Virus classification
Virus classification is the process of naming
Viruses are classified by
In 2021, the ICTV changed the International Code of Virus Classification and Nomenclature (ICVCN) to mandate a binomial format (genus|| ||species) for naming new viral species similar to that used for cellular organisms; the names of species coined prior to 2021 are gradually being converted to the new format, a process which has been stated to be planned to be completed by end 2023.
As at 2022, the ICTV taxonomy listed 11,273 named virus species (including some classed as satellite viruses and others as viroids) in 2,818 genera, 264 families, 72 orders, 40 classes, 17 phyla, 9 kingdoms and 6 realms. [1] However, the number of named viruses considerably exceeds the number of named virus species since, by contrast to the classification systems used elsewhere in biology, a virus "species" is a collective name for a group of (presumably related) viruses sharing certain common features (see below). Also, the use of the term "kingdom" in virology does not equate to its usage in other biological groups, where it reflects high level groupings that separate completely different kinds of organisms (see Kingdom (biology)).
Definitions
Virus definition
The currently accepted and formal definition of a 'virus' was accepted by the ICTV Executive Committee in November 2020 and ratified in March 2021, and is as follows:[2]
Viruses
sensu stricto are defined operationally by the ICTV as a type of MGEthat encodes at least one protein that is a major component of the virion encasing the nucleic acid of the respective MGE and therefore the gene encoding the major virion protein itself or MGEs that are clearly demonstrable to be members of a line of evolutionary descent of such major virion protein-encoding entities. Any monophyletic group of MGEs that originates from a virion protein-encoding ancestor should be classified as a group of viruses.
Species definition
Species form the basis for any biological classification system. Before 1982, it was thought that viruses could not be made to fit Ernst Mayr's reproductive concept of species, and so were not amenable to such treatment. In 1982, the ICTV started to define a species as "a cluster of strains" with unique identifying qualities. In 1991, the more specific principle that a virus species is a polythetic class of viruses that constitutes a replicating lineage and occupies a particular ecological niche was adopted.[3]
As at 2021 (the latest edition of the ICVCN), the ICTV definition of species states: "A species is the lowest taxonomic level in the hierarchy approved by the ICTV. A species is a
Below species rank (named viruses/virus strains/isolates)
Many individually named viruses (sometimes referred to as "virus strains") exist at below the rank of virus species. The ICVCN gives the examples of blackeye cowpea mosaic virus and peanut stripe virus, which are both classified in the species
ICTV classification
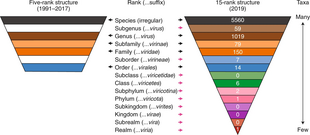
The International Committee on Taxonomy of Viruses began to devise and implement rules for the naming and classification of viruses early in the 1970s, an effort that continues to the present. The ICTV is the only body charged by the International Union of Microbiological Societies with the task of developing, refining, and maintaining a universal virus taxonomy, following the methods set out in the International Code of Virus Classification and Nomenclature.[4][6] The system shares many features with the classification system of cellular organisms, such as taxon structure. However, some differences exist, such as the universal use of italics for all taxonomic names, unlike in the International Code of Nomenclature for algae, fungi, and plants and International Code of Zoological Nomenclature.
