Variants of PCR
The versatility of polymerase chain reaction (PCR) has led to modifications of the basic protocol being used in a large number of variant techniques designed for various purposes. This article summarizes many of the most common variations currently or formerly used in molecular biology laboratories; familiarity with the fundamental premise by which PCR works and corresponding terms and concepts is necessary for understanding these variant techniques.
Basic modifications
Often only a small modification needs to be made to the standard PCR protocol to achieve a desired goal:

Variable Number of Tandem Repeats (VNTR) PCR targets repetitive areas of the genome that exhibit
Asymmetric PCR preferentially amplifies one strand of a double-stranded DNA target. It is used in some
Some modifications are needed to perform long PCR. The original Klenow-based PCR process did not generate products that were larger than about 400 bp. Taq polymerase can however amplify targets of up to several thousand bp long.[4] Since then, modified protocols with Taq enzyme have allowed targets of over 50 kb to be amplified.[5]
Quantitative Real-Time PCR (QRT-PCR), sometimes simply called Real-Time PCR (RT-PCR), refers to a collection of methods that use fluorescent dyes, such as Sybr Green, or fluorophore-containing DNA probes, such as TaqMan, to measure the amount of amplified product in real time as the amplification progresses.
Hot-start PCR is a technique performed manually by heating the reaction components to the DNA melting temperature (e.g. 95 °C) before adding the polymerase. In this way, non-specific amplification at lower temperatures is prevented.[6] Alternatively, specialized reagents inhibit the polymerase's activity at ambient temperature, either by the binding of an antibody, or by the presence of covalently bound inhibitors that only dissociate after a high-temperature activation step. 'Hot-start/cold-finish PCR' is achieved with new hybrid polymerases that are inactive at ambient temperature and are only activated at elevated temperatures.
In touchdown PCR, the annealing temperature is gradually decreased in later cycles. The annealing temperature in the early cycles is usually 3–5 °C above the standard Tm of the primers used, while in the later cycles it is a similar amount below the Tm. The initial higher annealing temperature leads to greater specificity for primer binding, while the lower temperatures permit more efficient amplification at the end of the reaction.[7]
Assembly PCR (also known as Polymerase Cycling Assembly or PCA) is the synthesis of long DNA structures by performing PCR on a pool of long oligonucleotides with short overlapping segments, to assemble two or more pieces of DNA into one piece. It involves an initial PCR with primers that have an overlap and a second PCR using the products as the template that generates the final full-length product. This technique may substitute for ligation-based assembly.[8]
In colony PCR, bacterial colonies are screened directly by PCR, for example, the screen for correct DNA vector constructs. Colonies are sampled with a sterile pipette tip and a small quantity of cells transferred into a PCR mix. To release the DNA from the cells, the PCR is either started with an extended time at 95 °C (when standard polymerase is used), or with a shortened denaturation step at 100 °C and special chimeric DNA polymerase.[9]
The digital polymerase chain reaction simultaneously amplifies thousands of samples, each in a separate droplet within an emulsion or partition within an micro-well.
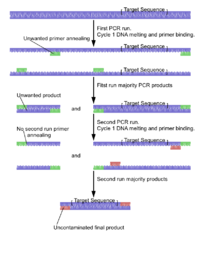
Suicide PCR is typically used in paleogenetics or other studies where avoiding false positives and ensuring the specificity of the amplified fragment is the highest priority. It was originally described in a study to verify the presence of the microbe Yersinia pestis in dental samples obtained from 14th-century graves of people supposedly killed by plague during the medieval Black Death epidemic.[10] The method prescribes the use of any primer combination only once in a PCR (hence the term "suicide"), which should never have been used in any positive-control PCR reaction, and the primers should always target a genomic region never amplified before in the lab using this or any other set of primers. This ensures that no contaminating DNA from previous PCR reactions is present in the lab, which could otherwise generate false positives.
