Yersinia pestis
| Yersinia pestis | |
|---|---|
| |
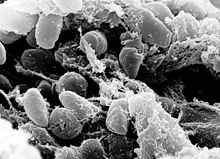
| A scanning electron micrograph depicting a mass of Yersinia pestis bacteria in the foregut of an infected flea | |
| Scientific classification | |
| Domain: | Bacteria |
| Phylum: | Pseudomonadota |
| Class: | Gammaproteobacteria |
| Order: | Enterobacterales |
| Family: | Yersiniaceae |
| Genus: | Yersinia |
| Species: | Y. pestis
|
| Binomial name | |
| Yersinia pestis (Lehmann & Neumann, 1896)
van Loghem, 1944 | |
| Synonyms | |
| |
Yersinia pestis (Y. pestis; formerly Pasteurella pestis) is a gram-negative, non-motile, coccobacillus bacterium without spores that is related to both Yersinia enterocolitica and Yersinia pseudotuberculosis, the pathogen from which Y. pestis evolved[1][2] and responsible for the Far East scarlet-like fever. It is a facultative anaerobic organism that can infect humans via the Oriental rat flea (Xenopsylla cheopis).[3] It causes the disease plague, which caused the Plague of Justinian and the Black Death, the deadliest pandemic in recorded history. Plague takes three main forms: pneumonic, septicemic, and bubonic. Yersinia pestis is a parasite of its host, the rat flea, which is also a parasite of rats, hence Y. pestis is a hyperparasite.
Y. pestis was discovered in 1894 by Alexandre Yersin, a Swiss/French physician and bacteriologist from the Pasteur Institute, during an epidemic of the plague in Hong Kong.[4][5] Yersin was a member of the Pasteur school of thought. Kitasato Shibasaburō, a Japanese bacteriologist who practised Koch's methodology, was also engaged at the time in finding the causative agent of the plague.[6] However, Yersin actually linked plague with a bacillus, initially named Pasteurella pestis; it was renamed Yersinia pestis in 1944.
Every year, between one thousand and two thousand cases of the plague are still reported to the World Health Organization.[7] With proper antibiotic treatment, the prognosis for victims is much better than before antibiotics were developed. A five- to six-fold increase in cases occurred in Asia during the time of the Vietnam War, possibly due to the disruption of ecosystems and closer proximity between people and animals. The plague is now commonly found in sub-Saharan Africa and Madagascar, areas that now account for over 95% of reported cases. The plague also has a detrimental effect on non-human mammals;[8] in the United States, these include the black-tailed prairie dog and the endangered black-footed ferret.
General features
Y. pestis is a non-motile
Genome and proteome
Genome
Several complete
| KIM | CO92 | 91001 | |
|---|---|---|---|
| length (bp) | 4,600,755 | 4,653,728 | 4,595,065 |
| proteins encoded | 4,198 | 4,012 | 4,037 |
| pseudogenes | 54 | 149 | 141 |
| tRNAs | 73 | 70 | 72 |
Plasmids
Like Y. pseudotuberculosis and Y. enterocolitica, Y. pestis is host to the
Proteome
A comprehensive and comparative proteomics analysis of Y. pestis strain KIM was performed in 2006.[17] The analysis focused on the transition to a growth condition mimicking growth in host cells.[citation needed]
Small noncoding RNA
Numerous bacterial small noncoding RNAs have been identified to play regulatory functions. Some can regulate the virulence genes. Some 63 novel putative sRNAs were identified through deep sequencing of the Y. pestis sRNA-ome. Among them was Yersinia-specific (also present in Y. pseudotuberculosis and Y. enterocolitica) Ysr141 (Yersinia small RNA 141). Ysr141 sRNA was shown to regulate the synthesis of the type III secretion system (T3SS) effector protein YopJ.[18] The Yop-Ysc T3SS is a critical component of virulence for Yersinia species.[19] Many novel sRNAs were identified from Y. pestis grown in vitro and in the infected lungs of mice suggesting they play role in bacterial physiology or pathogenesis. Among them sR035 predicted to pair with SD region and transcription initiation site of a thermo-sensitive regulator ymoA, and sR084 predicted to pair with fur, ferric uptake regulator.[20]
Pathogenesis and immunity

