Viral quasispecies
A viral quasispecies is a population structure of viruses with a large number of variant genomes (related by mutations). Quasispecies result from high mutation rates as mutants arise continually and change in relative frequency as viral replication and selection proceeds.[1]
The theory predicts that a viral quasispecies at a low but evolutionarily neutral and highly connected (that is, flat) region in the fitness landscape will outcompete a quasispecies located at a higher but narrower fitness peak in which the surrounding mutants are unfit.[2][3] This phenomenon has been called 'the quasispecies effect' or, more recently, the 'survival of the flattest'.[4]
The term quasispecies was adopted from a theory of the
The quasispecies model is most applicable when the genome size is limited and the mutation rate is high, and so is most relevant to
The relevance of quasispecies in virology has been the subject of extended debate. However, standard clonal analyses and
History
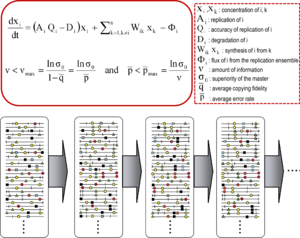
Quasispecies theory was developed in the 1970s by
The core quasispecies concepts are described by two fundamental equations: replication with production of error copies, and the error threshold relationship. They capture two major features of RNA viruses at the population level: the presence of a mutant spectrum, and the adverse effect of an increase of mutation rate on virus survival, each with several derivations.[1]
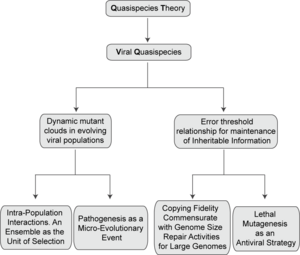
The existence of a mutant spectrum was experimentally evidenced first by clonal analyses of
High mutation rates and quasispecies were verified for other RNA viruses based on dissection of viral populations by molecular or biological cloning, and sequence analysis of individual clones. John Holland and colleagues were the first to recognize that a rapidly evolving RNA world inserted in a DNA-based biosphere had multiple evolutionary and medical implications.
Current scope
The first mathematical formulation of quasispecies was deterministic; it assumed steady state mutant distributions in
Dynamic heterogeneity
The molecular basis of high error rates is the limited template-copying fidelity of RNA-dependent RNA polymerases (RdRps) and RNA-dependent DNA polymerases (also termed reverse transcriptases, RTs). In addition, these enzymes are defective in proofreading[29] because they lack a 3’ to 5’ exonuclease domain present in replicative cellular DNA polymerases.[30] Also, postreplicative-repair pathways, abundant to correct genetic lesions in replicating cellular DNA, appear as ineffective for double-stranded RNA or RNA-DNA hybrids. The presence of a proofreading-repair activity in coronaviruses increases their copying accuracy in about 15-fold.[31] This and other repair activities, that may act on standard RNA or retroviral genomes,[32][33][34][35] do not prevent the formation of mutant spectra, although their amplitude may be lower than for other RNA viruses, at least in populations close to a clonal (single genome) origin. Quasispecies dynamics will operate in any viral or cellular system in which due to high mutation rates (as a result of low fidelity nucleic acid polymerases or environmental alterations) mutant spectra are rapidly generated.[10][36][37][38][39][40]
Studies with different
Phenotypic reservoirs
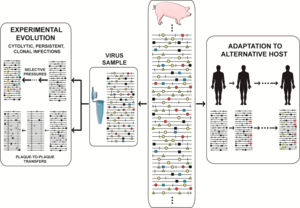
The crux of the matter regarding quasispecies implications is that at any given time, the viral population includes a reservoir not only of
Mutant spectrum dynamics has been depicted in different ways, and we have chosen one that encompasses frequent events in natural populations and research designs, such as
Limitations and indeterminacies
The nucleotide sequence of an individual genome from a population (no matter which the degree of population complexity might be), can be determined either following a biological or molecular cloning event or by deep sequencing of entire viral genomes, in a manner that mutation linkage (assignment of different mutations to the same genome molecule) can be established. Each of these procedures implies some limitations: biological cloning can bias the representation in favor of infectious genomes, while molecular cloning can introduce non-infectious (defective) genomes in the analysis.[22][58][59] Whole genome quasispecies description is still technically challenging due to the artifactual introduction of mutations. Most current deep sequencing platforms yield sequences of short reads for a given amplicon (sequence under analysis); minority mutations in an amplicon cannot be reliably linked to mutations in a different amplicon of the same genome; at most, statistical inferences on linkage can be proposed. Despite these limitations, control experiments and improvements of bioinformatic procedures support that the majority of sequence heterogeneity analyzed in viral populations indeed reflects differences in the natural template populations. If mutation linkage can be solved on a routine basis, a new wave of molecular information relevant to epistatic interactions will enter the picture.