COLD-PCR (co-amplification at lower denaturation temperature-PCR) is a modified protocol that enriches variant alleles from a mixture of wild-type and mutation-containing DNA samples.
Pretreatments and extensions
The basic PCR process can sometimes precede or follow another technique.
Two-tailed PCR uses a single primer that binds to a microRNA target with both 3' and 5' ends, known as hemiprobes.[11] Both ends must be complementary for binding to occur. The 3'-end is then extended by reverse transcriptase forming a long cDNA. The cDNA is then amplified using two target specific PCR primers. The combination of two hemiprobes, both targeting the short microRNA target, makes the Two-tailed assay exceedingly sensitive and specific.
Ligation-mediated PCR uses small DNA oligonucleotide 'linkers' (or adaptors) that are first ligated to fragments of the target DNA. PCR primers that anneal to the linker sequences are then used to amplify the target fragments. This method is deployed for DNA sequencing, genome walking, and DNA footprinting.[12] A related technique is amplified fragment length polymorphism, which generates diagnostic fragments of a genome.
Methylation-specific PCR (MSP) is used to identify patterns of
Other modifications
Adjustments of the components in PCR is commonly used for optimal performance.
The divalent magnesium ion (Mg++) is required for PCR polymerase activity. Lower concentrations Mg++ will increase replication fidelity, while higher concentrations will introduce more mutations.[15]
Denaturants(such as
DNA polymerases occasionally incorporate mismatch bases into the extending strand. High-fidelity PCR employs enzymes with 3'-5' exonuclease activity that decreases this rate of mis-incorporation. Examples of enzymes with proofreading activity include Pfu; adjustments of the Mg++ and dNTP concentrations may help maximize the number of products that exactly match the original target DNA. [citation needed]
Primer modifications
Adjustments to the synthetic oligonucleotides used as primers in PCR are a rich source of modification:
Normally PCR primers are chosen from an invariant part of the genome, and might be used to amplify a polymorphic area between them. In allele-specific PCR the opposite is done. At least one of the primers is chosen from a polymorphic area, with the mutations located at (or near) its 3'-end. Under stringent conditions, a mismatched primer will not initiate replication, whereas a matched primer will. The appearance of an amplification product therefore indicates the genotype. (For more information, see SNP genotyping.)
InterSequence-Specific PCR (or ISSR-PCR) is method for
Primers can also be designed to be 'degenerate' – able to initiate replication from a large number of target locations.
Normally the primers used in PCR are designed to be fully complementary to the target. However, the polymerase is tolerant to mis-matches away from the 3' end. Tailed-primers include non-complementary sequences at their 5' ends. A common procedure is the use of linker-primers, which ultimately place restriction sites at the ends of the PCR products, facilitating their later insertion into cloning vectors.
An extension of the 'colony-PCR' method (above), is the use of vector primers. Target DNA fragments (or cDNA) are first inserted into a cloning vector, and a single set of primers are designed for the areas of the vector flanking the insertion site. Amplification occurs for whatever DNA has been inserted.[4]
PCR can easily be modified to produce a labeled product for subsequent use as a hybridization probe. One or both primers might be used in PCR with a radioactive or fluorescent label already attached, or labels might be added after amplification. These labeling methods can be combined with 'asymmetric-PCR' (above) to produce effective hybridization probes.
RNase H-dependent PCR (rhPCR) can reduce primer-dimer formation, and increase the number of assays in multiplex PCR. The method utilizes primers with a cleavable block on the 3’ end that is removed by the action of a thermostable RNase HII enzyme.[17]
DNA Polymerases
There are several DNA polymerases that are used in PCR.