In the urban and sylvatic (forest) cycles of Y. pestis, most of the spreading occurs between rodents and fleas. In the sylvatic cycle, the rodent is wild, but in the urban cycle, the rodent is primarily the brown rat (Rattus norvegicus). In addition, Y. pestis can spread from the urban environment and back. Transmission to humans is usually through the bite of infected fleas. If the disease has progressed to the pneumonic form, humans can spread the bacterium to others by coughing, vomiting, and possibly sneezing.[citation needed]
Mammals as hosts
Several species of rodents serve as the main reservoir for Y. pestis in the environment. In the steppes, the natural reservoir is believed to be principally the marmot. In the western United States, several species of rodents are thought to maintain Y. pestis. However, the expected disease dynamics have not been found in any rodent. Several species of rodents are known to have a variable resistance, which could lead to an asymptomatic carrier status.[21] Evidence indicates fleas from other mammals have a role in human plague outbreaks.[22]
The lack of knowledge of the dynamics of plague in mammal species is also true among susceptible rodents such as the black-tailed prairie dog (Cynomys ludovicianus), in which plague can cause colony collapse, resulting in a massive effect on prairie food webs.[23] However, the transmission dynamics within prairie dogs do not follow the dynamics of blocked fleas; carcasses, unblocked fleas, or another vector could possibly be important, instead.[24]
The CO92 strain was isolated from a patient who died from pneumonia and who contracted the infection from an infected cat.[12]
In other regions of the world, the reservoir of the infection is not clearly identified, which complicates prevention and early-warning programs. One such example was seen in a 2003 outbreak in Algeria.[25]
Fleas as vector
The transmission of Y. pestis by fleas is well characterized.
The hemin storage system plays an important role in the transmission of Y. pestis back to a mammalian host.[27] While in the insect vector, proteins encoded by hemin storage system genetic loci induce biofilm formation in the proventriculus, a valve connecting the midgut to the esophagus.[28][29] The presence of this biofilm seems likely to be required for stable infection of the flea.[30] Aggregation in the biofilm inhibits feeding, as a mass of clotted blood and bacteria forms (referred to as "Bacot's block" after entomologist A.W. Bacot, the first to describe this phenomenon).[31] Transmission of Y. pestis occurs during the futile attempts of the flea to feed. Ingested blood is pumped into the esophagus, where it dislodges bacteria lodged in the proventriculus, which is regurgitated back into the host circulatory system.[31]
In humans and other susceptible hosts
In addition, the type-III secretion system (T3SS) allows Y. pestis to inject proteins into macrophages and other immune cells. These T3SS-injected proteins, called Yersinia outer proteins (Yops), include Yop B/D, which form pores in the host cell membrane and have been linked to
Y. pestis
]YopE functions as a
YopJ is an acetyltransferase that binds to a conserved α-helix of MAPK kinases.[36] YopJ acetylates MAPK kinases at serines and threonines that are normally phosphorylated during activation of the MAP kinase cascade.[37][38] YopJ is activated in eukaryotic cells by interaction with target cell phytic acid (IP6).[39] This disruption of host cell protein kinase activity causes apoptosis of macrophages, and this is proposed to be important for the establishment of infection and for evasion of the host immune response. YopO is a protein kinase also known as Yersinia protein kinase A (YpkA). YopO is a potent inducer of human macrophage apoptosis.[40]
It has also been suggested that a bacteriophage – Ypφ – may have been responsible for increasing the virulence of this organism.[41]
Depending on which form of the plague infects the individual, the plague develops a different illness; however, the plague overall affects the host cell's ability to communicate with the immune system, hindering the body bringing phagocytic cells to the area of infection.
