this instrument for the study of fundamental peptide fragmentation. Specifically, SID has been applied to the study of energetics and the kinetics of gas-phase fragmentation within an ICR instrument.[49] This approach has been used to understand the gas-phase fragmentation of protonated peptides, odd-electron peptide ions, non-covalent ligand-peptide complexes, and ligated metal clusters.
Quantitative proteomics
Quantitative proteomics is used to determine the relative or absolute amount of proteins in a sample.[50][51][52] Several quantitative proteomics methods are based on tandem mass spectrometry. MS/MS has become a benchmark procedure for the structural elucidation of complex biomolecules.[53]
One method commonly used for quantitative proteomics is isobaric tag labeling. Isobaric tag labeling enables simultaneous identification and quantification of proteins from multiple samples in a single analysis. To quantify proteins,
Isobaric tags for relative and absolute quantitation (iTRAQ)
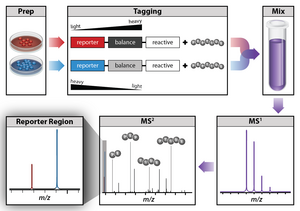
An isobaric tag for relative and absolute quantitation (iTRAQ) is a reagent for tandem mass spectrometry that is used to determine the amount of proteins from different sources in a single experiment.[54][55][56] It uses stable
Tandem mass tag (TMT)
A tandem mass tag (TMT) is an isobaric mass tag chemical label used for protein quantification and identification.[57] The tags contain four regions: mass reporter, cleavable linker, mass normalization, and protein reactive group. TMT reagents can be used to simultaneously analyze 2 to 11 different peptide samples prepared from cells, tissues or biological fluids. Recent developments allow up to 16 and even 18 samples (16plex or 18plex respectively) to be analyzed.[58][59] Three types of TMT reagents are available with different chemical reactivities: (1) a reactive NHS ester functional group for labeling primary amines (TMTduplex, TMTsixplex, TMT10plex plus TMT11-131C), (2) a reactive iodoacetyl functional group for labeling free sulfhydryls (iodoTMT) and (3) reactive alkoxyamine functional group for labeling of carbonyls (aminoxyTMT).
Multiplexed DIA (plexDIA)
The progress in data independent acquisition (DIA) enabled multiplexed quantitative proteomics with non-isobaric mass tags and a new method called plexDIA introduced in 2021.[60][61] This new approach increases the number of data points by parallelizing both samples and peptides, thus achieving multiplicative gains. It has the potential to continue scaling proteomic throughput with new mass tags and algorithms.[62][63] plexDIA is applicable to both bulk[64] and single-cell samples and is particularly powerful for single-cell proteomics.[65][66][67]
Applications
Peptides

Tandem mass spectrometry can be used for
Tandem mass spectrometry can produce a peptide sequence tag that can be used to identify a peptide in a protein database.[69][70][71] A notation has been developed for indicating peptide fragments that arise from a tandem mass spectrum.[72] Peptide fragment ions are indicated by a, b, or c if the charge is retained on the N-terminus and by x, y or z if the charge is maintained on the C-terminus. The subscript indicates the number of amino acid residues in the fragment. Superscripts are sometimes used to indicate neutral losses in addition to the backbone fragmentation, * for loss of ammonia and ° for loss of water. Although peptide backbone cleavage is the most useful for sequencing and peptide identification other fragment ions may be observed under high energy dissociation conditions. These include the side chain loss ions d, v, w and ammonium ions[73][74] and additional sequence-specific fragment ions associated with particular amino acid residues.[75]
Oligosaccharides
Oligosaccharides may be sequenced using tandem mass spectrometry in a similar manner to peptide sequencing.[76] Fragmentation generally occurs on either side of the glycosidic bond (b, c, y and z ions) but also under more energetic conditions through the sugar ring structure in a cross-ring cleavage (x ions). Again trailing subscripts are used to indicate position of the cleavage along the chain. For cross ring cleavage ions the nature of the cross ring cleavage is indicated by preceding superscripts.[77][78]
Oligonucleotides
Tandem mass spectrometry has been applied to
Newborn screening
Newborn screening is the process of testing newborn babies for treatable
Small molecule analysis
It has been shown that tandem mass spectrometry data is highly consistent across instrument and manufacturer platforms including quadrupole time-of-flight (QTOF) and Q Exactive instrumentation, especially at 20 eV.[85]
Limitation
Tandem mass spectrometry cannot be applied for single-cell analyses as it is insensitive to analyze such small amounts of a cell. These limitations are primarily due to a combination of inefficient ion production and ion losses within the instruments due to chemical noise sources of solvents.[86]
Future outlook
Tandem mass spectrometry will be a useful tool for protein characterization, nucleoprotein complexes, and other biological structures. However, some challenges left such as analyzing the characterization of the proteome quantitatively and qualitatively.[87]
See also
- Accelerator mass spectrometry
- Cross section (physics)
- Mass-analyzed ion-kinetic-energy spectrometry
- Unimolecular ion decomposition
References
- .