Viral classification starts at the level of realm and continues as follows, with the taxonomic suffixes in parentheses:[4]
- Realm (-viria)
In parallel to the system of binomial nomenclature adopted in cellular species, the ICTV has recently (2021) mandated that new virus species be named in a binomial format (genus|| species), and that pre-existing virus species names be progressively replaced with new names in the binomial format.[7] A mid-2023 review of the status of this changeover stated: "...a large number of proposals [concerning virus nomenclature, submitted to the ICTV Executive Committee (EC) for its consideration] renamed existing species for compliance with the recently mandated binomial nomenclature format. As a result, 8,982 out of the current 11,273 species (80%) now have binomial names. The process will be concluded in 2023, with the remaining 2,291 species being renamed."[8]
As of 2021, all levels of taxa except subrealm, subkingdom, and subclass are used. Six realms, one incertae sedis class, 22 incertae sedis families, and two incertae sedis genera are recognized:[9]
Realms:
Incertae sedis classes:
Incertae sedis families:
- Alphasatellitidae
- Ampullaviridae
- Anelloviridae
- Avsunviroidae
- Bartogtaviriformidae
- Bicaudaviridae
- Brachygtaviriformidae
- Clavaviridae
- Fuselloviridae
- Globuloviridae
- Guttaviridae
- Halspiviridae
- Itzamnaviridae
- Ovaliviridae
- Plasmaviridae
- Polydnaviriformidae
- Portogloboviridae
- Pospiviroidae
- Rhodogtaviriformidae
- Spiraviridae
- Thaspiviridae
- Tolecusatellitidae
Incertae sedis genera:
Structure-based virus classification
It has been suggested that similarity in virion assembly and structure observed for certain viral groups infecting hosts from different domains of life (e.g., bacterial tectiviruses and eukaryotic adenoviruses or prokaryotic Caudovirales and eukaryotic herpesviruses) reflects an evolutionary relationship between these viruses.[10] Therefore, structural relationship between viruses has been suggested to be used as a basis for defining higher-level taxa – structure-based viral lineages – that could complement the ICTV classification scheme of 2010.[11]
The ICTV has gradually added many higher-level taxa using relationships in protein folds. All four realms defined in the 2019 release are defined by the presence of a protein of a certain structural family.[12]
Baltimore classification
Baltimore classification (first defined in 1971) is a classification system that places viruses into one of seven groups depending on a combination of their nucleic acid (DNA or RNA), strandedness (single-stranded or double-stranded), sense, and method of replication. Named after David Baltimore, a Nobel Prize-winning biologist, these groups are designated by Roman numerals. Other classifications are determined by the disease caused by the virus or its morphology, neither of which are satisfactory due to different viruses either causing the same disease or looking very similar. In addition, viral structures are often difficult to determine under the microscope. Classifying viruses according to their genome means that those in a given category will all behave in a similar fashion, offering some indication of how to proceed with further research. Viruses can be placed in one of the seven following groups:[13]
- I: Poxviruses)
- II: Parvoviruses)
- III: Reoviruses)
- IV:Togaviruses)
- V: Rhabdoviruses)
- VI: ssRNA-RT viruses (+ strand or sense) RNA with DNA intermediate in life-cycle (e.g. Retroviruses)
- VII: Hepadnaviruses)

DNA viruses
Viruses with a DNA genome, except for the DNA reverse transcribing viruses, are members of three of the four recognized viral realms: Duplodnaviria, Monodnaviria, and Varidnaviria. But the incertae sedis order Ligamenvirales, and many other incertae sedis families and genera, are also used to classify DNA viruses. The domains Duplodnaviria and Varidnaviria consist of double-stranded DNA viruses; other double-stranded DNA viruses are incertae sedis. The domain Monodnaviria consists of single-stranded DNA viruses that generally encode a HUH endonuclease; other single-stranded DNA viruses are incertae sedis.[14]
- Group I: viruses possess double-stranded DNA. Viruses that cause chickenpox and herpes are found here.
- Group II: viruses possess single-stranded DNA.
| Virus family | Examples (common names) | Virion naked/enveloped |
Capsid symmetry |
Nucleic acid type | Group |
|---|---|---|---|---|---|
| 1. Adenoviridae | Canine hepatitis virus, Some types of the common cold
|
Naked | Icosahedral | ds | I |
| 2. Papovaviridae
|
JC virus, HPV
|
Naked | Icosahedral | ds circular | I |
| 3. Parvoviridae | Human parvovirus B19, canine parvovirus | Naked | Icosahedral | ss | II |
| 4. Herpesviridae | varicella-zoster virus, cytomegalovirus, Epstein–Barr virus
|
Enveloped | Icosahedral | ds | I |
| 5. Poxviridae | vaccinia virus
|
Complex coats | Complex | ds | I |
| 6. Anelloviridae | Torque teno virus | Naked | Icosahedral | ss circular | II |
| 7. Pleolipoviridae | HHPV1, HRPV1 | Enveloped | ss/ds linear/circular | I/II |
RNA viruses
All viruses that have an RNA genome, and that encode an RNA-dependent RNA polymerase (RdRp), are members of the kingdom Orthornavirae, within the realm Riboviria.[15]
- Group III: viruses possess double-stranded RNA genomes, e.g. rotavirus.
- Group IV: viruses possess positive-sense single-stranded RNA genomes. Many well known viruses are found in this group, including the virus.
