Ubiquitin
| Ubiquitin family | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| |||||||||
Ubiquitin is a small (8.6
The addition of ubiquitin to a substrate protein is called ubiquitylation (or ubiquitination or ubiquitinylation). Ubiquitylation affects proteins in many ways: it can mark them for
The protein modifications can be either a single ubiquitin protein (monoubiquitylation) or a chain of ubiquitin (polyubiquitylation). Secondary ubiquitin molecules are always linked to one of the seven
The discovery that ubiquitin chains target proteins to the proteasome, which degrades and recycles proteins, was honored with the Nobel Prize in Chemistry in 2004.[8][11][12]
Identification

Ubiquitin (originally, ubiquitous immunopoietic polypeptide) was first identified in 1975
The ubiquitylation system was initially characterised as an
The protein
| Number of residues | 76 |
|---|---|
| Molecular mass | 8564.8448 Da |
| Isoelectric point (pI) | 6.79 |
| Gene names | |
| Sequence (single-letter) | MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPD QQRLIFAGKQLEDGRTLSDYNIQKESTLHLVLRLRGG |
Ubiquitin is a small
Genes
Ubiquitin is encoded in mammals by 4 different genes.
Ubiquitylation
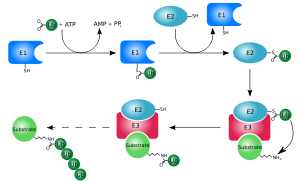
Ubiquitylation (also known as ubiquitination or ubiquitinylation) is an enzymatic
Ubiquitination requires three types of enzyme: ubiquitin-activating enzymes, ubiquitin-conjugating enzymes, and ubiquitin ligases, known as E1s, E2s, and E3s, respectively. The process consists of three main steps:
- Activation: Ubiquitin is activated in a two-step reaction by an E1 ubiquitin-activating enzyme, which is dependent on ATP. The initial step involves production of a ubiquitin-adenylate intermediate. The E1 binds both ATP and ubiquitin and catalyses the acyl-adenylation of the C-terminus of the ubiquitin molecule. The second step transfers ubiquitin to an active site cysteine residue, with release of AMP. This step results in a thioester linkage between the C-terminal carboxyl group of ubiquitin and the E1 cysteine sulfhydryl group.[15][53] The human genome contains two genes that produce enzymes capable of activating ubiquitin: UBA1 and UBA6.[54]
- Conjugation: E2 eukaryotic organisms have between 16 and 35. They are characterised by their highly conserved structure, known as the ubiquitin-conjugating catalytic (UBC) fold.[55]

Glycine and lysine linked by an isopeptide bond. The isopeptide bond is highlighted yellow. - Ligation: E3 substrate recognition modules of the system and are capable of interaction with both E2 and substrate. Some E3 enzymes also activate the E2 enzymes. E3 enzymes possess one of two domains: the homologous to the E6-AP carboxyl terminus (HECT) domain and the really interesting new gene (RING) domain (or the closely related U-box domain). HECT domain E3s transiently bind ubiquitin in this process (an obligate thioester intermediate is formed with the active-site cysteine of the E3), whereas RING domain E3s catalyse the direct transfer from the E2 enzyme to the substrate.[56] The anaphase-promoting complex (APC) and the SCF complex (for Skp1-Cullin-F-box protein complex) are two examples of multi-subunit E3s involved in recognition and ubiquitylation of specific target proteins for degradation by the proteasome.[57]
In the ubiquitylation cascade, E1 can bind with many E2s, which can bind with hundreds of E3s in a hierarchical way. Having levels within the cascade allows tight regulation of the ubiquitylation machinery.[7] Other ubiquitin-like proteins (UBLs) are also modified via the E1–E2–E3 cascade, although variations in these systems do exist.[58]
E4 enzymes, or ubiquitin-chain elongation factors, are capable of adding pre-formed polyubiquitin chains to substrate proteins.[59] For example, multiple monoubiquitylation of the tumor suppressor p53 by Mdm2[60] can be followed by addition of a polyubiquitin chain using p300 and CBP.[61][62]
Types
Ubiquitylation affects cellular process by regulating the degradation of proteins (via the proteasome and lysosome), coordinating the cellular localization of proteins, activating and inactivating proteins, and modulating protein–protein interactions.[4][5][6] These effects are mediated by different types of substrate ubiquitylation, for example the addition of a single ubiquitin molecule (monoubiquitylation) or different types of ubiquitin chains (polyubiquitylation).[63]
Monoubiquitylation
Monoubiquitylation is the addition of one ubiquitin molecule to one substrate protein residue. Multi-monoubiquitylation is the addition of one ubiquitin molecule to multiple substrate residues. The monoubiquitylation of a protein can have different effects to the polyubiquitylation of the same protein. The addition of a single ubiquitin molecule is thought to be required prior to the formation of polyubiquitin chains.
