Chargaff's rules
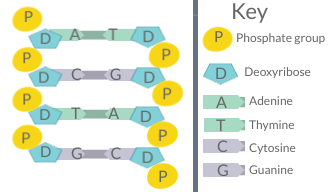
Chargaff's rules (given by Erwin Chargaff) state that in the DNA of any species and any organism, the amount of guanine should be equal to the amount of cytosine and the amount of adenine should be equal to the amount of thymine. Further, a 1:1 stoichiometric ratio of purine and pyrimidine bases (i.e., A+G=T+C) should exist. This pattern is found in both strands of the DNA. They were discovered by Austrian-born chemist Erwin Chargaff[1][2] in the late 1940s.
Definitions
First parity rule
The first rule holds that a double-stranded
Second parity rule
The second rule holds that both Α% ≈ Τ% and G% ≈ C% are valid for each of the two DNA strands.[3] This describes only a global feature of the base composition in a single DNA strand.[4]
Research
The second parity rule was discovered in 1968.[3] It states that, in single-stranded DNA, the number of adenine units is approximately equal to that of thymine (%A ≈ %T), and the number of cytosine units is approximately equal to that of guanine (%C ≈ %G).
The first empirical generalization of Chargaff's second parity rule, called the Symmetry Principle, was proposed by Vinayakumar V. Prabhu[5] in 1993. This principle states that for any given oligonucleotide, its frequency is approximately equal to the frequency of its complementary reverse oligonucleotide. A theoretical generalization[6] was mathematically derived by Michel E. B. Yamagishi and Roberto H. Herai in 2011.[7]
In 2006, it was shown that this rule applies to four
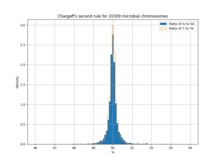

The rule itself has consequences. In most bacterial genomes (which are generally 80-90% coding) genes are arranged in such a fashion that approximately 50% of the coding sequence lies on either strand.
The combined effect of Chargaff's second rule and Szybalski's rule can be seen in bacterial genomes where the coding sequences are not equally distributed. The
Multivariate statistical analysis of codon use within genomes with unequal quantities of coding sequences on the two strands has shown that codon use in the third position depends on the strand on which the gene is located. This seems likely to be the result of Szybalski's and Chargaff's rules. Because of the asymmetry in pyrimidine and purine use in coding sequences, the strand with the greater coding content will tend to have the greater number of purine bases (Szybalski's rule). Because the number of purine bases will, to a very good approximation, equal the number of their complementary pyrimidines within the same strand and, because the coding sequences occupy 80–90% of the strand, there appears to be (1) a selective pressure on the third base to minimize the number of purine bases in the strand with the greater coding content; and (2) that this pressure is proportional to the mismatch in the length of the coding sequences between the two strands.
The origin of the deviation from Chargaff's rule in the organelles has been suggested to be a consequence of the mechanism of replication.[13] During replication the DNA strands separate. In single stranded DNA, cytosine spontaneously slowly deaminates to adenosine (a C to A transversion). The longer the strands are separated the greater the quantity of deamination. For reasons that are not yet clear the strands tend to exist longer in single form in mitochondria than in chromosomal DNA. This process tends to yield one strand that is enriched in guanine (G) and thymine (T) with its complement enriched in cytosine (C) and adenosine (A), and this process may have given rise to the deviations found in the mitochondria. [citation needed][dubious ]
Chargaff's second rule appears to be the consequence of a more complex parity rule: within a single strand of DNA any oligonucleotide (
| First codon | Second codon | Relation proposed | Details |
|---|---|---|---|
Twx (1st base position is T) |
yzA (3rd base position is A) |
% Twx % yzA |
Twx and yzA are mirror codons, e.g. TCG and CGA
|
Cwx (1st base position is C) |
yzG (3rd base position is G) |
% Cwx % yzG |
Cwx and yzG are mirror codons, e.g. CTA and TAG
|
wTx (2nd base position is T) |
yAz (2nd base position is A) |
% wTx % yAz |
wTx and yAz are mirror codons, e.g. CTG and CAG
|
wCx (2nd base position is C) |
yGz (2nd base position is G) |
% wCx % yGz |
wCx and yGz are mirror codons, e.g. TCT and AGA
|
wxT (3rd base position is T) |
Ayz (1st base position is A) |
% wxT % Ayz |
wxT and Ayz are mirror codons, e.g. CTT and AAG
|
wxC (3rd base position is C) |
Gyz (1st base position is G) |
% wxC % Gyz |
wxC and Gyz are mirror codons, e.g. GGC and GCC
|
Examples — computing whole human genome using the first codons reading frame provides:
36530115 TTT and 36381293 AAA (ratio % = 1.00409). 2087242 TCG and 2085226 CGA (ratio % = 1.00096), etc...
In 2020, it is suggested that the physical properties of the dsDNA (double stranded DNA) and the tendency to maximum entropy of all the physical systems are the cause of Chargaff's second parity rule.[16] The symmetries and patterns present in the dsDNA sequences can emerge from the physical peculiarities of the dsDNA molecule and the maximum entropy principle alone, rather than from biological or environmental evolutionary pressure.