There are additional levels of indeterminacy in the sequential analysis of viral populations, in particular those replicating in vivo. Components of the mutant spectrum represented at a given time in the sample taken for sequencing may differ from those in the next time point, due either to sampling uncertainties or bona fide fluctuations of genome frequencies. It is not justified to accept a rough similarity because even a single mutation in a given sequence context may affect biological properties.[10] In the words of John Holland and colleagues: "It is important to remember that every quasispecies genome swarm in an infected individual is unique and "new" in the sense that no identical population of genomes has ever existed before and none such will ever exist again".[63] On top of the fleeting nature of any mutant distribution, the standard methods available for quasispecies characterization provide genomic sequences of a minority of the population (estimated in 10−8 to 10−13 for molecular cloning-Sanger sequencing, and in 10−6 to 10−11 for deep sequencing).[58] We can only have an approximate representation of viral populations and their dynamics, as evidenced by many experimental studies.[15][20][10][43][44][58][60][64]
Non-consensus-based descriptors
The points summarized in previous sections fully justifies addressing analytical tools towards the mutant spectrum rather than ignoring it or considering its presence a side issue. Use of consensus sequences to describe the genome of a virus isolate, despite being warranted by the difficulties of conveying the information recapitulated in a mutant spectrum, blurs and enfeebles biological interpretations. Experimental results have demonstrated that minority genomes from a mutant spectrum (that cannot be identified by examining the consensus sequence) can include mutations that confer resistance to
Collective response
Mutant spectra are not mere aggregates of mutants acting independently. They are often engaged in collective responses.[1] Two major types are those that depend on the presence of sets of variants, and those that rely on intra-mutant spectrum interactions.
Variants that drive responses to selective constraints
Behavior of reconstructed quasispecies
In some cases of
Quasispecies memory
Quasispecies memory is a type of molecular memory dependent on the recent history of the evolutionary lineage and the integrity of the mutant spectrum.[69][70] The search for memory was prompted by the complex adaptive system behavior of a viral quasispecies, suggested by the presence of core information (considered the one that defines viral identity) despite variation of constitutive elements (the mutant spectrum). A well-known example is memory in the immune system that mobilizes and expands minority components in response to stimuli previously faced by the system.[71] In the experiments designed to identify memory in viral quasispecies, members of the mutant spectrum increased in frequency as a consequence of their replication during a selection event that drove them towards dominance. When the selective constraint was withdrawn, memory genomes remained at levels that were 10- to 100-fold higher than the basal levels attributable solely to their generation by mutation, as documented with independent FMDV genetic markers, and with HIV-1 in vivo.[69][70][72][73] Thus, memory is a history-dependent, collective property of the quasispecies that confers a selective advantage to respond to environmental changes previously experienced by the same evolutionary lineage. It can be manifested only if the mutant spectrum maintains its completeness, since memory is lost when the population undergoes a bottleneck event that excludes minorities. A relevant example of the consequences of memory occurs in antiviral pharmacology with the administration for a second time of the same or a related antiviral agent (capable of evoking shared resistance mutations) used in a previous treatment. The second intervention may face inhibitor-resistant memory genomes from the earlier treatment, thus contributing to virus escape.[69] This is an aspect that has not received adequate attention in the planning of antiviral interventions for patients who fail a first treatment and have to be subjected to a second treatment.