Polyubiquitin chains


Polyubiquitylation is the formation of a ubiquitin chain on a single lysine residue on the substrate protein. Following addition of a single ubiquitin moiety to a protein substrate, further ubiquitin molecules can be added to the first, yielding a polyubiquitin chain.[63] These chains are made by linking the glycine residue of a ubiquitin molecule to a lysine of ubiquitin bound to a substrate. Ubiquitin has seven lysine residues and an N-terminus that serves as points of ubiquitination; they are K6, K11, K27, K29, K33, K48, K63 and M1, respectively.[8] Lysine 48-linked chains were the first identified and are the best-characterised type of ubiquitin chain. K63 chains have also been well-characterised, whereas the function of other lysine chains, mixed chains, branched chains, M1-linked linear chains, and heterologous chains (mixtures of ubiquitin and other ubiquitin-like proteins) remains more unclear.[17][39][63][64][65]
Lysine 48-linked polyubiquitin chains target proteins for destruction, by a process known as proteolysis. Multi-ubiquitin chains at least four ubiquitin molecules long must be attached to a lysine residue on the condemned protein in order for it to be recognised by the 26S proteasome.[66] This is a barrel-shape structure comprising a central proteolytic core made of four ring structures, flanked by two cylinders that selectively allow entry of ubiquitylated proteins. Once inside, the proteins are rapidly degraded into small peptides (usually 3–25 amino acid residues in length). Ubiquitin molecules are cleaved off the protein immediately prior to destruction and are recycled for further use.[67] Although the majority of protein substrates are ubiquitylated, there are examples of non-ubiquitylated proteins targeted to the proteasome.[68] The polyubiquitin chains are recognised by a subunit of the proteasome: S5a/Rpn10. This is achieved by a ubiquitin-interacting motif (UIM) found in a hydrophobic patch in the C-terminal region of the S5a/Rpn10 unit.[4]
Lysine 63-linked chains are not associated with proteasomal degradation of the substrate protein. Instead, they allow the coordination of other processes such as endocytic trafficking, inflammation, translation, and DNA repair.[10] In cells, lysine 63-linked chains are bound by the ESCRT-0 complex, which prevents their binding to the proteasome. This complex contains two proteins, Hrs and STAM1, that contain a UIM, which allows it to bind to lysine 63-linked chains.[69][70]
Methionine 1-linked (or linear) polyubiquitin chains are another type of non-degradative ubiquitin chains. In this case, ubiquitin is linked in a head-to-tail manner, meaning that the C-terminus of the last ubiquitin molecule binds directly to the N-terminus of the next one. Although initially believed to target proteins for proteasomal degradation,[71] linear ubiquitin later proved to be indispensable for NF-kB signaling.[72] Currently, there is only one known E3 ubiquitin ligase generating M1-linked polyubiquitin chains - linear ubiquitin chain assembly complex (LUBAC).[39][73]
Less is understood about atypical (non-lysine 48-linked) ubiquitin chains but research is starting to suggest roles for these chains.[64] There is evidence that atypical chains linked by lysine 6, 11, 27, 29 and methionine 1 can induce proteasomal degradation.[68][74]
Branched ubiquitin chains containing multiple linkage types can be formed.[75] The function of these chains is unknown.[8]
