User:Rkelly66/sandbox
Chromomere
A chromomere, also known as an idiomere, is one of the serially aligned beads or granules of a eukaryotic chromosome, resulting from local coiling of a continuous DNA thread.[1] In areas of chromatin with the absence of transcription, condensing of DNA and protein complexes will result in the formation of chromomeres.[1] It is visible on a chromosome during the prophase of meiosis and mitosis. Giant banded (Polytene) chromosomes resulting from the replication of the chromosomes and the synapsis of homologs without cell division is a process called endomitosis. These chromosomes consist of more than 1000 copies of the same chromatid that are aligned and produce alternating dark and light bands when stained. The dark bands are the chromomere.
It is unknown when chromomeres first appear on the chromosome. Chromomeres can be observed best when chromosomes are highly condensed.[1]The chromomeres are present during leptotene phase of prophase I during meiosis. During zygotene phase of prophase I, the chromomeres of homologs align with each other to form homologous rough pairing (homology searching). These chromomeres helps provide a unique identity for each homologous pairs.
There are more than 2000 chromomeres on 20 chromosomes of maize.
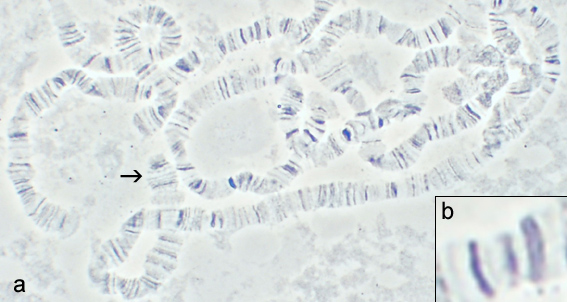
Physical Properties
Chromomeres are organized in a discontinuous linear pattern along the condensed chromosomes (pachytene chromosomes) during the prophase stage of
transcription.[1]
Functions
Chromomeres are known as the structural subunit of a chromosome.The arrangement of chromomere structure can aid in control of gene expression. [2]
Maps of chromomeres can be made for use in genetic and evolutionary studies. Chromomeric maps can be used to locate the exact position of genes on a chromosome. Chromomeric maps can be used to analyze chromosome aberrations, and find correlations between the aberrations and their effects on genes near breakpoints.[2]
Lampbrush Chromosomes and Chromomeres
Chromomeres display different properties and behaviours when associated with
Methyl-CpG-binding protein 2 (MeCP2) is a protein that binds to methylated DNA. MeCP2 has been found to associate most strongly with transcriptionally inactive chromomere domains. MeCP2 binding patterns to chromomere domains are proportional to the density of chromomeric DNA found on a chromosome. The binding of MeCP2 to chromomeric regions represents regions of chromatin that are highly dynamic on the lampbrush chromosome.[5]
References
- ^ . Retrieved 2015-09-24.
- ^ a b c d Judd, Burke (September 1998). "Genes and chromomeres: A puzzle in three dimensions". Genetics. Retrieved 2015-10-16.
- ^ Sheval, E. (July 2002). "ORGANIZATION OF HIGHER-LEVEL CHROMATIN STRUCTURES (CHROMOMERE, CHROMONEMA AND CHROMATIN BLOCK) EXAMINED USING VISIBLE LIGHT-INDUCED CHROMATIN PHOTO-STABILIZATION". Cell Biology International. Retrieved 2015-10-16.
- ^ PMID 19556778. Retrieved 2015-10-16.
- ^ PMID 23149574. Retrieved 2015-10-16.
 | This is a user sandbox of Rkelly66. A user sandbox is a subpage of the user's user page. It serves as a testing spot and page development space for the user and is not an encyclopedia article. |
