Topologically associating domain

A topologically associating domain (TAD) is a self-interacting genomic region, meaning that
The functions of TADs are not fully understood and are still a matter of debate. Most of the studies indicate TADs regulate gene expression by limiting the enhancer-promoter interaction to each TAD;[5] however, a recent study uncouples TAD organization and gene expression.[6] Disruption of TAD boundaries are found to be associated with wide range of diseases such as cancer,[7][8][9] variety of limb malformations such as synpolydactyly, Cooks syndrome, and F-syndrome,[10] and number of brain disorders like Hypoplastic corpus callosum and Adult-onset demyelinating leukodystrophy.[10]
The mechanisms underlying TAD formation are also complex and not yet fully elucidated, though a number of protein complexes and DNA elements are associated with TAD boundaries. However, the handcuff model and the loop extrusion model describe the TAD formation by the aid of CTCF and cohesin proteins.[11] Furthermore, it has been proposed that the stiffness of TAD boundaries itself could cause the domain insulation and TAD formation.[11]
Discovery and diversity
TADs are defined as regions whose DNA sequences preferentially contact each other. They were discovered in 2012 using chromosome conformation capture techniques including Hi-C.[3][12][4] They have been shown to be present in multiple species,[13] including fruit flies (Drosophila),[14] mouse,[3] plants, fungi and human[4] genomes. In bacteria, they are referred to as Chromosomal Interacting Domains (CIDs).[13]
Analytical tools and databases
TAD locations are defined by applying an algorithm to Hi-C data. For example, TADs are often called according to the so-called "directionality index".[4] The directionality index is calculated for individual 40kb bins, by collecting the reads that fall in the bin, and observing whether their paired reads map upstream or downstream of the bin (read pairs are required to span no more than 2Mb). A positive value indicates that more read pairs lie downstream than upstream, and a negative value indicates the reverse. Mathematically, the directionality index is a signed chi-square statistic.
The development of specialized genome browsers and visualization tools[15] such as Juicebox,[16] HiGlass[17]/HiPiler,[18] The 3D Genome Browser,[19] 3DIV,[20] 3D-GNOME,[21] and TADKB[22] have enabled us to visualize the TAD organization of regions of interest in different cell types.
Mechanisms of formation
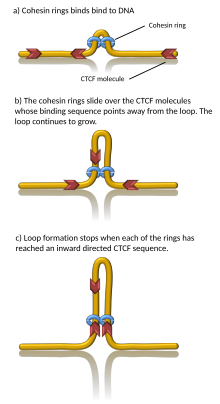
A number of proteins are known to be associated with TAD formation including the protein
Computer simulations have shown that chromatin loop extrusion driven by cohesin motors can generate TADs.[23][24] In the loop extrusion model, cohesin binds chromatin, pulls it in, and extrudes chromatin to progressively grow a loop. Chromatin on both sides of the cohesin complex is extruded until cohesin encounters a chromatin-bound CTCF protein, typically located at the boundary of a TAD. In this way, TAD boundaries can be brought together as the anchors of a chromatin loop.[25] Indeed, in vitro, cohesin has been observed to processively extrude DNA loops in an ATP-dependent manner[26][27][28] and stall at CTCF.[29][30] However, some in vitro data indicates that the observed loops may be artifacts.[31][32] Importantly, since cohesins can dynamically unbind from chromatin, this model suggests that TADs (and associated chromatin loops) are dynamic, transient structures,[23] in agreement with in vivo observations.[33][34][35][36]
Other mechanisms for TAD formation have been suggested. For example, some simulations suggest that transcription-generated supercoiling can relocalize cohesin to TAD boundaries[37][38] or that passively diffusing cohesin “slip links”[39][40] can generate TADs.
Properties
Conservation
TADs have been reported to be relatively constant between different cell types (in stem cells and blood cells, for example), and even between species in specific cases.[4][41][42][43]
Relationship with promoter-enhancer contacts
The majority of observed interactions between promoters and enhancers do not cross TAD boundaries. Removing a TAD boundary (for example, using CRISPR to delete the relevant region of the genome) can allow new promoter-enhancer contacts to form. This can affect gene expression nearby - such misregulation has been shown to cause limb malformations (e.g. polydactyly) in humans and mice.[42]
Computer simulations have shown that transcription-induced supercoiling of chromatin fibres can explain how TADs are formed and how they can assure very efficient interactions between enhancers and their cognate promoters located in the same TAD.[37]
Relationship with other structural features of the genome
Replication timing domains have been shown to be associated with TADs as their boundary is co localized with the boundaries of TADs that are located at either sides of compartments.[44] Insulated neighborhoods, DNA loops formed by CTCF/cohesin-bound regions, are proposed to functionally underlie TADs.[45]
Role in disease
Disruption of TAD boundaries can affect the expression of nearby genes, and this can cause disease.[46]
For example, genomic structural variants that disrupt TAD boundaries have been reported to cause developmental disorders such as human limb malformations.[47][48][49] Additionally, several studies have provided evidence that the disruption or rearrangement of TAD boundaries can provide growth advantages to certain cancers, such as T-cell acute lymphoblastic leukemia (T-ALL),[50] gliomas,[51] and lung cancer.[52]
Lamina-associated domains

Lamina-associated domains (LADs) are parts of the chromatin that heavily interact with the lamina, a network-like structure at the
See also
References
- ^ S2CID 6713103.
- ^ PMID 28783961.
- ^ PMID 22495304.
- ^ PMID 22495300.
- S2CID 11484886.
- PMID 31308546.
- PMID 26855137.
- PMID 27111891.
- PMID 27936932.
- ^ S2CID 22325904.
- ^ PMID 27259200.
- S2CID 4468533.
- ^ PMID 30989119.
- PMID 22265598.
- PMID 31558569.
- PMID 27467250.
- PMID 30143029.
- PMID 28866592.
- PMID 30286773.
- PMID 29106613.
- PMID 27185892.
- ^ Liu, T., Porter, J., Zhao, C. et al. TADKB: Family classification and a knowledge base of topologically associating domains. BMC Genomics 20, 217 (2019). https://doi.org/10.1186/s12864-019-5551-2
- ^ PMID 27210764.
- PMID 26499245.
- S2CID 203653572.
- PMID 32396063.
- S2CID 208228309.
- PMID 31780627.
- bioRxiv 10.1101/2022.09.08.507093.
- bioRxiv 10.1101/2022.09.15.508177.
- PMID 35770038.
- PMID 33568486.
- PMID 35420890.
- bioRxiv 10.1101/2021.04.12.439407.
- bioRxiv 10.1101/2022.03.03.482826.
- PMID 28355183.
- ^ PMID 30395328.
- PMID 29140466.
- S2CID 14706723.
- PMID 29346962.
- PMID 25732821.
- ^ PMID 28040646.
- PMID 31229558.
- S2CID 201714312.
- PMID 26686465.
- PMID 26862051.
- PMID 25959774.
- ^ Angier N (2017-01-09). "A Family's Shared Defect Sheds Light on the Human Genome". The New York Times.
- S2CID 4463482.
- PMID 26940867.
- PMID 26700815.
- PMID 27869826.
- ^ PMID 27312344.
- S2CID 22524502.