Y. pestis is a versatile killer. In addition to rodents and humans, it is known to have killed camels, chickens, and pigs.[42] Domestic dogs and cats are susceptible to plague, as well, but cats are more likely to develop illness when infected. In either, the symptoms are similar to those experienced by humans, and can be deadly to the animal. People can be exposed by coming into contact with an infected animal (dead or alive), or inhaling infectious droplets that a sick dog or cat has coughed into the air.[43][44]
Immunity
A formalin-inactivated vaccine was available in the United States for adults in 1993[45] at high risk of contracting the plague until removal from the market by the Food and Drug Administration. It was of limited effectiveness and could cause severe inflammation. Experiments with genetic engineering of a vaccine based on F1 and V antigens are underway and show promise. However, bacteria lacking antigen F1 are still virulent, and the V antigens are sufficiently variable such that vaccines composed of these antigens may not be fully protective.[46] The United States Army Medical Research Institute of Infectious Diseases has found that an experimental F1/V antigen-based vaccine protects crab-eating macaques, but fails to protect African green monkey species.[47] A systematic review by the Cochrane Collaboration found no studies of sufficient quality to make any statement on the efficacy of the vaccine.[48]
Isolation and identification

In 1894, two bacteriologists, Alexandre Yersin of Switzerland and Kitasato Shibasaburō of Japan, independently isolated in Hong Kong the bacterium responsible for the 1894 Hong Kong plague. Though both investigators reported their findings, a series of confusing and contradictory statements by Kitasato eventually led to the acceptance of Yersin as the primary discoverer of the organism. Yersin named it Pasteurella pestis in honor of the Pasteur Institute, where he worked. In 1967, it was moved to a new genus and renamed Yersinia pestis in his honor. Yersin also noted that rats were affected by plague not only during plague epidemics, but also often preceding such epidemics in humans and that plague was regarded by many locals as a disease of rats; villagers in China and India asserted that when large numbers of rats were found dead, plague outbreaks soon followed.[citation needed]
In 1898, French scientist
Three main strains are recognised: Y. p. antiqua, which caused a plague pandemic in the sixth century; Y. p. medievalis, which caused the Black Death and subsequent epidemics during the second pandemic wave; and Y. p. orientalis, which is responsible for current plague outbreaks.[52]
21st century
On January 15, 2018, researchers at the University of Oslo and the University of Ferrara suggested that humans and their parasites (most likely fleas and lice at the time) were the biggest carriers of the plague.[53][54]
Ancient DNA evidence
In 2010, researchers in Germany definitively established, using PCR evidence from samples obtained from Black Death victims, that Y. pestis was the cause of the medieval Black Death.[55]
In 2011, the first genome of Y. pestis isolated from Black Death victims was published, and concluded that this medieval strain was ancestral to most modern forms of Y. pestis.[56]
In 2015,
DNA evidence published in 2015 indicates Y. pestis infected humans 5,000 years ago in Bronze Age Eurasia,[57] but genetic changes that made it highly virulent did not occur until about 4,000 years ago.[60] The highly virulent version capable of transmission by fleas through rodents, humans, and other mammals was found in two individuals associated with the Srubnaya culture from the Samara region in Russia from around 3,800 years ago and an Iron Age individual from Kapan, Armenia, from around 2,900 years ago.[60][57] This indicates that at least two lineages of Y. pestis were circulating during the Bronze Age in Eurasia.[60] The Y. pestis bacterium has a relatively large number of nonfunctioning genes and three "ungainly" plasmids, suggesting an origin less than 20,000 years ago.[42] One such strain has been identified from about 4000 BP (the "LNBA lineage" (Late Neolithic and Bronze Age lineage)) in western Britain, indicating that this highly transmissible form spread from Eurasia to the far north-western edges of Europe.[61]
On September 8, 2016, the Y. pestis bacterium was identified from DNA in teeth found at a Crossrail building site in London. The human remains were found to be victims of the Great Plague of London, which lasted from 1665 to 1666.[62]
In 2021, researchers found a 5,000-year-old victim of Y. pestis, the world's oldest-known, in hunter-gatherer remains in the modern Latvian and Estonian border area.[63]
Events
Between 1970 and 2020, 496 cases were reported in the United States. Cases have been found predominantly in New Mexico, Arizona, Colorado, California, Oregon, and Nevada.[64]
In 2008, plague was commonly found in sub-Saharan Africa and Madagascar, areas that accounted for over 95% of the reported cases.[8]
In September 2009, the death of
On November 3, 2019, two cases of pneumonic plague were diagnosed at a hospital in Beijing's
In July 2020, officials increased precautions after a case of bubonic plague was confirmed in Bayannur, a city in China's Inner Mongolia autonomous region. The patient was quarantined and treated. According to China's Global Times, a second suspected case was also investigated, and a level 3 alert was issued, in effect until the end of the year. It forbade hunting and eating of animals that could carry plague, and called on the public to report suspected cases.[69]
References
- PMID 10570195.
- S2CID 21267985.
- ISBN 978-0-8385-8529-0.
- ^ Yersin, Alexandre (1894). "La peste bubonique à Hong-Kong". Annales de l'Institut Pasteur (in French). 8: 662–67.
- PMID 7959865.
- S2CID 31767623.
- ^ "Plague FAQ". CDC. 15 November 2021.
- ^ a b "The Plague", Centers for Disease Control and Prevention, October 2017
 This article incorporates text from this source, which is in the public domain.
This article incorporates text from this source, which is in the public domain.
- ^ ISBN 978-0-9631172-1-2.
- ISBN 978-0-387-25499-9.
- PMID 12142430.
