De Bruijn graph: Difference between revisions
144,422 edits Importing Wikidata short description: "Node-link graph representing overlaps between sequences of symbols" (Shortdesc helper) |
created "Implementations" section Tag: Reverted |
||
| Line 27: | Line 27: | ||
The Bernoulli map (also called the [[2x mod 1 map]] for ''m'' = 2) is an [[ergodic]] dynamical system, which can be understood to be a single [[shift operator|shift]] of a [[p-adic|m-adic number]].<ref name="Leroux2002"/> The trajectories of this dynamical system correspond to walks in the De Bruijn graph, where the correspondence is given by mapping each real ''x'' in the interval [0,1) to the vertex corresponding to the first ''n'' digits in the [[Radix|base]]-''m'' representation of ''x''. Equivalently, walks in the De Bruijn graph correspond to trajectories in a one-sided [[subshift of finite type]]. |
The Bernoulli map (also called the [[2x mod 1 map]] for ''m'' = 2) is an [[ergodic]] dynamical system, which can be understood to be a single [[shift operator|shift]] of a [[p-adic|m-adic number]].<ref name="Leroux2002"/> The trajectories of this dynamical system correspond to walks in the De Bruijn graph, where the correspondence is given by mapping each real ''x'' in the interval [0,1) to the vertex corresponding to the first ''n'' digits in the [[Radix|base]]-''m'' representation of ''x''. Equivalently, walks in the De Bruijn graph correspond to trajectories in a one-sided [[subshift of finite type]]. |
||
| ⚫ | |||
Embeddings resembling this one can be used to show that the binary De Bruijn graphs have [[queue number]] 2<ref name=HeathRosenberg/> and that they have [[book thickness]] at most 5.<ref name=Obrenic/> |
Embeddings resembling this one can be used to show that the binary De Bruijn graphs have [[queue number]] 2<ref name=HeathRosenberg/> and that they have [[book thickness]] at most 5.<ref name=Obrenic/> |
||
== Implementations == |
|||
[[File:DeBruijnGraph24.svg|thumb|Show the 4-dimension, 2-symbol De Bruijn Graph made in the [[Wolfram Language]].]] |
|||
In the [[Wolfram Language]], a De Bruijn Graph with ''n'' dimensions and ''m'' symbols can be generated with the function {{code|DeBruijnGraph[m,n]}}.<ref>{{cite web |last1=Wolfram Research |title=DeBruijnGraph—Wolfram Language Documentation |url=http://reference.wolfram.com/language/ref/DeBruijnGraph.html |website=reference.wolfram.com}}</ref> For example, {{code|DeBruijnGraph[2,4]}} creates the graph shown to the right: |
|||
== Uses == |
== Uses == |
||
| ⚫ | |||
* Some [[grid network]] topologies are De Bruijn graphs. |
* Some [[grid network]] topologies are De Bruijn graphs. |
||
* The [[distributed hash table]] protocol [[Koorde]] uses a De Bruijn graph. |
* The [[distributed hash table]] protocol [[Koorde]] uses a De Bruijn graph. |
||
Revision as of 20:07, 5 March 2021
In graph theory, an n-dimensional De Bruijn graph of m symbols is a directed graph representing overlaps between sequences of symbols. It has mn vertices, consisting of all possible length-n sequences of the given symbols; the same symbol may appear multiple times in a sequence. If we have the set of m symbols then the set of vertices is:
If one of the vertices can be expressed as another vertex by shifting all its symbols by one place to the left and adding a new symbol at the end of this vertex, then the latter has a directed edge to the former vertex. Thus the set of arcs (that is, directed edges) is
Although De Bruijn graphs are named after Nicolaas Govert de Bruijn, they were discovered independently by both De Bruijn[1] and I. J. Good.[2] Much earlier, Camille Flye Sainte-Marie[3] implicitly used their properties.
Properties
- If then the condition for any two vertices forming an edge holds vacuously, and hence all the vertices are connected forming a total of edges.
- Each vertex has exactly incoming and outgoing edges.
- Each -dimensional De Bruijn graph is the line digraph of the -dimensional De Bruijn graph with the same set of symbols.[4]
- Each De Bruijn graph is Hamiltonian. The Euler cycles and Hamiltonian cycles of these graphs (equivalent to each other via the line graph construction) are De Bruijn sequences.
The line graph construction of the three smallest binary De Bruijn graphs is depicted below. As can be seen in the illustration, each vertex of the -dimensional De Bruijn graph corresponds to an edge of the -dimensional De Bruijn graph, and each edge in the -dimensional De Bruijn graph corresponds to a two-edge path in the -dimensional De Bruijn graph.