Structure
Differently linked chains have specific effects on the protein to which they are attached, caused by differences in the conformations of the protein chains. K29-, K33-,[76] K63- and M1-linked chains have a fairly linear conformation; they are known as open-conformation chains. K6-, K11-, and K48-linked chains form closed conformations. The ubiquitin molecules in open-conformation chains do not interact with each other, except for the covalent isopeptide bonds linking them together. In contrast, the closed conformation chains have interfaces with interacting residues. Altering the chain conformations exposes and conceals different parts of the ubiquitin protein, and the different linkages are recognized by proteins that are specific for the unique topologies that are intrinsic to the linkage. Proteins can specifically bind to ubiquitin via ubiquitin-binding domains (UBDs). The distances between individual ubiquitin units in chains differ between lysine 63- and 48-linked chains. The UBDs exploit this by having small spacers between ubiquitin-interacting motifs that bind lysine 48-linked chains (compact ubiquitin chains) and larger spacers for lysine 63-linked chains. The machinery involved in recognising polyubiquitin chains can also differentiate between K63-linked chains and M1-linked chains, demonstrated by the fact that the latter can induce proteasomal degradation of the substrate.[8][10][74]
Function
The ubiquitylation system functions in a wide variety of cellular processes, including:[77]
- Antigen processing
- Apoptosis
- Biogenesisof organelles
- Cell cycle and division
- DNA transcriptionand repair
- Differentiation and development
- Immune response and inflammation
- Neural and muscular degeneration
- Maintenance of pluripotency[78]
- Morphogenesis of neural networks
- Modulation of cell surface receptors, ion channels and the secretory pathway
- Response to stress and extracellular modulators
- Ribosome biogenesis
- Viral infection
Membrane proteins
Multi-monoubiquitylation can mark transmembrane proteins (for example, receptors) for removal from membranes (internalisation) and fulfil several signalling roles within the cell. When cell-surface transmembrane molecules are tagged with ubiquitin, the subcellular localization of the protein is altered, often targeting the protein for destruction in lysosomes. This serves as a negative feedback mechanism, because often the stimulation of receptors by ligands increases their rate of ubiquitylation and internalisation. Like monoubiquitylation, lysine 63-linked polyubiquitin chains also has a role in the trafficking some membrane proteins.[10][63][66][79]
Genomic maintenance
Ubiquitylation of histone H2AX is involved in DNA damage recognition of DNA double-strand breaks. Lysine 63-linked polyubiquitin chains are formed on H2AX histone by the E2/E3 ligase pair, Ubc13-Mms2/RNF168.[82][83] This K63 chain appears to recruit RAP80, which contains a UIM, and RAP80 then helps localize BRCA1. This pathway will eventually recruit the necessary proteins for homologous recombination repair.[84]
Transcriptional regulation
Deubiquitination
Ubiquitin-binding domains
| Domain | Number of proteins
in proteome |
Length
(amino acids) |
Ubiquitin binding
Affinity |
|---|---|---|---|
| CUE | S. cerevisiae: 7
H. sapiens: 21 |
42–43 | ~2–160 μM |
| GATII | S. cerevisiae: 2
H. sapiens: 14 |
135 | ~180 μM |
| GLUE | S. cerevisiae: ?
