-
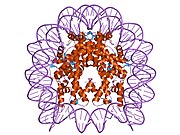
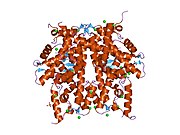
1aoi: COMPLEX BETWEEN NUCLEOSOME CORE PARTICLE (H3,H4,H2A,H2B) AND 146 BP LONG DNA FRAGMENT
-
1eqz: X-RAY STRUCTURE OF THE NUCLEOSOME CORE PARTICLE AT 2.5 A RESOLUTION
-
1f66: 2.6 A CRYSTAL STRUCTURE OF A NUCLEOSOME CORE PARTICLE CONTAINING THE VARIANT HISTONE H2A.Z
-
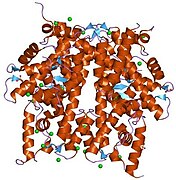
1hq3: CRYSTAL STRUCTURE OF THE HISTONE-CORE-OCTAMER IN KCL/PHOSPHATE
-
1kx3: X-Ray Structure of the Nucleosome Core Particle, NCP146, at 2.0 A Resolution
-
1kx4: X-Ray Structure of the Nucleosome Core Particle, NCP146b, at 2.6 A Resolution
-
1kx5: X-Ray Structure of the Nucleosome Core Particle, NCP147, at 1.9 A Resolution
-
1m18: LIGAND BINDING ALTERS THE STRUCTURE AND DYNAMICS OF NUCLEOSOMAL DNA
-
1m19: LIGAND BINDING ALTERS THE STRUCTURE AND DYNAMICS OF NUCLEOSOMAL DNA
-
1m1a: LIGAND BINDING ALTERS THE STRUCTURE AND DYNAMICS OF NUCLEOSOMAL DNA
-
1p34: Crystallographic Studies of Nucleosome Core Particles containing Histone 'Sin' Mutants
-
1p3a: Crystallographic Studies of Nucleosome Core Particles containing Histone 'Sin' Mutants
-
1p3b: Crystallographic Studies of Nucleosome Core Particles containing Histone 'Sin' Mutants
-
1p3f: Crystallographic Studies of Nucleosome Core Particles containing Histone 'Sin' Mutants
-
1p3g: Crystallographic Studies of Nucleosome Core Particles containing Histone 'Sin' Mutants
-
1p3i: Crystallographic Studies of Nucleosome Core Particles containing Histone 'Sin' Mutants
-
1p3k: Crystallographic Studies of Nucleosome Core Particles containing Histone 'Sin' Mutants
-
1p3l: Crystallographic Studies of Nucleosome Core Particles containing Histone 'Sin' Mutants
-
1p3m: Crystallographic Studies of Nucleosome Core Particles containing Histone 'Sin' Mutants
-
1p3o: Crystallographic Studies of Nucleosome Core Particles containing Histone 'Sin' Mutants
-
1p3p: Crystallographic Studies of Nucleosome Core Particles containing Histone 'Sin' Mutants
-
1s32: Molecular Recognition of the Nucleosomal 'Supergroove'
-
1tzy: Crystal Structure of the Core-Histone Octamer to 1.90 Angstrom Resolution
-
1u35: Crystal structure of the nucleosome core particle containing the histone domain of macroH2A
-
1zbb: Structure of the 4_601_167 Tetranucleosome
-
1zla: X-ray Structure of a Kaposi's sarcoma herpesvirus LANA peptide bound to the nucleosomal core
-
2aro: Crystal Structure Of The Native Histone Octamer To 2.1 Angstrom Resolution, Crystalised In The Presence Of S-Nitrosoglutathione
-
2cv5: Crystal structure of human nucleosome core particle
-
2f8n: 2.9 Angstrom X-ray structure of hybrid macroH2A nucleosomes
-
2fj7: Crystal structure of Nucleosome Core Particle Containing a Poly (dA.dT) Sequence Element
-
2hio: HISTONE OCTAMER (CHICKEN), CHROMOSOMAL PROTEIN
-
2hue: Structure of the H3-H4 chaperone Asf1 bound to histones H3 and H4
-
2io5: Crystal structure of the CIA- histone H3-H4 complex
-
2nzd: Nucleosome core particle containing 145 bp of DNA