Percentages of bases in DNA
The following table is a representative sample of Erwin Chargaff's 1952 data, listing the base composition of DNA from various organisms and support both of Chargaff's rules.[17] An organism such as φX174 with significant variation from A/T and G/C equal to one, is indicative of single stranded DNA.
| Organism | Taxon | %A | %G | %C | %T | A / T | G / C | %GC | %AT |
|---|---|---|---|---|---|---|---|---|---|
| Maize | Zea | 26.8 | 22.8 | 23.2 | 27.2 | 0.99 | 0.98 | 46.1 | 54.0 |
| Octopus | Octopus | 33.2 | 17.6 | 17.6 | 31.6 | 1.05 | 1.00 | 35.2 | 64.8 |
| Chicken | Gallus |
28.0 | 22.0 | 21.6 | 28.4 | 0.99 | 1.02 | 43.7 | 56.4 |
| Rat | Rattus | 28.6 | 21.4 | 20.5 | 28.4 | 1.01 | 1.00 | 42.9 | 57.0 |
| Human | Homo | 29.3 | 20.7 | 20.0 | 30.0 | 0.98 | 1.04 | 40.7 | 59.3 |
| Grasshopper | Orthoptera | 29.3 | 20.5 | 20.7 | 29.3 | 1.00 | 0.99 | 41.2 | 58.6 |
| Sea urchin | Echinoidea |
32.8 | 17.7 | 17.3 | 32.1 | 1.02 | 1.02 | 35.0 | 64.9 |
| Wheat | Triticum |
27.3 | 22.7 | 22.8 | 27.1 | 1.01 | 1.00 | 45.5 | 54.4 |
| Yeast | Saccharomyces | 31.3 | 18.7 | 17.1 | 32.9 | 0.95 | 1.09 | 35.8 | 64.4 |
| E. coli | Escherichia | 24.7 | 26.0 | 25.7 | 23.6 | 1.05 | 1.01 | 51.7 | 48.3 |
φX174 |
PhiX174 |
24.0 | 23.3 | 21.5 | 31.2 | 0.77 | 1.08 | 44.8 | 55.2 |
See also
- Genetic codes
References
- S2CID 36803326.
- ^ S2CID 11358561.
- ^ PMID 4970114.
- PMID 12651717.
- PMID 8332488.
- S2CID 16742066.
- S2CID 16742066.
- PMID 16364245.
- PMID 4966069.
- ^ Cristillo AD (1998). Characterization of G0/G1 switch genes in cultured T lymphocytes. Kingston, Ontario, Canada: Queen's University.
- PMID 10036208.
- PMID 10673280.
- PMID 16893615.
- ^ PMID 17093051.
- S2CID 54565279.
- PMID 32266404.)
{{cite journal}}: CS1 maint: multiple names: authors list (link - ^ Bansal M (2003). "DNA structure: Revisiting the Watson-Crick double helix" (PDF). Current Science. 85 (11): 1556–1563. Archived from the original (PDF) on 2014-07-26. Retrieved 2013-07-26.
Further reading
- Szybalski W, Kubinski H, Sheldrick P (1966). "Pyrimidine clusters on the transcribing strands of DNA and their possible role in the initiation of RNA synthesis". Cold Spring Harbor Symposia on Quantitative Biology. 31: 123–127. PMID 4966069.
- Lobry JR (1996). "Asymmetric substitution patterns in the two DNA strands of bacteria". Mol. Biol. Evol. 13 (5): 660–665. PMID 8676740.
- Lafay B, Lloyd AT, McLean MJ, Devine KM, Sharp PM, Wolfe KH (1999). "Proteome composition and codon usage in spirochaetes: species-specific and DNA strand-specific mutational biases". Nucleic Acids Res. 27 (7): 1642–1649. PMID 10075995.
- McLean MJ, Wolfe KH, Devine KM (1998). "Base composition skews, replication orientation, and gene orientation in 12 prokaryote genomes". J Mol Evol. 47 (6): 691–696. S2CID 12917481.
- McInerney JO (1998). "Replicational and transcriptional selection on codon usage in Borrelia burgdorferi". Proc Natl Acad Sci USA. 95 (18): 10698–10703. PMID 9724767.
External links
- CBS Genome Atlas Database Archived 2016-05-16 at the Portuguese Web Archive — contains hundreds of examples of base skews and had problems.[1]
- The Z curve database of genomes — a 3-dimensional visualization and analysis tool of genomes.[2]