H. sapiens: ? |
~135 | ~460 μM |
| NZF | S. cerevisiae: 1
H. sapiens: 25 |
~35 | ~100–400 μM |
| PAZ | S. cerevisiae: 5
H. sapiens: 16 |
~58 | Not known |
| UBA | S. cerevisiae: 10
H. sapiens: 98 |
45–55 | ~0.03–500 μM |
| UEV | S. cerevisiae: 2
H. sapiens: ? |
~145 | ~100–500 μM |
| UIM | S. cerevisiae: 8
H. sapiens: 71 |
~20 | ~100–400 μM |
| VHS | S. cerevisiae: 4
H. sapiens: 28 |
150 | Not known |
Ubiquitin-binding domains (UBDs) are modular protein domains that non-covalently bind to ubiquitin, these motifs control various cellular events. Detailed molecular structures are known for a number of UBDs, binding specificity determines their mechanism of action and regulation, and how it regulates cellular proteins and processes.[89][90]
Disease associations
Pathogenesis
The ubiquitin pathway has been implicated in the pathogenesis of a wide range of diseases and disorders, including:[91]
- Neurodegeneration
- Infection and immunity
- Genetic disorders
- Cancer
Neurodegeneration
Ubiquitin is implicated in neurodegenerative diseases associated with proteostasis dysfunction, including
Infection and immunity
Ubiquitin and ubiquitin-like molecules extensively regulate immune signal transduction pathways at virtually all stages, including steady-state repression, activation during infection, and attenuation upon clearance. Without this regulation, immune activation against pathogens may be defective, resulting in chronic disease or death. Alternatively, the immune system may become hyperactivated and organs and tissues may be subjected to autoimmune damage.
On the other hand, viruses must block or redirect host cell processes including immunity to effectively replicate, yet many viruses relevant to disease have informationally limited genomes. Because of its very large number of roles in the cell, manipulating the ubiquitin system represents an efficient way for such viruses to block, subvert or redirect critical host cell processes to support their own replication.[95]
The retinoic acid-inducible gene I (RIG-I) protein is a primary immune system sensor for viral and other invasive RNA in human cells.[96] The RIG-I-like receptor (RLR) immune signaling pathway is one of the most extensively studied in terms of the role of ubiquitin in immune regulation.[97]
Genetic disorders
- Angelman syndrome is caused by a disruption of UBE3A, which encodes a ubiquitin ligase (E3) enzyme termed E6-AP.
- Von Hippel–Lindau syndrome involves disruption of a ubiquitin E3 ligase termed the VHL tumor suppressor, or VHLgene.
- Fanconi anemia: Eight of the thirteen identified genes whose disruption can cause this disease encode proteins that form a large ubiquitin ligase (E3) complex.
- 3-M syndrome is an autosomal-recessive growth retardation disorder associated with mutations of the Cullin7 E3 ubiquitin ligase.[98]
Diagnostic use
- Neurofibrillary tangles in Alzheimer's disease
- Lewy body in Parkinson's disease
- Pick's disease
- Inclusions in motor neuron disease and Huntington's disease
- Mallory bodies in alcoholic liver disease
- Rosenthal fibers in astrocytes
Link to cancer
Direct loss of function mutation of E3 ubiquitin ligase
Renal cell carcinoma
The VHL (
Breast cancer
The BRCA1 gene is another tumor suppressor gene in humans which encodes the BRCA1 protein that is involved in response to DNA damage. The protein contains a RING motif with E3 Ubiquitin Ligase activity. BRCA1 could form dimer with other molecules, such as BARD1 and BAP1, for its ubiquitylation activity. Mutations that affect the ligase function are often found and associated with various cancers.[110][114]
Cyclin E
As processes in cell cycle progression are the most fundamental processes for cellular growth and differentiation, and are the most common to be altered in human carcinomas, it is expected for cell cycle-regulatory proteins to be under tight regulation. The level of cyclins, as the name suggests, is high only at certain a time point during the cell cycle. This is achieved by continuous control of cyclins or CDKs levels through ubiquitylation and degradation. When cyclin E is partnered with CDK2 and gets phosphorylated, an SCF-associated F-box protein Fbw7 recognizes the complex and thus targets it for degradation. Mutations in Fbw7 have been found in more than 30% of human tumors, characterizing it as a tumor suppressor protein.[113]
Increased ubiquitination activity
Cervical cancer
Oncogenic types of the
p53 regulation
Gene amplification often occur in various tumor cases, including of MDM2, a gene encodes for a RING E3 Ubiquitin ligase responsible for downregulation of p53 activity. MDM2 targets p53 for ubiquitylation and proteasomal degradation thus keeping its level appropriate for normal cell condition. Overexpression of MDM2 causes loss of p53 activity and therefore allowing cells to have a limitless replicative potential.[109][114]
p27
Another gene that is a target of gene amplification is SKP2. SKP2 is an F-box protein with a role in substrate recognition for ubiquitylation and degradation. SKP2 targets p27Kip-1, an inhibitor of cyclin-dependent kinases (CDKs). CDKs2/4 partner with the cyclins E/D, respectively, forming a family of cell cycle regulators which control cell cycle progression through the G1 phase. Low level of p27Kip-1 protein is often found in various cancers and is due to overactivation of ubiquitin-mediated proteolysis through overexpression of SKP2.[111][114]
Efp
Efp, or estrogen-inducible RING-finger protein, is an E3 ubiquitin ligase whose overexpression has been shown to be the major cause of estrogen-independent breast cancer.[105][115] Efp's substrate is 14-3-3 protein which negatively regulates cell cycle.
