Thymidylate synthase
Ensembl | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| UniProt | |||||||||
| RefSeq (mRNA) | |||||||||
| RefSeq (protein) | |||||||||
| Location (UCSC) | Chr 18: 0.66 – 0.67 Mb | Chr 5: 30.26 – 30.28 Mb | |||||||
| PubMed search | [3] | [4] | |||||||
| View/Edit Human | View/Edit Mouse |
| thymidylate synthase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
ExPASy NiceZyme view | | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| Thymidylate synthase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | Thymidylat_synt | ||||||||
SCOP2 | 1tys / SCOPe / SUPFAM | ||||||||
| |||||||||
Thymidylate synthase (TS) (
Function
The following reaction is catalyzed by thymidylate synthase:
- 5,10-methylenetetrahydrofolate + dUMP dihydrofolate + dTMP
By means of reductive
This provides the sole de novo pathway for production of dTMP and is the only enzyme in folate metabolism in which the 5,10-methylenetetrahydrofolate is oxidised during one-carbon transfer.[8] The enzyme is essential for regulating the balanced supply of the 4 DNA precursors in normal DNA replication: defects in the enzyme activity affecting the regulation process cause various biological and genetic abnormalities, such as thymineless death.[9] The enzyme is an important target for certain chemotherapeutic drugs. Thymidylate synthase is an enzyme of about 30 to 35 kDa in most species except in protozoan and plants where it exists as a bifunctional enzyme that includes a dihydrofolate reductase domain.[8] A cysteine residue is involved in the catalytic mechanism (it covalently binds the 5,6-dihydro-dUMP intermediate). The sequence around the active site of this enzyme is conserved from phages to vertebrates.
Thymidylate synthase is induced by a transcription factor LSF/TFCP2 and LSF is an oncogene in hepatocellular carcinoma. LSF and Thymidylate synthase plays significant role in Liver Cancer proliferation and progression and Drug resistance.[10]
Clinical significance
Thymidylate synthase (TS) plays a crucial role in the early stages of
Using TS as a drug target
The use of
Experimentally, it has been shown that low levels of TS expression leads to a better response to 5-FU and higher success rates and survival of colon and liver cancer patients.
TS's relation to the cell cycle also contributes to its use in cancer treatment. Several cell-cycle dependent
Interactive pathway map
Click on genes, proteins and metabolites below to link to respective articles.[§ 1]
- ^ The interactive pathway map can be edited at WikiPathways: "FluoropyrimidineActivity_WP1601".
Mechanism description

In the proposed mechanism, TS forms a covalent bond to the substrate dUMP through a 1,4-addition involving a cysteine nucleophile. The substrate tetrahydrofolate donates a methyl group to the alpha carbon while reducing the new methyl on dUMP to form dTMP.[15]

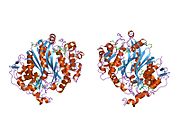
It has been proven that the imine formed through reaction with THF and dUMP is an intermediate in the reaction with dUMP through mutations in the structure of TS that inhibit the completion of the mechanism. V316Am TS, a mutant with deletion of C terminal valines from both subunits, allows the catalysis of dehalogenation of BrdUMP preceding the mechanism described above and the covalent bond to THF and dUMP. The mutant TS is unable to accomplish the C-terminal conformational change needed to break covalent bonds to form dTMP, thus showing the proposed mechanism to be true. The structure was deduced through x-ray crystallography of V316Am TS to illustrate full homodimer TS structure (Figure 1). In addition, it showed possible interactions of the 175Arg and 174Arg between dimers. These arginines are thought to stabilize the UMP structures within the active sites by creating hydrogen bonds to the phosphate group (Figure 2). [Stroud and Finer-Moore][citation needed] 5-FU is an inhibitor of TS. Upon entering the cell, 5-fluorouracil (5-FU) is converted to a variety of active metabolites, intracellularly. One such metabolite is FdUMP which differs from dUMP by a fluorine in place of a hydrogen on the alpha carbon. FdUMP is able to inhibit TS by binding to the nucleotide-binding site of dUMP. This competitive binding inhibits the normal function of dTMP synthesis from dUMP [Longley].[citation needed] Thus the dUMP is unable to have an elimination reaction and complete the methyl donation from THF.