Dynamical systems
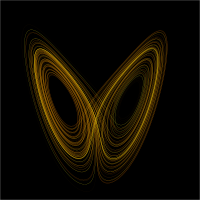
Binary De Bruijn graphs can be drawn (below, left) in such a way that they resemble objects from the theory of
This analogy can be made rigorous: the n-dimensional m-symbol De Bruijn graph is a model of the
The Bernoulli map (also called the
Embeddings resembling this one can be used to show that the binary De Bruijn graphs have
Implementations

In the Wolfram Language, a De Bruijn Graph with n dimensions and m symbols can be generated with the function DeBruijnGraph[m,n].[8] For example, DeBruijnGraph[2,4] creates the graph shown to the right:
Uses

- Some grid network topologies are De Bruijn graphs.
- The distributed hash table protocol Koorde uses a De Bruijn graph.
- In bioinformatics, De Bruijn graphs are used for de novo assembly of sequencing reads into a genome.[9][10][11][12][13]
See also
References
- ^ de Bruijn; N. G. (1946). "A Combinatorial Problem". Koninklijke Nederlandse Akademie V. Wetenschappen. 49: 758–764.
- ^ .
- ^ Flye Sainte-Marie, C. (1894). "Question 48". L'Intermédiaire des Mathématiciens. 1: 107–110.
- ^ Zhang, Fu Ji; Lin, Guo Ning (1987). "On the de Bruijn-Good graphs". Acta Math. Sinica. 30 (2): 195–205.
- ^
Leroux, Philippe (2002). "Coassociative grammar, periodic orbits and quantum random walk over Z". arXiv:quant-ph/0209100.
- ^
Heath, Lenwood S.; MR 1181408.
- ^
Obrenić, Bojana (1993). "Embedding de Bruijn and shuffle-exchange graphs in five pages". MR 1241401.
- ^ Wolfram Research. "DeBruijnGraph—Wolfram Language Documentation". reference.wolfram.com.
- ^
Pevzner, Pavel A.; Tang, Haixu; Waterman, Michael S. (2001). "An Eulerian path approach to DNA fragment assembly". PNAS. 98 (17): 9748–9753. PMID 11504945.
- ^
Pevzner, Pavel A.; Tang, Haixu (2001). "Fragment Assembly with Double-Barreled Data". Bioinformatics. 17 (Suppl 1): S225–S233. PMID 11473013.
- ^
Zerbino, Daniel R.; Birney, Ewan (2008). "Velvet: algorithms for de novo short read assembly using de Bruijn graphs". Genome Research. 18 (5): 821–829. PMID 18349386.
- ^
Chikhi, R; Limasset, A; Jackman, S; Simpson, J; Medvedev, P (2014). "On the representation of de Bruijn graphs". Journal of Computational Biology : A Journal of Computational Molecular Cell Biology. 22 (5): 336–52. PMID 25629448.
- ^
Iqbal, Zamin; Caccamo, Mario; Turner, Isaac; Flicek, Paul; McVean, Gil (2012). "De novo assembly and genotyping of variants using colored de Bruijn graphs". Nature Genetics. 44 (2): 226–32. PMID 22231483.