Evasion of ubiquitination
Colorectal cancer
The gene associated with
Glioblastoma
As the most aggressive cancer originated in the brain, mutations found in patients with glioblastoma are related to the deletion of a part of the extracellular domain of the epidermal growth factor receptor (EGFR). This deletion causes CBL E3 ligase unable to bind to the receptor for its recycling and degradation via a ubiquitin-lysosomal pathway. Thus, EGFR is constitutively active in the cell membrane and activates its downstream effectors that are involved in cell proliferation and migration.[112]
Phosphorylation-dependent ubiquitylation
The interplay between ubiquitylation and phosphorylation has been an ongoing research interest since phosphorylation often serves as a marker where ubiquitylation leads to degradation.[99] Moreover, ubiquitylation can also act to turn on/off the kinase activity of a protein.[116] The critical role of phosphorylation is largely underscored in the activation and removal of autoinhibition in the Cbl protein.[117] Cbl is an E3 ubiquitin ligase with a RING finger domain that interacts with its tyrosine kinase binding (TKB) domain, preventing interaction of the RING domain with an E2 ubiquitin-conjugating enzyme. This intramolecular interaction is an autoinhibition regulation that prevents its role as a negative regulator of various growth factors and tyrosine kinase signaling and T-cell activation.[117] Phosphorylation of Y363 relieves the autoinhibition and enhances binding to E2.[117] Mutations that render the Cbl protein dysfunctional due to the loss of its ligase/tumor suppressor function and maintenance of its positive signaling/oncogenic function have been shown to cause the development of cancer.[118][119]
As a drug target
Screening for ubiquitin ligase substrates
Deregulation of E3-substrate interactions is a key cause of many human disorders, therefore identifying E3 ligase substrates is crucial. In 2008, 'Global Protein Stability (GPS) Profiling' was developed to discover E3 ubiquitin ligase substrates.[120] This high-throughput system made use of reporter proteins fused with thousands of potential substrates independently. By inhibition of the ligase activity (through the making of Cul1 dominant negative thus renders ubiquitination not to occur), increased reporter activity shows that the identified substrates are being accumulated. This approach added a large number of new substrates to the list of E3 ligase substrates.
Possible therapeutic applications
Blocking of specific substrate recognition by the E3 ligases, e.g. bortezomib.[115]
Challenge
Finding a specific molecule that selectively inhibits the activity of a certain E3 ligase and/or the protein–protein interactions implicated in the disease remains as one of the important and expanding research area. Moreover, as ubiquitination is a multi-step process with various players and intermediate forms, consideration of the much complex interactions between components needs to be taken heavily into account while designing the small molecule inhibitors.[105]
Similar proteins
Ubiquitin is the most-understood post-translation modifier, however, several family of
These related molecules have novel functions and influence diverse biological processes. There is also cross-regulation between the various conjugation pathways, since some proteins can become modified by more than one UBL, and sometimes even at the same lysine residue. For instance, SUMO modification often acts antagonistically to that of ubiquitination and serves to stabilize protein substrates. Proteins conjugated to UBLs are typically not targeted for degradation by the proteasome but rather function in diverse regulatory activities. Attachment of UBLs might, alter substrate conformation, affect the affinity for ligands or other interacting molecules, alter substrate localization, and influence protein stability.