Figure 1. This figure depicts the homodimer that is TS. As you can see the orange and teal backbones never connect or intertwine, but there are side chains interactions between the dimers. On the orange protein, you can visibly detect two long side chains that enter the teal protein (this is located within the yellow circle). The other beige parts are side chains that interact within the active site. Just below the yellow circle, you are able to see the same pattern of side chains and configuration.

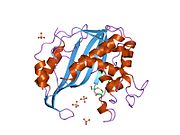
Figure 2. This figure shows the possible H-bond interactions between the arginines and the UMP in the active site of thymidylate synthase. This can be seen by the faint lines between the blue tips and the red tips. These arginines are used to hold the position of the UMP molecule so that the interaction may occur correctly. The two arginines in the top right corner that are located next to each other on the back bone are actually from the other protein of this dimer enzyme. This interaction is one of the many intermolecular forces that holds these two tertiary structures together. The yellow stand in the top-middle region shows a sulfur bond that forms between a cysteine side chain and UMP. This covalently holds the UMP within the active site until it is reacted to yield TMP.
See also
- Pyrimidine analogues
- Thymidylate synthase inhibitor
- Thymidine kinase
- Thymidine kinase in clinical chemistry
- Thymidylate kinase
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000176890 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000025747 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Entrez Gene: TYMS thymidylate synthetase".
- ^ "DNA: Form and Function" (PDF). Archived from the original (PDF) on 2013-05-25. Retrieved 2014-05-02.
- ^ "DNA Synthesis".
- ^ PMID 3099389.
- PMID 2243092.
- PMID 22679558.
- ^ PMID 12084461.
- ^ "Leucovorin". MedlinePlus Drug Information. U.S. National Library of Medicine.
- ^ PMID 10631692.
- PMID 21622151.
- PMID 7574499.
Further reading
- Carreras CW, Santi DV (1995). "The Catalytic Mechanism and Structure of Thymidylate Synthase". Annual Review of Biochemistry. 64 (1): 721–762. PMID 7574499.
- Banerjee D, Mayer-Kuckuk P, Capiaux G, et al. (2002). "Novel aspects of resistance to drugs targeted to dihydrofolate reductase and thymidylate synthase". Biochim. Biophys. Acta. 1587 (2–3): 164–73. PMID 12084458.
- Liu J, Schmitz JC, Lin X, et al. (2002). "Thymidylate synthase as a translational regulator of cellular gene expression". Biochim. Biophys. Acta. 1587 (2–3): 174–82. PMID 12084459.
- Chu J, Dolnick BJ (2002). "Natural antisense (rTSalpha) RNA induces site-specific cleavage of thymidylate synthase mRNA". Biochim. Biophys. Acta. 1587 (2–3): 183–93. PMID 12084460.
- Peters GJ, Backus HH, Freemantle S, et al. (2002). "Induction of thymidylate synthase as a 5-fluorouracil resistance mechanism". Biochim. Biophys. Acta. 1587 (2–3): 194–205. PMID 12084461.
- Costi MP, Tondi D, Rinaldi M, et al. (2002). "Structure-based studies on species-specific inhibition of thymidylate synthase". Biochim. Biophys. Acta. 1587 (2–3): 206–14. S2CID 19676651.
- Lin D, Li H, Tan W, et al. (2007). Genetic polymorphisms in folate- metabolizing enzymes and risk of gastroesophageal cancers: a potential nutrient-gene interaction in cancer development. Forum of Nutrition. Vol. 60. pp. 140–5. PMID 17684410.
External links
- Thymidylate+synthetase at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- PDBe-KB provides an overview of all the structure information available in the PDB for Human Thymidylate synthase