Ubiquitin ligase
| Ubiquitin—protein ligase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
ExPASy NiceZyme view | | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| Ubiquitin ligase | |
|---|---|
| Identifiers | |
| Symbol | Ubiquitin ligase |
| OPM superfamily | 471 |
| OPM protein | 4v6p |
| Membranome | 240 |
A ubiquitin ligase (also called an E3 ubiquitin ligase) is a
Ubiquitination system
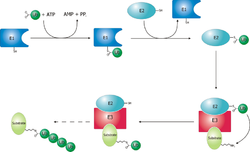
The ubiquitin ligase is referred to as an E3, and operates in conjunction with an
The ubiquitylation reaction proceeds in three or four steps depending on the mechanism of action of the E3 ubiquitin ligase. In the conserved first step, an E1 cysteine residue attacks the ATP-activated C-terminal glycine on ubiquitin, resulting in a thioester Ub-S-E1 complex. The energy from ATP and diphosphate hydrolysis drives the formation of this reactive thioester, and subsequent steps are thermoneutral. Next, a transthiolation reaction occurs, in which an E2 cysteine residue attacks and replaces the E1. HECT domain type E3 ligases will have one more transthiolation reaction to transfer the ubiquitin molecule onto the E3, whereas the much more common RING finger domain type ligases transfer ubiquitin directly from E2 to the substrate.[5] The final step in the first ubiquitylation event is an attack from the target protein lysine amine group, which will remove the cysteine, and form a stable isopeptide bond.[6] One notable exception to this is p21 protein, which appears to be ubiquitylated using its N-terminal amine, thus forming a peptide bond with ubiquitin.[7]
Ubiquitin ligase families
Humans have an estimated 500-1000 E3 ligases, which impart substrate specificity onto the E1 and E2.
Mono- and poly-ubiquitylation

Ubiquitin signaling relies on the diversity of ubiquitin tags for the specificity of its message. A protein can be tagged with a single ubiquitin molecule (monoubiquitylation), or variety of different chains of ubiquitin molecules (polyubiquitylation).[11] E3 ubiquitin ligases catalyze polyubiquitination events much in the same way as the single ubiquitylation mechanism, using instead a lysine residue from a ubiquitin molecule currently attached to substrate protein to attack the C-terminus of a new ubiquitin molecule.[6][11] For example, a common 4-ubiquitin tag, linked through the lysine at position 48 (K48) recruits the tagged protein to the proteasome, and subsequent degradation.[11] However, all seven of the ubiquitin lysine residues (K6, K11, K27, K29, K33, K48, and K63), as well as the N-terminal methionine are used in chains in vivo.[11]
Monoubiquitination has been linked to membrane protein
Monoubiquitination also can regulate cytosolic protein localization. For example, the E3 ligase
Substrate recognition
Ubiquitin ligases are the final, and potentially the most important determinant of substrate specificity in ubiquitination of proteins.[14] The ligases must simultaneously distinguish their protein substrate from thousands of other proteins in the cell, and from other (ubiquitination-inactive) forms of the same protein. This can be achieved by different mechanisms, most of which involve recognition of degrons: specific short amino acid sequences or chemical motifs on the substrate.[15]
N-degrons
Phosphodegrons

A degron can be converted into its active form by a post-translational modification[19] such as phosphorylation of a tyrosine, serine or threonine residue.[20] In this case, the ubiquitin ligase exclusively recognizes the phosphorylated version of the substrate due to stabilization within the binding site. For example, FBW7, the F-box substrate recognition unit of an SCFFBW7ubiquitin ligase, stabilizes a phosphorylated substrate by hydrogen binding its arginine residues to the phosphate, as shown in the figure to the right. In absence of the phosphate, residues of FBW7 repel the substrate.[18]
Oxygen and small molecule dependent degrons
The presence of
Misfolded and sugar degrons
In addition to recognizing amino acids, ubiquitin ligases can also detect unusual features on substrates that serve as signals for their destruction.[14] For example, San1 (Sir antagonist 1), a nuclear protein quality control in yeast, has a disordered substrate binding domain, which allows it to bind to hydrophobic domains of misfolded proteins.[14] Misfolded or excess unassembled glycoproteins of the ERAD pathway, on the other hand, are recognized by Fbs1 and Fbs2, mammalian F-box proteins of E3 ligases SCFFbs1and SCFFbs2.[23] These recognition domains have small hydrophobic pockets allowing them to bind high-mannose containing glycans.
Structural motifs
In addition to linear
Disease relevance
E3 ubiquitin ligases regulate homeostasis, cell cycle, and DNA repair pathways, and as a result, a number of these proteins are involved in a variety of cancers, including famously MDM2,
Examples
- A RING (Really Interesting New Gene) domain binds the E2 conjugase and might be found to mediate enzymatic activity in the E2-E3 complex[29]
- An F-box domain (as in the SCF complex) binds the ubiquitinated substrate. (e.g., Cdc 4, which binds the target protein Sic1; Grr1, which binds Cln).[30]
- A HECT domain, which is involved in the transfer of ubiquitin from the E2 to the substrate.
Individual E3 ubiquitin ligases
- E3A
- mdm2
- Anaphase-promoting complex (APC)
- UBR5 (EDD1)
- LNXp80
- CBX4, CBLL1
- HACE1
- HECTD1, HECTD2, HECTD3, HECTD4
- HECW1, HECW2
- HERC1, HERC2, HERC3, HERC4, HERC5, HERC6
- HUWE1, ITCH
- NEDD4, NEDD4L
- PPIL2
- PRPF19
- RANBP2
- RNF4
- RBX1
- SMURF1, SMURF2
- STUB1
- TOPORS
- TRIP12
- UBE4A, UBE4B
- UBOX5
- UBR5
- VHL
- WWP1, WWP2
- Parkin
- MKRN1
See also
References
- PMID 22266821.
- PMID 9759494.
- PMID 23495935.
- PMID 18213395.
- PMID 22389392.
- ^ ISBN 978-0-9747077-3-0.[page needed]
- PMID 14532004.
- ^ S2CID 19594293.
- PMID 15520277.
- PMID 3041007.
- ^ S2CID 19237120.
- PMID 12651740.
- S2CID 43124248.
- ^ PMID 28375744.
- PMID 18698327.
- S2CID 10555455.
- PMID 22524314.
- ^ PMID 28130986.
- PMID 26268526.
- S2CID 149598.
- S2CID 20914281.
- S2CID 41227590.
- PMID 30837888.
- PMID 21863050.
- ^ PMID 25624711.
- PMID 25415135.
- S2CID 16385927.
- S2CID 85527795.
- PMID 16250895.
- PMID 8706131.
External links
- Quips article describing E3 Ligase function Archived 2012-11-30 at the Wayback Machine at PDBe
- Ubiquitin-Protein+Ligases at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- EC 6.3.2.19