UBLs are structurally similar to ubiquitin and are processed, activated, conjugated, and released from conjugates by enzymatic steps that are similar to the corresponding mechanisms for ubiquitin. UBLs are also translated with C-terminal extensions that are processed to expose the invariant C-terminal LRGG. These modifiers have their own specific E1 (activating), E2 (conjugating) and E3 (ligating) enzymes that conjugate the UBLs to intracellular targets. These conjugates can be reversed by UBL-specific isopeptidases that have similar mechanisms to that of the deubiquitinating enzymes.[77]
Within some species, the recognition and destruction of sperm mitochondria through a mechanism involving ubiquitin is responsible for sperm mitochondria's disposal after fertilization occurs.[124]
Prokaryotic origins
Ubiquitin is believed to have descended from bacterial proteins similar to
Prokaryotic ubiquitin-like protein (Pup) and ubiquitin bacterial (UBact)
Prokaryotic ubiquitin-like protein (Pup) is a functional analog of ubiquitin which has been found in the gram-positive bacterial phylum Actinomycetota. It serves the same function (targeting proteins for degradations), although the enzymology of ubiquitylation and pupylation is different, and the two families share no homology. In contrast to the three-step reaction of ubiquitylation, pupylation requires two steps, therefore only two enzymes are involved in pupylation.
In 2017, homologs of Pup were reported in five phyla of
The finding of the Pup/UBact-proteasome system in both gram-positive and gram-negative bacteria suggests that either the Pup/UBact-proteasome system evolved in bacteria prior to the split into gram positive and negative clades over 3000 million years ago or,[131] that these systems were acquired by different bacterial lineages through horizontal gene transfer(s) from a third, yet unknown, organism. In support of the second possibility, two UBact loci were found in the genome of an uncultured anaerobic methanotrophic Archaeon (ANME-1;locus CBH38808.1 and locus CBH39258.1).
Human proteins containing ubiquitin domain
These include ubiquitin-like proteins.
;Related proteins
Prediction of ubiquitination
Currently available prediction programs are:
- UbiPred is a SVM-based prediction server using 31 physicochemical properties for predicting ubiquitylation sites.[132]
- UbPred is a random forest-based predictor of potential ubiquitination sites in proteins. It was trained on a combined set of 266 non-redundant experimentally verified ubiquitination sites available from our experiments and from two large-scale proteomics studies.[133]
- CKSAAP_UbSite is SVM-based prediction that employs the composition of k-spaced amino acid pairs surrounding a query site (i.e. any lysine in a query sequence) as input, uses the same dataset as UbPred.[134]
Podcast
Investigating the ubiquitin proteasome system was the focus of a Dementia Researcher Podcast.[135] The podcast was published on 16 August 2021, hosted by Professor Selina Wray from University College London.
See also
- Autophagy
- Autophagin
- Endoplasmic-reticulum-associated protein degradation
- JUNQ and IPOD
- Prokaryotic ubiquitin-like protein
- SUMO enzymes
References
- ^ PMID 1078892.
- PMID 16230621.
- ^ PMID 20418328.
- ^ PMID 11917093.
- ^ S2CID 35434448.
- ^ PMID 12860974.
- ^ PMID 15571809.
- ^ S2CID 30693177.
- ^ PMID 23732108.
- ^ PMID 17609522.
- ^ a b "The Nobel Prize in Chemistry 2004". Nobelprize.org. Retrieved 2010-10-16.
- ^ "The Nobel Prize in Chemistry 2004: Popular Information". Nobelprize.org. Retrieved 2013-12-14.
- PMID 22925675.
- PMID 21227875.
- ^ PMID 11395416.
- PMID 11955054.
- ^ S2CID 11992443.
- PMID 9774340.
- S2CID 15114828.
- PMID 23696636.
- PMID 18836078.