- Group V: viruses possess negative-sense single-stranded RNA genomes. .
| Virus Family | Examples (common names) | Capsid naked/enveloped |
Capsid Symmetry |
Nucleic acid type | Group |
|---|---|---|---|---|---|
| 1. Reoviridae
|
Reovirus, rotavirus
|
Naked | Icosahedral | ds | III |
| 2. Picornaviridae
|
Enterovirus, rhinovirus, hepatovirus, cardiovirus, aphthovirus, poliovirus, parechovirus, erbovirus, kobuvirus, teschovirus, coxsackie | Naked | Icosahedral | ss | IV |
| 3. Caliciviridae | Norwalk virus | Naked | Icosahedral | ss | IV |
| 4. Togaviridae
|
Eastern equine encephalitis, Chikungunya | Enveloped | Icosahedral | ss | IV |
| 5. Arenaviridae
|
Lymphocytic choriomeningitis virus, Lassa fever
|
Enveloped | Complex | ss(−) | V |
| 6. Flaviviridae | Dengue virus, hepatitis C virus, yellow fever virus, Zika virus | Enveloped | Icosahedral | ss | IV |
| 7. Orthomyxoviridae | isavirus, thogotovirus
|
Enveloped | Helical | ss(−) | V |
| 8. Paramyxoviridae | canine distemper virus
|
Enveloped | Helical | ss(−) | V |
| 9. Bunyaviridae
|
Sin nombre virus
|
Enveloped | Helical | ss(−) | V |
| 10. Rhabdoviridae | Vesicular stomatitis
|
Enveloped | Helical | ss(−) | V |
| 11. Filoviridae | Ebola virus, Marburg virus
|
Enveloped | Helical | ss(−) | V |
| 12. Coronaviridae | Severe acute respiratory syndrome coronavirus 2
|
Enveloped | Helical | ss | IV |
| 13. Astroviridae
|
Astrovirus | Naked | Icosahedral | ss | IV |
| 14. Bornaviridae | Borna disease virus | Enveloped | Helical | ss(−) | V |
| 15. Arteriviridae | Arterivirus, equine arteritis virus
|
Enveloped | Icosahedral | ss | IV |
| 16. Hepeviridae | Hepatitis E virus
|
Naked | Icosahedral | ss | IV |
Reverse transcribing viruses
All viruses that encode a reverse transcriptase (also known as RT or RNA-dependent DNA polymerase) are members of the class Revtraviricetes, within the phylum Arterviricota, kingdom Pararnavirae, and realm Riboviria. The class Blubervirales contains the single family Hepadnaviridae of DNA RT (reverse transcribing) viruses; all other RT viruses are members of the class Ortervirales.[16]
- Group VI: viruses possess single-stranded RNA viruses that replicate through a DNA intermediate. The retroviruses are included in this group, of which HIV is a member.
- Group VII: viruses possess double-stranded DNA genomes and replicate using reverse transcriptase. The hepatitis B virus can be found in this group.
| Virus Family | Examples (common names) | Capsid naked/enveloped |
Capsid Symmetry |
Nucleic acid type | Group |
|---|---|---|---|---|---|
| 1. Retroviridae | HIV | Enveloped | dimer RNA | VI | |
| 2. Caulimoviridae | Cacao swollen-shoot virus (CSSV)
|
Naked | VII | ||
| 3. Hepadnaviridae | Hepatitis B virus | Enveloped | Icosahedral | circular, partially ds | VII |
Historical systems
Holmes classification
Holmes (1948) used a Linnaean taxonomy with binomial nomenclature to classify viruses into 3 groups under one order, Virales. They are placed as follows:
- Group I: Phaginae (attacks bacteria)
- Group II: Phytophaginae (attacks plants)
- Group III: Zoophaginae (attacks animals)
The system was not accepted by others due to its neglect of morphological similarities.[17]
Subviral agents
Viroids and virus-dependent agents
Viroids
- Family Avsunviroidae[21]
- Genus Avsunviroid; type species: Avocado sunblotch viroid
- Genus Pelamoviroid; type species: Peach latent mosaic viroid
- Genus Elaviroid; type species: Eggplant latent viroid
- Genus
- Family Pospiviroidae[22]
- Genus Pospiviroid; type species: Potato spindle tuber viroid
- Genus Hostuviroid; type species: Hop stunt viroid
- Genus Coconut cadang-cadang viroid
- Genus Apscaviroid; type species: Apple scar skin viroid
- Genus Coleviroid; type species: Coleus blumei viroid 1
Satellites
Satellites depend on co-infection of a host cell with a helper virus for productive multiplication. Their nucleic acids have substantially distinct nucleotide sequences from either their helper virus or host. When a satellite subviral agent encodes the coat protein in which it is encapsulated, it is then called a satellite virus.