Intra-mutant spectrum interactions for interference, complementation or cooperation
Individual genomes surrounded by a cloud of related mutants can be either suppressed to be kept at low frequency, or helped to be maintained in the population. The two alternative fates are dependent on several factors, one being the surrounding mutant spectrum in those steps of the infectious cycle in which an effective competition among variants is established, for example within replication complexes. This important concept was first derived theoretically,[12][74] and then approached experimentally with several viruses. In an early study, Juan Carlos de la Torre and John Holland described suppression of high fitness VSV by mutant spectra of inferior fitness.[12] Suppressive effects have since been documented with standard and mutagenized viral populations. Some examples are:
- Suppression of high fitness antigenic variants of FMDV by low fitness antibody-escape mutants.[75]
- Suppression of virulent poliovirus vaccines.[65]
- Suppression of pathogenic lymphocytic choriomengitis virus (LCMV) (that cause growth hormone deficiency in mice) by non-pathogenic LCMV variants.[76]
- Suppression of FMDV by a mutagenized FMDV population.[77]
- Suppression of FMDV by capsid and polymerase FMDV mutants.[78]
- Suppression of drug-resistant viral mutants during antiviral therapy.[79][80]
Opposite to suppression is maintenance of a mutant either by a favorable position in a fitness landscape or by interactions of
Collective behavior of viruses was documented with mutant RNA viruses resistant to nucleotide analogues. The study of this class of mutants has been instrumental for the understanding of the molecular basis of template copying fidelity, and the consequences of fidelity alterations in the adaptive capacity and pathogenic potential of RNA viruses.
Complementation (often occurring when a functional protein encoded by a set of genomes is used by another set of genomes whose encoded protein is not functional) may underlie some collective responses of quasispecies such as fitness of individuals isolated from a population being inferior to fitness of the population.[15][30] Complementation was described between two truncated FMDV genomic forms.[87] The genomes with internal deletions became detectable upon high multiplicity passage of a clonal population of standard FMDV, a virus with a monopartite single stranded RNA genome. Infectivity was generated by complementation of the two truncated forms, in absence of standard, full length FMDV genomes. For complementation to be effective, prior exploration of sequence space through point mutations was a requirement.[88] The system underwent a remarkable evolutionary transition akin to genome segmentation. Drastic genetic lesions in viral genomes are difficult to observe unless a mechanism such as complementation comes into the rescue of the deviant genomes. Additional examples of complementation among RNA viruses have been reported.[89][90][91][42][44] Complementation is a means to maintain defective genomes at detectable frequencies in viral populations.
A distinction has been made between complementation and cooperation, in which two different genomes give rise to a new phenotype through the interaction between two variant proteins.[92] An example of cooperation was characterized during studies with measles virus on membrane fusion which is essential for virus entry into cells. For this virus fusion is mediated by two proteins termed H and F. A truncated H was deficient in cell fusion but the activity was regained when the truncated H was accompanied by two forms of F but not one of the forms individually.[92]
Therefore, complementation, cooperation, interference and suppression can emerge from interactions among components of mutant spectra that have their origin in random mutations. Selection acts on whatever sets of mutants can provide a useful trait, to turn random occurrences into biological meaning.
Bottlenecks

A means to interrupt the participation of individual genomes in interactions with their mutant spectrum is for the quasispecies swarm to undergo drastic reductions in population size that isolate one or few individual genomes from their surroundings. Such reductions are termed bottlenecks, and they have an important participation in shaping evolutionary lineages for all kinds of organisms, and also for viruses. They occur frequently not only upon host-to host transmission but also inside infected hosts,[93][94][95] and they can perturb positive and negative selection events in processes that are difficult to identify and characterize.
Drastic bottleneck events have been reproduced with laboratory populations of viruses in the form of plaque-to-plaque transfers.