- PMID 15226418.
- PMID 15289458.
- PMID 15254040.
- PMID 18981217.
- PMID 11127826.
- PMID 10807912.
- PMID 12202215.
- S2CID 32330510.
- PMID 21345952.
- PMID 15163661.
- PMID 20713359.
- PMID 14733935.
- ^ PMID 19336407.
- ^ PMID 20807509.
- PMID 23560854.
- PMID 25436519.
- PMID 7615550.
- ^ PMID 17006537.
- ^ PMID 17502423.
- PMID 15994556.
- PMID 18272573.
- PMID 17550898.
- PMID 17726030.
- PMID 17209040.
- ^ PMID 18166654.
- ^ PMID 23259061.
- PMID 20442859.
- PMID 20980512.
- PMID 20519503.
- PMID 21172657.
- PMID 22420755.
- PMID 19352404.
- PMID 18353650.
- S2CID 21280193.
- PMID 22389392.
- PMID 19775879.
- PMID 16753028.
- PMID 10089879.
- PMID 11397792.
- S2CID 11526100.
- PMID 19805293.
- ^ PMID 19754430.
- ^ PMID 18516089.
- PMID 18351785.
- ^ S2CID 205013847.
- PMID 16738015.
- ^ PMID 22389393.
- PMID 23314748.
- PMID 12551915.
- PMID 17069764.
- S2CID 23733705.
- S2CID 4384869.
- ^ PMID 20385835.
- PMID 17426036.
- PMID 25752577.
- ^ a b "Ubiquitin Proteasome Pathway Overview". Archived from the original on 2008-03-30. Retrieved 2008-04-30.
- PMID 31200561.
- PMID 29796309.
- PMID 20847899.
- PMID 23416108.
- PMID 22589545.
- PMID 17709392.
- S2CID 31636419.
- PMID 19213613.
- PMID 22033037.
- PMID 19489724.
- S2CID 15575576.
- ^ S2CID 3056635.
- PMID 22482907.
- ^ S2CID 205394130.
- S2CID 216095994.
- ^ "UBQLN1 ubiquilin 1 [ Homo sapiens ]". Gene. National Center for Biotechnology Information. Retrieved 9 May 2012.
- ^ PMID 21852239.
- "Alzheimer's brains found to have lower levels of key protein". ScienceDaily (Press release). September 1, 2011.
- PMID 26712804.
- S2CID 223338.
- PMID 29354136.
- S2CID 44003147.
- ^ PMID 23902637.
- PMID 18793634.
- PMID 22194680.
- PMID 18929679.
- PMID 11504494.
- PMID 11498385.
- ^ PMID 12531181.
- ^ PMID 11331613.
- ^ PMID 9515795.
- ^ S2CID 27437768.
- ^ PMID 9671804.
- ^ PMID 11278247.
- ^ PMID 15041716.
- ^ PMID 12734385.
- ^ PMID 10487825.
- ^ PMID 16034054.
- ^ PMID 12738243.
- PMID 12456688.
- ^ PMID 22158902.
- PMID 20694012.
- PMID 20501843.
- S2CID 23586705.
- PMID 16831869.
- S2CID 7373421.
- PMID 19253967.
- PMID 10906068.
- S2CID 29632248.
- S2CID 3224437.
- PMID 19325621.
- PMID 23479445.
- PMID 30002364.
- ^ PMID 28087277.
- PMID 27965376.
- PMID 18625080.
- PMID 19722269.
- PMID 21829559.
- ^ "Stream episode Investigating the ubiquitin proteasome system by Dementia Researcher podcast | Listen online for free on SoundCloud".
External links
- GeneReviews/NCBI/NIH/UW entry on Angelman syndrome
- OMIM entries on Angelman syndrome
- UniProt entry for ubiquitin
- "7.340 Ubiquitination: The Proteasome and Human Disease". MIT OpenCourseWare. 2004. Notes from MIT course.
- Ubiquitin at the U.S. National Library of Medicine Medical Subject Headings (MeSH)

