Transactivation domain
The transactivation domain or trans-activating domain (TAD) is a transcription factor scaffold domain which contains binding sites for other proteins such as transcription coregulators. These binding sites are frequently referred to as activation functions (AFs).[1] TADs are named after their amino acid composition. These amino acids are either essential for the activity or simply the most abundant in the TAD. Transactivation by the Gal4 transcription factor is mediated by acidic amino acids, whereas hydrophobic residues in Gcn4 play a similar role. Hence, the TADs in Gal4 and Gcn4 are referred to as acidic or hydrophobic, respectively.[2][3][4][5][6][7][8][9]
In general we can distinguish four classes of TADs:[10]
- acidic domains (called also “acid blobs” or “negative noodles", rich in D and E amino acids, present in Gal4, Gcn4 and VP16).[11]
- glutamine-rich domains (contains multiple repetitions like "QQQXXXQQQ", present in SP1)[12]
- proline-rich domains (contains repetitions like "PPPXXXPPP" present in
- isoleucine-rich domains (repetitions "IIXXII", present in NTF-1)[14]
Alternatively, since similar amino acid compositions does not necessary mean similar activation pathways, TADs can be grouped by the process they stimulate, either initiation or elongation.[15]
Acidic/9aaTAD
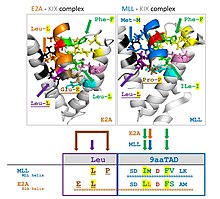
Nine-amino-acid transactivation domain (9aaTAD) defines a domain common to a large superfamily of eukaryotic transcription factors represented by Gal4, Oaf1, Leu3, Rtg3, Pho4, Gln3, Gcn4 in yeast, and by p53, NFAT, NF-κB and VP16 in mammals. The definition largely overlaps with an "acidic" family definition. A 9aaTAD prediction tool is available.[16] 9aaTADs tend to have an associated 3-aa hydrophobic (usually Leu-rich) region immediately to its N-terminal.[17]
9aaTAD transcription factors
The KIX domain of general coactivators Med15(Gal11) interacts with 9aaTAD transcription factors
| Source | 9aaTAD | Peptide-KIX interaction (NMR) |
|---|---|---|
| p53 TAD1 | E TFSD LWKL | LSPEETFSDLWKLPE |
| p53 TAD2 | D DIEQ WFTE | QAMDDLMLSPDDIEQWFTEDPGPD |
| MLL | S DIMD FVLK | DCGNILPSDIMDFVLKNTP |
| E2A | D LLDF SMMF | PVGTDKELSDLLDFSMMFPLPVT |
| Rtg3 | E TLDF SLVT | E2A homolog |
| CREB | R KILN DLSS | RREILSRRPSYRKILNDLSSDAP |
| CREBaB6 | E AILA ELKK | CREB-mutant binding to KIX |
| Gli3 | D DVVQ YLNS | TAD homology to CREB/KIX |
| Gal4 | D DVYN YLFD | Pdr1 and Oaf1 homolog |
| Oaf1 | D LFDY DFLV | DLFDYDFLV |
| Pip2 | D FFDY DLLF | Oafl homolog |
| Pdr1 | E DLYS ILWS | EDLYSILWSDWY |
| Pdr3 | T DLYH TLWN | Pdr1 homolog |
Glutamine-rich
Glutamine (Q)-rich TADs are found in
See also
References
- PMID 12893880.
- PMID 3115591.
- S2CID 4276393.
- PMID 9742254.
- PMID 3115592.
- S2CID 2635634.
- S2CID 40730692.
- ^ PMID 7862116.
- PMID 8381535.
- PMID 2667136.
- S2CID 4276393.
- ^ S2CID 2910480.
- S2CID 22817940.
- PMID 8330738.
- ^ PMID 21557087.
- ^ PMID 17467953.
- ^ PMID 27618436.
- PMID 10611293.
- PMID 9271577.
- PMID 10821850.
- PMID 19220000.
- PMID 12205094.
- S2CID 17268267.
- PMID 12196545.
- PMID 19203263.
- PMID 18241676.
- PMID 16914730.
- PMID 9625762.
- PMID 9343424.
- PMID 17438265.
- PMID 19363523.
- PMID 19217391.
- PMID 19357310.
- PMID 18250150.
- PMID 11258797.
- S2CID 19287634.
- S2CID 5002125.
- PMID 3062377.
- PMID 2685570.
- PMID 11073972.
- PMID 12084920.
- PMID 12665580.
- PMID 12769853.
- PMID 15866178.
- PMID 17919657.
- PMID 17020582.
- PMID 12753200.
- S2CID 882139.
- S2CID 205212715.
- PMID 19056732.
- PMID 12501245.
- PMID 16118188.
- PMID 27515574.