Deprivation of an individual virus from possible suppression, complementation or cooperation, may represent a liberation to initiate a new evolutionary process, or a condemnation to extinction. If liberated from suppression, the isolated genome must replicate and be able to reconstruct a mutant cloud to regain adaptive capability. This has led to the suggestion that high mutation rates evolved to allow such mutant spectrum recovery following bottlenecks. Other models attribute high mutation rates to adaptive optimization independent of bottlenecks, or to a mechanistic consequence of rapid replication.[58] Whatever their ultimate origins, high mutation rates serve the purpose of adaptation in multiple circumstances, not only following bottlenecks. A founder virus can introduce a different phenotype for the ensuing evolution. Evolution of viruses in nature and as disease agents can be viewed as succession of mutant spectrum alterations, subjected to expansions and reductions of population size in a continuous interplay of positive and negative selection and random drift. While short-term (for example, intra-host) evolution is observable and measurable, viruses may appear to be relatively static in the long term for decades (as seen with antigenic variants of FMDV [102]) or longer. Intra-host evolution is generally more rapid than inter-host evolution, as documented with viruses[10] and other biological systems.[103] Apparent invariance may be the result of selection for long-term survival of populations that have previously frenziedly tested evolutionary outcomes in short-term processes.[58]
Viral disease
Soon after quasispecies was evidenced for viruses, some medical implications were made explicit.[20][104] Several specific and general points below.[10][43][66][28]
- High mutation rates and population heterogeneity endow viruses with the potential to escape immune pressures (including those due to vaccination) and antiviral inhibitors used in therapy. It is an open question if vaccination can promote long-term evolution of antigenic determinants.
- Attenuated RNA virus vaccines can revert to virulent forms. RNA viruses released in nature for pest control purposes can mutate to new phenotypes.
- Virus attenuation and virulence is dependent on viral genetic traits. Variant forms of a given virus may display increased virulence or atypical disease.
- Components of a mutant spectrum can exhibit a different host rangethan most genomes in the same population, with implications for the emergence and re-emergence of viral disease.
- Viral pathogenesis is influenced by microevolutionary processes in which some viral subpopulations are replaced by others to persist or to invade new cell types, tissues or organs.
- The larger the actively replicating (effective) population size and the replication rate, the most effective is exploration of sequence space for phenotypic expansions that favor survival and persistence.
- There is a connection between four parameters that characterize viruses during infection processes: replication rate (the rate at which viral RNA or DNA is synthesized intracellularly for viral progeny production), viral load (the total amount of virus quantified in an infected host or host compartment), genetic heterogeneity, and replicative fitness (the yield of infectious particles that can contribute to the next generation). They can influence disease progression, and any of them can be targeted for disease control.
In all interactions conductive to disease, the host cells individually and as groups in tissues and organs play decisive roles. The consequences of a viral infection are always host-dependent. However, the virus itself poses a major challenge that a deeper understanding of quasispecies dynamics is helping to confront.[28]
Antiviral strategies
There is an increasing perception that Darwinian principles should assist in the planning of antiviral designs.[105] The aim of vaccination is to evoke a protective response that either prevents virus replication or disease. The aim of an antiviral pharmacological intervention is to inhibit virus replication to provide the immune system with an opportunity to clear the virus. Expressed simply, the direct danger for vaccination and treatment is that the virus can escape through selection of mutants resistant to vaccine-triggered defense components or to the externally administered inhibitors. This has led to several proposals to confront viral disease, that can be summarized below.[58]
Vaccine exposure of multiple B cell and T cell epitopes
Vaccines should include repertoires of B cell and T cell epitopes to evoke an ample immune response. The broad response should minimize selection of escape mutants that may be present as minority components in mutant spectra, as repeatedly documented experimentally.[10][22][44][69] With the current types of available vaccines, those that best comply with the multiple epitope requirement are, in the order of expected efficacy to confer protection against highly variable viruses: attenuated > inactivated whole virus > several expressed proteins > one expressed protein > multiple synthetic peptide antigens > single peptide antigen. The scarcity of effective synthetic vaccines for RNA viral pathogens despite huge scientific and economic efforts is a reflection of the underlying problems.