- .
- .
- PMID 24243061.
- .
- ISBN 978-0-471-48566-7.
- PMID 30681830.
- S2CID 214768212.
- S2CID 1030006.
- PMID 16401509.
- ^ PMID 15481084.
- PMID 16478115.
- S2CID 16506573.
- PMID 18511480.
- ^ S2CID 2538231.
- PMID 24251866.
- PMID 28608681.
- PMID 20873877.
- PMID 29949056.
- )
- PMID 15389856.
- ^ PMID 15210983.
- PMID 17118725.
- ^ US patent 7534622, Donald F. Hunt, Joshua J. Coon, John E.P. Syka, Jarrod A. Marto, "Electron transfer dissociation for biopolymer sequence mass spectrometric analysis", issued 2009-05-19
- PMID 10360331.
- PMID 17287358.
- PMID 23106539.
- PMID 23347405.
- PMID 15907703.
- .
- PMID 24236851.
- S2CID 1400057.
- PMID 16401510.
- PMID 7526742.
- ^ PMID 16525512.
- PMID 14732935.
- .
- .
- ^ PMID 30412666.
- PMID 26499904.
- PMID 15389858.
- PMID 18181247.
- S2CID 19837927.
- S2CID 32054251.
- PMID 17668192.
- S2CID 33009117.
- .
- S2CID 1979097.
- PMID 16574745.
- PMID 17162667.
- PMID 12713048.
- PMID 32203386.
- PMID 33900084.
- , retrieved 14 April 2024
- PMID 35835881.
- PMID 36735898.
- PMID 34808355.
- PMID 37695820.
- PMID 34597050.
- PMID 36864200.
- PMID 38076795, retrieved 14 April 2024
- PMID 22498958.
- PMID 17492750.
- S2CID 38068737.
- PMID 8710858.
- PMID 6525415.
- .
- S2CID 38931760.
- .
- PMID 14966796.
- S2CID 206787925.
- PMID 10973004.
- PMID 15884792.
- PMID 16483781.
- .
- PMID 17679658.
- S2CID 28372085.
- PMID 14578311.
- PMID 38529642.
- PMID 22498958.
- PMID 18718552.
Bibliography
- ISBN 978-0-89573-275-0.
- ISBN 978-0-471-18699-1.
- ISBN 978-0-471-86597-1.
- Sherman NE, Kinter M (2000). Protein sequencing and identification using tandem mass spectrometry. New York: John Wiley. ISBN 978-0-471-32249-8.
External links
- An Introduction to Mass Spectrometry by Dr Alison E. Ashcroft Archived 8 August 2020 at the Wayback Machine


![{\displaystyle [{\ce {M}}+n{\ce {H}}]^{n+}+{\ce {e^{-}->}}\left[[{\ce {M}}+(n-1){\ce {H}}]^{(n-1)+}\right]^{*}{\ce {->fragments}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/80535d501352b7854827d81bd70a7811d203d639)
![{\displaystyle [{\ce {M}}+n{\ce {H}}]^{n+}+{\ce {A^{-}->}}\left[[{\ce {M}}+(n-1){\ce {H}}]^{(n-1)+}\right]^{*}+{\ce {A->fragments}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/a0f39c0e76fd3f9d3862b8b49cb144a43c1cd907)
![{\displaystyle [{\ce {M}}-n{\ce {H}}]^{n-}+{\ce {A+->}}\left[[{\ce {M}}-n{\ce {H}}]^{(n+1)-}\right]^{*}+{\ce {A->fragments}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/383244602d9ce4607f68c0696b8c55fb10951a3f)
![{\displaystyle {\ce {{[{M}+H]^{1}+}+He+->}}\left[{\ce {[{M}+H]^{2}+}}\right]^{*}+{\ce {He^{0}->fragments}}}](https://wikimedia.org/api/rest_v1/media/math/render/svg/a8ccd097f31299c546cd6e591efb6ed4f8556e2e)

