Histone methylation
Histone methylation is a process by which
Function
Histone methylation, as a mechanism for modifying
Animal models have shown methylation and other epigenetic regulation mechanisms to be associated with conditions of aging,
- Histone methylation can be associated with either transcriptional repression or activation. For example, trimethylation of histone H3 at lysine 4 (H3K4me3) is an active mark for transcription and is upregulated in hippocampus one hour after contextual fear conditioning in rats. However, dimethylation of histone H3 at lysine 9 (H3K9me2), a signal for transcriptional silencing, is increased after exposure to either the fear conditioning or a novel environment alone.[5]
- Methylation of some .
- Addition of methyl groups to histones by histone methyltransferases, can either activate or further repress transcription, depending on the amino acid being methylated and the presence of other methyl or acetyl groups in the vicinity.[6]
- Most often, for histone lysine methylations, histone methyltransferases (which methylate) are characterized as "writers", whereas demethylases are characterized as "erasers".[2][4]
Mechanism
The fundamental unit of
Lysine and arginine residues both contain amino groups, which confer basic and hydrophobic characteristics. Lysine is able to be mono-, di-, or trimethylated with a methyl group replacing each hydrogen of its NH3+ group. With a free NH2 and NH2+ group, arginine is able to be mono- or dimethylated. This dimethylation can occur asymmetrically on the NH2 group or symmetrically with one methylation on each group.[9] Each addition of a methyl group on each residue requires a specific set of protein enzymes with various substrates and cofactors. Generally, methylation of an arginine residue requires a complex including protein arginine methyltransferase (PRMT) while lysine requires a specific histone methyltransferase (HMT), usually containing an evolutionarily conserved SET domain.[10]
Different degrees of residue methylation can confer different functions, as exemplified in the methylation of the commonly studied H4K20 residue. Monomethylated H4K20 (H4K20me1) is involved in the compaction of chromatin and therefore transcriptional repression. However, H4K20me2 is vital in the repair of damaged DNA. When dimethylated, the residue provides a platform for the binding of protein 53BP1 involved in the repair of double-stranded DNA breaks by non-homologous end joining. H4K20me3 is observed to be concentrated in heterochromatin and reductions in this trimethylation are observed in cancer progression. Therefore, H4K20me3 serves an additional role in chromatin repression.[10] Repair of DNA double-stranded breaks in chromatin also occurs by homologous recombination and also involves histone methylation (H3K9me3) to facilitate access of the repair enzymes to the sites of damage.[11]
Histone methyltransferase

The genome is tightly condensed into chromatin, which needs to be loosened for
The histone methyltransferases are specific to either lysine or arginine. The lysine-specific transferases are further broken down into whether or not they have a SET domain or a non-SET domain. These domains specify exactly how the enzyme catalyzes the transfer of the methyl from SAM to the transfer protein and further to the histone residue.[12] The methyltransferases can add 1-3 methyls on the target residues.
These methyls that are added to the histones act to regulate transcription by blocking or encouraging DNA access to transcription factors. In this way the integrity of the genome and epigenetic inheritance of genes are under the control of the actions of histone methyltransferases. Histone methylation is key in distinguishing the integrity of the genome and the genes that are expressed by cells, thus giving the cells their identities.
Methylated histones can either repress or activate transcription.
Epigenetics
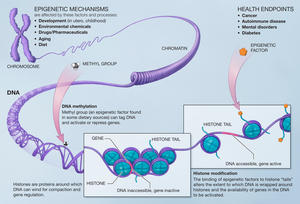
Modifications made on the histone have an effect on the genes that are expressed in a cell and this is the case when methyls are added to the histone residues by the histone methyltransferases.