Antiviral agents used in combination
Antiviral
- Inhibitors used in combination should target different viral gene products.
- Splitting a treatment into two steps: first an induction regimen, and a second maintenance regimen. Drugs administered in the two steps should be different.
- Targeting of cellular functions needed for the virus life cycle.
- Use of innate immune response-stimulating drugs (for example, inhibitors of enzymes involved in pyrimidine biosynthesis).
- Combined use of immunotherapy and chemotherapy.
- Lethal mutagenesis or virus extinction by excess of mutations introduced during viral replication.
These strategies have as their main objective to avoid selection of treatment-escape mutants by multiple selective constraints that cannot be surmounted by the virus.[58][106] Control is effective either because exploration of sequence space cannot reach the required multiple mutations (even when recombination is available) or because the multiple mutations inflict a severe fitness cost.[106] Vaccines exposing multiple epitopes and combination therapies follow the same strategy whose aim is to limit possible escape routes to viral quasispecies in the face of the suppressive constraint.
Lethal mutagenesis
Defense mechanisms based on genome modification of invading genetic parasites such as editing cellular activities that are recruited as part of the innate immune response (ADAR, APOBEC, RIP, etc.)[110] represent a natural counterpart of the principle utilized by lethal mutagenesis. Applicability to pathogenic cellular elements is a real possibility, and lethal mutagenesis to control tumor cells is an active field of investigation.[111][112] Thus, the recognition of quasispecies dynamics has suggested some fundamental guidelines for disease prevention and control that are gradually permeating clinical practice. This is in line with the recognized need to apply Darwinian principles to the control of infectious disease.
Error threshold
This may be defined as "The inability of a genetic element to be maintained in a population as the fidelity of its replication machinery decreases beyond a certain threshold value".[113]
In theory, if the mutation rate was sufficiently high, the viral population would not be able to maintain the genotype with the highest fitness, and therefore the ability of the population to adapt to its environment would be compromised. A practical application of this dynamic is in antiviral drugs employing lethal mutagenesis. For example, increased doses of the mutagen ribavirin reduces the infectivity of poliovirus.[114]
However, these models assume that only the mutations that occur in the fittest sequence are deleterious, and furthermore that they are non-lethal. It has been argued that, if we take into account the deleterious effect of mutations on the population of variants and the fact that many mutations are lethal, then the error threshold disappears, i.e. the fittest sequence always maintains itself.[115][113][116] Empirical data on the effect of mutations in viruses is rare, but appears to correspond with this scenario.[117]
Possible evolutionary consequences
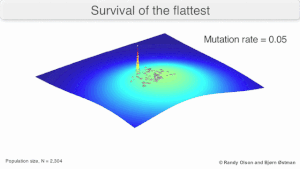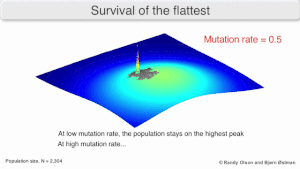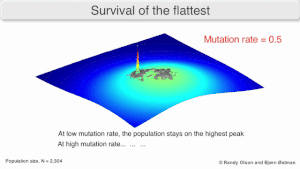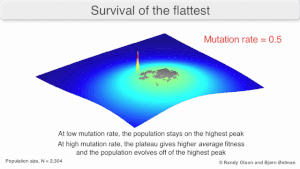
Mutational robustness
The long-term evolution of the virus may be influenced in that it may be a better evolutionarily stable strategy to generate a broad quasispecies with members of approximately equal fitness than to have a sharply defined 'most fit' single genotype (with mutational neighbours substantially less fit). This has been called 'survival of the flattest' - referring to the fitness profiles of the two strategies respectively.[3]
Over the long-term, a flatter fitness profile might better allow a quasispecies to exploit changes in
However, whether
Cooperation
Experimental manipulation of
References
![]() This article was adapted from the following source under a CC BY 4.0 license (2000) (reviewer reports):
Esteban Domingo Solans, Celia Perales (17 October 2019). "Viral quasispecies".
