Curacin A
Appearance
 | |
| Identifiers | |
|---|---|
| |
JSmol) | |
| |
| |
| (verify) | |
Curacin A is a hybrid
Biosynthesis
The synthetic
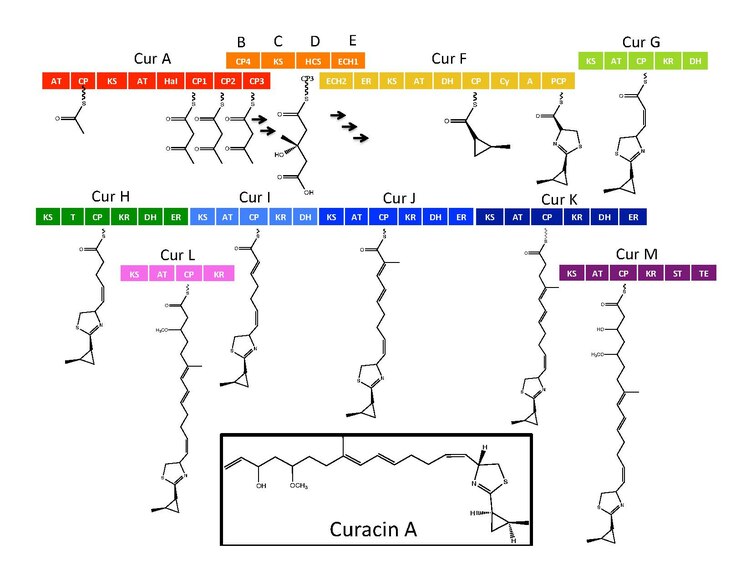
enzymes for Curacin A are found in a gene cluster with 14 open reading frames (ORFs) with the nomenclature CurA through CurN.[1] Analysis of the pathway demonstrated the presence of one NRPS/PKS hybrid module located on CurF, one HMG-CoA synthase cassette located on CurD, and seven monomodular PKS modules.[1] CurA contains a unique GCN5-related N-acetyltransferase (GNAT) loading domain and an associated acyl carrier protein (ACP).[2] The loading module tethers an acetyl group to the ACP that then condenses with one of three tandem ACPs present in the adjacent module of CurA.[1][2][5] An hydroxymethylglutaryl-CoA synthase cassette (mevalonate pathway) catlyzes the formation of hydroxymethylglutaryl acid by the addition of an malonyl-CoA unit to the terminal ketide of the aceto-acetyl-ACP moiety of ACP1,ACP2, or ACP3.[5] subsequent enzymes, including a unique heme independent halogenase (HaI) catalyze the formation of a cyclopropyl ring.[1][5][6] A cysteine specific NRPS module located on CurF follows after cyclopropyl ring formation, and due to the activity of a cyclizing condensation domain, forms a thiazole ring attached to the cylcopropyl moiety from previous reactions in the pathway.[1][5][6] Seven standalone PKS modules follow to extend the growing polyketide chain with S-adenosyl methionine (SAM) dependent methylations occurring at positions 10 and 13.[1] A rare offloading strategy involving a sulfotransferase is employed by the final curacin synthase module. The sulfotransferase sulfates the hydroxyl group of carbon 15, which activates the molecule for decarboxylation and terminal alkene formation.[7]
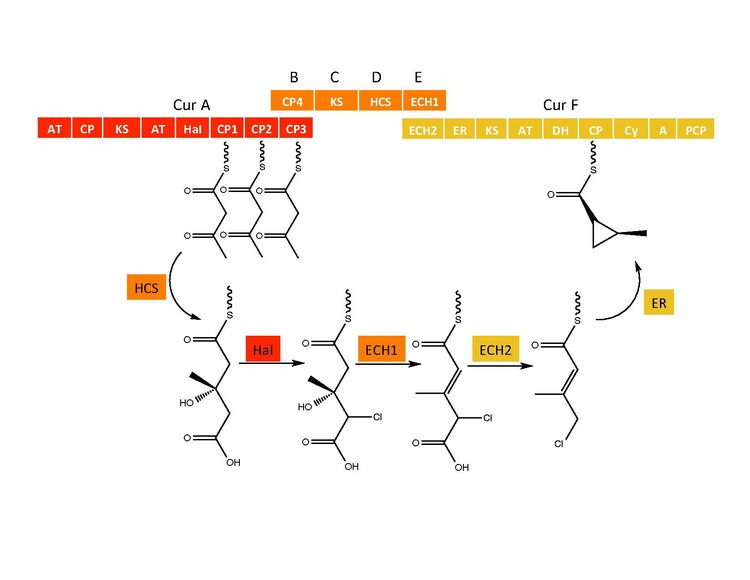
Cyclopropyl ring formation
The CurB (ACP), CurC (ketosynthase), and CurD (HMG-CoA reductase) are responsible for the formation of (S)HMG-ACP3.[6] HaI, from the CurA gene, is a unique non-heme halogenase that goes through a purported Fe(IV)=O intermediate to add a chlorine atom onto an unactivated carbon atom.[6] After chlorination, ECH1 acting as a dehydratates HMG-ACP3 to 3-methylgultaconyl-ACP3 and ECH2 performs the required decarboxylation.[6] Finally,an unusual ER catalyzed cyclization reaction, purported to go through a substitution like mechanism, forms the cyclopropane ring.[6] The added chlorine atom assists in the decarboxylation step and likely serves as the leaving group during cyclopropane ring formation.[6]