The activities of histone methyltransferases are offset by the activity of histone demethylases. This allows for the switching on or off of transcription by reversing pre-existing modifications. It is necessary for the activities of both histone methyltransferases and histone demethylases to be regulated tightly. Misregulation of either can lead to gene expression that leads to increased susceptibility to disease. Many cancers arise from the inappropriate epigenetic effects of misregulated methylation.[15] However, because these processes are at times reversible, there is interest in utilizing their activities in concert with anti-cancer therapies.[15]
In X chromosome inactivation
In female organisms, a sperm containing an X chromosome fertilizes the egg, giving the embryo two copies of the X chromosome. Females, however, do not initially require both copies of the X chromosome as it would only double the amount of protein products transcribed as shown by the hypothesis of dosage compensation. The paternal X chromosome is quickly inactivated during the first few divisions.[16] This inactive X chromosome (Xi) is packed into an incredibly tight form of chromatin called heterochromatin.[17] This packing occurs due to the methylation of the different lysine residues that help form different histones. In humans X inactivation is a random process, that is mediated by the non-coding RNA XIST.[18]
Although methylation of lysine residues occurs on many different histones, the most characteristic of Xi occurs on the ninth lysine of the third histone (H3K9). While a single methylation of this region allows for the genes bound to remain transcriptionally active,
Mutations
Due to the fact that histone methylation regulates much of what genes become transcribed, even slight changes to the methylation patterns can have dire effects on the organism. Mutations that occur to increase and decrease methylation have great changes on gene regulation, while mutations to enzymes such as methyltransferase and demethyltransferase can completely alter which proteins are transcribed in a given cell. Over methylation of a chromosome can cause certain genes that are necessary for normal cell function, to become inactivated. In a certain yeast strain, Saccharomyces cerevisiae, a mutation that causes three lysine residues on the third histone, H3K4, H3K36, and H3K79, to become methylated causes a delay in the mitotic cell cycle, as many genes required for this progression are inactivated. This extreme mutation leads to the death of the organism. It has been discovered that the deletion of genes that will eventually allow for the production of histone methyltransferase allows this organism to live as its lysine residues are not methylated.[21]
In recent years it has come to the attention of researchers that many types of cancer are caused largely due to epigenetic factors. Cancer can be caused in a variety of ways due to differential methylation of histones. Since the discovery of
One-carbon metabolism modifies histone methylation
In one-carbon metabolism, the amino acids glycine and serine are converted via the folate and methionine cycles to nucleotide precursors and SAM. Multiple nutrients fuel one-carbon metabolism, including glucose, serine, glycine, and threonine. High levels of the methyl donor SAM influence histone methylation, which may explain how high SAM levels prevent malignant transformation.[24]
See also
- Histone code
- Histone acetylation and deacetylation
- Histone methyltransferase
- Methylation
- Methyllysine
- Genetic imprinting
- DNA methylation
References
- ^ S2CID 6195513.
- ^ PMID 31267065.
- PMID 22986149.
- ^ PMID 28450737.
- PMID 20219993.
- PMID 22473383.
- ^ Jia, Songtao; Wang, Yu (2009). "Degrees make all the difference" (PDF). Epigenetics. 4 (5): 1–4.
- ^ Gilbert, S. F. (2010). Developmental biology. (9th ed.). Sinauer Associates, Inc. 35-37.
- PMID 28061334.
- ^ PMID 11562345.
- PMID 29937925.
- ^ PMID 14690610.
- S2CID 26181430.
- PMID 15677708.
- ^ PMID 19892027.
- S2CID 4190616.
- S2CID 9728548.
- ^ Ahn, J.; Lee, J. (2008). "X chromosome: X inactivation". Nature Education. 1 (1): 24.
- S2CID 6326093.
- PMID 19477626.
- PMID 17664279.
- PMID 16404435.
- PMID 22343901.
- PMID 29784032.
Further reading
- Orouji, Elias & Utikal, Jochen. (2018). Tackling malignant melanoma epigenetically: histone lysine methylation. Clinical Epigenetics 2018 10:145 https://clinicalepigeneticsjournal.biomedcentral.com/articles/10.1186/s13148-018-0583-z
- Gozani, O., & Shi, Y. (2014). Histone Methylation in Chromatin Signaling. In: Fundamentals of Chromatin (pp. 213–256). Springer New York.
- Shi, Yang; Whetstine, Johnathan R. (January 12, 2007). "Dynamic Regulation of Histone Lysine Methylation by Demethylases". Molecular Cell. 25 (1): 1–14. PMID 17218267.