This article was adapted from the following source under a CC BY 4.0 license (2000) (reviewer reports):
Esteban Domingo Solans, Celia Perales (17 October 2019). "Viral quasispecies". {{cite journal}}: CS1 maint: unflagged free DOI (link
- ^ PMID 34586874.
- ^ PMID 10449760.
- ^ S2CID 1482925.
- ^ PMID 18253158.
- S2CID 96727272.
- PMID 21235976.
- ^ PMID 10570172.
- OCLC 958495192.
- ^ S2CID 1812273.
- ^ PMID 22688811.
- S2CID 38296619.
- ^ PMID 7159681.
- S2CID 52947045.
- PMID 26597856.
- ^ S2CID 19109157.
- ^ PMID 8207804.
- ISBN 9783527608010.
- PMID 1052321.
- PMID 23852383.
- ^ PMID 7041255.
- PMID 3912262.
- ^ ISBN 9781351076432.
- ^ PMID 16568907.
- ^ S2CID 35903181.
- ^ S2CID 260313185.
- S2CID 42131267.
- PMID 26342705.
- ^ S2CID 19514308.
- PMID 1336756.
- ^ S2CID 23203156.
- PMID 17804504.
- PMID 9037015.
- PMID 11277927.
- PMID 24348241.
- PMID 26958717.
- PMID 26373410.
- PMID 28416670.
- PMID 27795410.
- PMID 30335851.
- ^ PMID 30013589.
- ^ PMID 30768648.
- ^ PMID 29540453.
- ^ ISSN 0931-7597.
- ^ S2CID 13239618.
- PMID 27392606.
- ^ PMID 20661479.
- PMID 28539654.
- PMID 30944212.
- PMID 29593040.
- PMID 30452113.
- PMID 11222259.
- PMID 29304222.
- ^ PMID 24945141.
- ^ PMID 28910639.
- PMID 22949662.
- PMID 26195777.
- ^ PMID 24340131.
- ^ ISBN 978-0-12-800837-9.
- ^ PMID 24284629.
- ^ PMID 28275194.
- PMID 20080701.
- ^ S2CID 52006443.
- ^ S2CID 46530529.
- ^ PMID 16797586.
- ^ PMID 1846038.
- ^ OCLC 851813241.
- S2CID 52980171.
- ^ PMID 18436553.
- ^ PMID 18615120.
- ^ PMID 10729128.
- S2CID 8000538.
- PMID 12875847.
- PMID 15136042.
- )
- PMID 7690417.
- PMID 8806545.
- PMID 15016853.
- PMID 17481660.
- S2CID 16501575.
- PMID 27764731.
- PMID 11462003.
- PMID 17196038.
- ^ PMID 26210988.
- ^ PMID 16220146.
- ^ PMID 16327776.
- PMID 26499340.
- PMID 15479809.
- PMID 24757055.
- S2CID 21605827.
- PMID 22541042.
- PMID 26978794.
- ^ PMID 26162566.
- PMID 16567621.
- PMID 22921636.
- PMID 21912520.
- S2CID 4235839.
- PMID 1321432.
- PMID 14195748.
- PMID 8951375.
- PMID 9878424.
- PMID 15269370.
- PMID 1316467.
- S2CID 14433111.
- )
- PMID 20036799.
- ^ ISBN 9781493906932.
- ^ PMID 9990051.
- ^ S2CID 201673871.
- PMID 12370416.
- PMID 29324969.
- PMID 20934515.
- PMID 21593786.
- ^ PMID 16352527.
- PMID 11371613.
- PMID 16107214.
- S2CID 122142345.
- PMID 15159545.
- ^ Using fitness landscapes to visualize evolution in action, retrieved 2019-10-22
External links
- Video: Using fitness landscapes to visualize evolution in action—contains an example of "survival of the flattest"