The
The bacteriophage T4 DNA polymerase (family A) was also initially used in PCR. It has a higher fidelity of replication than the Klenow fragment, but is also destroyed by heat. T7 DNA polymerase (family B) has similar properties and purposes. It has been applied to site-directed mutagenesis[18] and Sanger sequencing.[19]
Taq polymerase, the DNA Polymerase I from Thermus aquaticus, was the first thermostable polymerase used in PCR, and is still the one most commonly used. The enzyme can be isolated from its native source, or from its cloned gene expressed in E. coli.[4] A 61kDa truncated from lacking 5'-3' exonuclease activity is known as the Stoffel fragment, and is expressed in E. coli.[20] The lack of exonuclease activity may allow it to amplify longer targets than the native enzyme. It has been commercialized as AmpliTaq and Klentaq.[21] A variant designed for hot-start PCR called the "Faststart polymerase" has also been produced. It requires strong heat activation, thereby avoiding non-specific amplification due to polymerase activity at low temperature. Many other variants have been created.[22]
Other Thermus polymerases, such as Tth polymerase I (P52028) from Thermus thermophilus, has seen some use. Tth has reverse transcriptase activity in the presence of Mn2+ ions, allowing PCR amplification from RNA targets.[23]
The archean genus Pyrococcus has proven a rich source of thermostable polymerases with proofreading activity. Pfu DNA polymerase, isolated from the P. furiosus shows a 5-fold decrease in the error rate of replication compared to Taq.[24] Since errors increase as PCR progresses, Pfu is the preferred polymerase when products are to be individually cloned for sequencing or expression. Other lesser used polymerases from this genus include Pwo (P61876) from Pyrococcus woesei, Pfx from an unnamed species, "Deep Vent" polymerase (Q51334) from strain GB-D.[25]
Vent or
Mechanism modifications
Sometimes even the basic mechanism of PCR can be modified.

Unlike normal PCR, Inverse PCR allows amplification and sequencing of DNA that surrounds a known sequence. It involves initially subjecting the target DNA to a series of restriction enzyme digestions, and then circularizing the resulting fragments by self ligation. Primers are designed to be extended outward from the known segment, resulting in amplification of the rest of the circle. This is especially useful in identifying sequences to either side of various genomic inserts.[26]
Similarly, thermal asymmetric interlaced PCR (or TAIL-PCR) is used to isolate unknown sequences flanking a known area of the genome. Within the known sequence, TAIL-PCR uses a nested pair of primers with differing annealing temperatures. A 'degenerate' primer is used to amplify in the other direction from the unknown sequence.[27]
Isothermal amplification methods
Some DNA amplification protocols have been developed that may be used alternatively to PCR. They are isothermal, meaning that they are run at a constant temperature.[28]
Nicking enzyme amplification reaction (NEAR) and its cousin strand displacement amplification (SDA) are isothermal, replicating DNA at a constant temperature using a polymerase and nicking enzyme.[28]
Other types of isothermal amplification include whole genome amplification (WGA), Nucleic acid sequence-based amplification (NASBA), and transcription-mediated amplification (TMA).[28]
See also
References
- PMID 18282271.
- PMID 3200828.
- )
- ^ PMID 2448875.
- PMID 8202550.
- PMID 1579465.
- PMID 1861999.
- PMID 7590320.
- ISBN 978-0-7637-3383-4.
- PMID 11058154.
- PMID 28911110.
- PMID 2814500.
- PMID 8790415.
- PMID 24107250.
- PMID 11835531.
- S2CID 41802285.
- PMID 21831278.
- PMID 2726477.
- ^ "Thermo Sequenase DNA Polymerase".
- PMID 8324500.
- ^ "Applied Biosystems - Support". www6.appliedbiosystems.com.
- PMID 11054459.
- ^ https://www.promega.in/products/pcr/rt-pcr/tth-dna-polymerase/ [dead link]
- PMID 8836181.
- ^ ISBN 978-1-4020-6240-7.
- PMID 2852134.
- PMID 7759102.
- ^ a b c "Isothermal Amplification - Application Overview". New England BioLabs, Inc. 2020. Retrieved 13 August 2020.
- PMID 15247927.
- PMID 10871386.
- PMID 16756388.
- PMID 20300675.