- ^ PMID 11586360.
- PMID 16740952.
- ^ PMID 15368893.
- ^ S2CID 34770234.
- ^ S2CID 39881239.
- PMID 17081052.
- PMID 24532772.
- PMID 9841674.
- PMID 24040259.
- PMID 13453634.
- PMID 908848.
- S2CID 9716200.
- PMID 16603630.
- PMID 18257987.
- PMID 16182593.
- S2CID 37512575.
- S2CID 36769530.
- PMID 15272407.
- PMID 16428415.
- ^ PMID 18453279.
- ^ Salyers AA, Whitt DD (2002). Bacterial Pathogenesis: A Molecular Approach (2nd ed.). ASM Press. pp. 207–212.
- PMID 15847602.
- PMID 19221593.
- PMID 18193942.
- PMID 18167536.
- S2CID 13101320.
- PMID 17116858.
- PMID 20430892.
- PMID 17475872.
- S2CID 30862265.
- ^ ISBN 978-0007150694.
- ^ "Cats – Healthy Pets Healthy People". Centers for Disease Control and Prevention. 2016-05-13. Retrieved 2016-11-25.
- ^ "Dogs – Healthy Pets Healthy People". Centers for Disease Control and Prevention. 2020-02-21. Retrieved 2020-06-30.
- ^ Sheet, The Pink (21 Nov 1994). "GREER LABORATORIES BUBONIC PLAGUE VACCINE APPROVED". Pink Sheet.
- PMID 12009274.
- ^ Pitt ML (13 October 2004). Non-human primates as a model for pneumonic plague (PDF). Animals Models and Correlates of Protection for Plague Vaccines Workshop, Gaithersburg, Maryland. Center for Biologics Evaluation and Research (Food and Drug Administration, Department of Health and Human Resources). pp. 222–248. Archived from the original (PDF) on 25 December 2004.
- PMID 10796565. Art. No. CD000976.
- ^ "The Plague". Association Amicale Sante Navale et d'Outremer. Archived from the original on 4 September 2012.
- ^ "On This Day: San Francisco Bubonic Plague Outbreak Begins". Finding Dulcinea. 27 May 2011. Retrieved 2017-11-25.
- ^ Chase, M. (2004). The Barbary Plague: The Black Death in Victorian San Francisco. Random House Trade Paperbacks.
- PMID 10570195.
- ^ "Don't blame the rats: Human fleas and lice likely spread Black Death". CBC News.
- PMID 29339508.
- PMID 20949072.
- PMID 21993626.
- ^ PMID 26496604.license.
 This article contains quotations from this source, which is available under the Creative Commons Attribution 4.0 International (CC BY 4.0)
This article contains quotations from this source, which is available under the Creative Commons Attribution 4.0 International (CC BY 4.0) - ISSN 0362-4331. Retrieved 2020-04-16.
- ^ PMID 30528431.
- ^ PMID 29884871.license.
 This article contains quotations from this source, which is available under the Creative Commons Attribution 4.0 International (CC BY 4.0)
This article contains quotations from this source, which is available under the Creative Commons Attribution 4.0 International (CC BY 4.0) - PMID 37253742.
- ^ "DNA of bacteria responsible for London Great Plague of 1665 identified for first time". 29 December 2022. Archived from the original on September 10, 2016.
- S2CID 235697166.
- ^ CDC (2022-11-16). "Plague surveillance | CDC". Centers for Disease Control and Prevention. Retrieved 2023-02-11.
- ^ Sadovi, Carlos (2009-09-19). "U. of C. researcher dies after exposure to plague bacteria". Chicago Breaking News Center. Retrieved 2010-03-03.
- ^ Randall T (Feb 25, 2011). "Plague Death Came Within Hours, Spurred by Scientist's Medical Condition".
- ^ ISSN 0362-4331. Retrieved 2019-11-13.
- ^ "Bubonic plague: Third case reported in China". Medical News Today. 19 November 2019.
- ^ "China bubonic plague: Inner Mongolia takes precautions after case". BBC News. 2020-07-06. Retrieved 2020-07-06.
External links
- A list of variant strains and information on synonyms (and much more) is available through the NCBI taxonomy browser.
- CDC's Home page for Plague
- IDSA's resource page on Plague: Current, comprehensive information on pathogenesis, microbiology, epidemiology, diagnosis, and treatment
- Plague (Yersinia Pestis) at Drugs.com
- Wyndham Lathem speaking on "From Mild to Murderous: How Yersinia pestis Evolved to Cause Pneumonic Plague"