Satellite-like nucleic acids resemble satellite nucleic acids, in that they replicate with the aid of helper viruses. However they differ in that they can encode functions that can contribute to the success of their helper viruses; while they are sometimes considered to be genomic elements of their helper viruses, they are not always found within their helper viruses.[18]
- Satellite viruses[23]
- Single-stranded RNA satellite viruses
- (unnamed family)
- Aumaivirus – Maize white line mosaic satellite virus
- Papanivirus – Panicum mosaic satellite virus
- Tobacco mosaic satellite virus
- Tobacco necrosis satellite virus
- Family Sarthroviridae
- Macrobrachium satellite virus 1(extra small virus)
- (unnamed genus) – Nilaparvata lugens commensal X virus
- (unnamed genus) – Chronic bee-paralysis satellite virus
- (unnamed family)
- Double-stranded DNA satellite viruses
- Family Lavidaviridae– Virophages
- Family
- Single-stranded DNA satellite viruses
- Genus Dependoparvovirus – Adeno-associated virus group
- Single-stranded RNA satellite viruses
- Satellite nucleic acids
- Single-stranded satellite DNAs
- Family Alphasatellitidae(encoding a replication initiator protein)
- Family Tolecusatellitidae (encoding a pathogenicity determinant βC1)
- Family
- Double-stranded satellite RNAs
- Single-stranded satellite RNAs
- Subgroup 1: Large satellite RNAs
- Subgroup 2: Small linear satellite RNAs
- Subgroup 3: Circular satellite RNAs (virusoids)
- Genus Deltavirus
- Polerovirus-associated RNAs
- Satellite-like RNA
- Satellite-like DNA
- Single-stranded satellite DNAs
Defective interfering particles
Defective interfering particles are defective viruses that have lost their ability to replicate except in the presence of a helper virus, which is normally the parental virus. They can also interfere with the helper virus.
- Defective interfering particles (RNA)
- Defective interfering particles (DNA)
Viriforms
Viriforms are a polyphyletic category of
See also
Notes
- ^ "Virus Taxonomy: 2022 v3 Release". ictv.global. International Committee on Taxonomy of Viruses. Retrieved 5 January 2024.
- PMID 34468181.
- ^ Alimpiev E (March 15, 2019). Rethinking the Virus Species Concept (PDF). Archived (PDF) from the original on 2020-09-22.
- ^ a b c d "The International Code of Virus Classification and Nomenclature (ICVCN), March 2021 edition". International Committee on Taxonomy of Viruses. Retrieved 6 January 2024.
- PMID 32123347.
- PMID 29040670.
- PMID 34231026.
- PMID 37296227.
- ^ "Virus Taxonomy: 2021 Release". ictv.global. International Committee on Taxonomy of Viruses. Retrieved 26 January 2023.
- PMID 12798226.
- PMID 20926569.
- ^ "Virus Taxonomy: 2019 Release". talk.ictvonline.org. International Committee on Taxonomy of Viruses. Retrieved 26 April 2020.
- ^ "Baltimore Classification of Viruses". www.web-books.com. Retrieved 2023-01-02.
- ^ "Virus Taxonomy: 2019 Release". talk.ictvonline.org. International Committee on Taxonomy of Viruses. Retrieved 26 April 2020.
- ^ "Virus Taxonomy: 2019 Release". talk.ictvonline.org. International Committee on Taxonomy of Viruses. Retrieved 25 April 2020.
- ^ Koonin EV, Dolja VV, Krupovic M, Varsani A, Wolf YI, Yutin N, Zerbini M, Kuhn JH. "Proposal: Create a megataxonomic framework, filling all principal taxonomic ranks, for realm Riboviria". International Committee on Taxonomy of Viruses (ICTV). Retrieved 2020-05-21.
{{cite web}}: CS1 maint: multiple names: authors list (link) - PMC 7157452.
- S2CID 80872659.
- ^ TaxoProp 2015.002aG
- ^ Di Serio F, Li SF, Matoušek J, Owens RA, Pallás V, Randles JW, Sano T, Verhoeven JT, Vidalakis G, Flores R (June 2020). "Family: Avsunviroidae". The 10th ICTV Report on Virus Classification and Taxon Nomenclature. ICTV. Retrieved 16 January 2023.
- ^ Di Serio F, Owens RA, Li SF, Matoušek J, Pallás V, Randles JW, Sano T, Verhoeven JT, Vidalakis G, Flores R (November 2020). "Family: Pospiviroidae". The 10th ICTV Report on Virus Classification and Taxon Nomenclature. ICTV. Retrieved 16 January 2023.
- PMID 26446887.
- PMID 36830658.
- PMID 36381234.