Medical image computing
Medical image computing (MIC) is an interdisciplinary field at the intersection of
. This field develops computational and mathematical methods for solving problems pertaining to medical images and their use for biomedical research and clinical care.The main goal of MIC is to extract clinically relevant information or knowledge from medical images. While closely related to the field of medical imaging, MIC focuses on the computational analysis of the images, not their acquisition. The methods can be grouped into several broad categories: image segmentation, image registration, image-based physiological modeling, and others.[1]
Data forms
Medical image computing typically operates on uniformly sampled data with regular x-y-z spatial spacing (images in 2D and volumes in 3D, generically referred to as images). At each sample point, data is commonly represented in
Segmentation
Segmentation is the process of partitioning an image into different meaningful segments. In medical imaging, these segments often correspond to different tissue classes,
- Atlas-Based Segmentation: For many applications, a clinical expert can manually label several images; segmenting unseen images is a matter of extrapolating from these manually labeled training images. Methods of this style are typically referred to as atlas-based segmentation methods. Parametric atlas methods typically combine these training images into a single atlas image,[3] while nonparametric atlas methods typically use all of the training images separately.[4] Atlas-based methods usually require the use of image registration in order to align the atlas image or images to a new, unseen image.
- Shape-Based Segmentation: Many methods parametrize a template shape for a given structure, often relying on control points along the boundary. The entire shape is then deformed to match a new image. Two of the most common shape-based techniques are Active Shape Models [5] and Active Appearance Models.[6] These methods have been very influential, and have given rise to similar models.[7]
- Image-Based segmentation: Some methods initiate a template and refine its shape according to the image data while minimizing integral error measures, like the Active contour model and its variations.[8]
- Interactive Segmentation: Interactive methods are useful when clinicians can provide some information, such as a seed region or rough outline of the region to segment. An algorithm can then iteratively refine such a segmentation, with or without guidance from the clinician. Manual segmentation, using tools such as a paint brush to explicitly define the tissue class of each pixel, remains the gold standard for many imaging applications. Recently, principles from feedback control theory have been incorporated into segmentation, which give the user much greater flexibility and allow for the automatic correction of errors.[9]
- Subjective surface Segmentation: This method is based on the idea of evolution of segmentation function which is governed by an advection-diffusion model.[10] To segment an object, a segmentation seed is needed (that is the starting point that determines the approximate position of the object in the image). Consequently, an initial segmentation function is constructed. The idea behind the subjective surface method [11][12][13] is that the position of the seed is the main factor determining the form of this segmentation function.
- Convolutional neural networks (CNN's): The computer-assisted fully automated segmentation performance has been improved due to the advancement of machine learning models. CNN based models such as SegNet,[14] UNet,[15] ResNet,[16] AATSN,[17] Transformers[18] and GANs[19] have fastened the segmentation process. In the future, such models may replace manual segmentation due to their superior performance and speed.
However, there are some other classification of image segmentation methods which are similar to above categories. Moreover, we can classify another group as "Hybrid" which is based on combination of methods.[20]
Registration

Image registration is a process that searches for the correct alignment of images.[21][22][23][24] In the simplest case, two images are aligned. Typically, one image is treated as the target image and the other is treated as a source image; the source image is transformed to match the target image. The optimization procedure updates the transformation of the source image based on a similarity value that evaluates the current quality of the alignment. This iterative procedure is repeated until a (local) optimum is found. An example is the registration of CT and PET images to combine structural and metabolic information (see figure).
Image registration is used in a variety of medical applications:
- Studying temporal changes. Longitudinal studies acquire images over several months or years to study long-term processes, such as disease progression. Time seriescorrespond to images acquired within the same session (seconds or minutes). They can be used to study cognitive processes, heart deformations and respiration.
- Combining complementary information from different imaging modalities. An example is the fusion of anatomical and functional information. Since the size and shape of structures vary across modalities, it is more challenging to evaluate the alignment quality. This has led to the use of similarity measures such as mutual information.[25]
- Characterizing a population of subjects. In contrast to intra-subject registration, a one-to-one mapping may not exist between subjects, depending on the structural variability of the organ of interest. Inter-subject registration is required for atlas construction in computational anatomy.[26] Here, the objective is to statistically model the anatomy of organs across subjects.
- Computer-assisted surgery. In computer-assisted surgery pre-operative images such as CT or MRI are registered to intra-operative images or tracking systems to facilitate image guidance or navigation.
There are several important considerations when performing image registration:
- The diffeomorphisms, which are invertible transformations with a smooth inverse.
- The similarity metric. A distance or similarity function is used to quantify the registration quality. This similarity can be calculated either on the original images or on features extracted from the images. Common similarity measures are sum of squared distances (SSD), correlation coefficient, and mutual information. The choice of similarity measure depends on whether the images are from the same modality; the acquisition noise can also play a role in this decision. For example, SSD is the optimal similarity measure for images of the same modality with Gaussian noise.[27] However, the image statistics in ultrasound are significantly different from Gaussian noise, leading to the introduction of ultrasound specific similarity measures.[28] Multi-modal registration requires a more sophisticated similarity measure; alternatively, a different image representation can be used, such as structural representations[29] or registering adjacent anatomy.[30][31] A recent study[32] employed contrastive coding to learn shared, dense image representations, referred to as CoMIRs (Contrastive Multi-modal Image Representations) which enabled the registration of multi-modal images where existing registration methods often fail due to a lack of sufficiently similar image structures. It reduced the multi-modal registration problem to a mono-modal one, in which general intensity based, as well as feature-based, registration algorithms can be applied.
- The optimization procedure. Either continuous or discrete optimization is performed. For continuous optimization, gradient-based optimization techniques are applied to improve the convergence speed.
Visualization
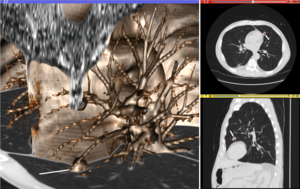
Visualization plays several key roles in Medical Image Computing. Methods from
The figure "Visualization of Medical Imaging" illustrates several types of visualization: 1. the display of cross-sections as gray scale images; 2. reformatted views of gray scale images (the sagittal view in this example has a different orientation than the original direction of the image acquisition; and 3. A 3D volume rendering of the same data. The nodular lesion is clearly visible in the different presentations and has been annotated with a white line.
Atlases
Medical images can vary significantly across individuals due to people having organs of different shapes and sizes. Therefore, representing medical images to account for this variability is crucial. A popular approach to represent medical images is through the use of one or more atlases. Here, an atlas refers to a specific model for a population of images with parameters that are learned from a training dataset.[33][34]
The simplest example of an atlas is a mean intensity image, commonly referred to as a template. However, an atlas can also include richer information, such as local image statistics and the probability that a particular spatial location has a certain label. New medical images, which are not used during training, can be mapped to an atlas, which has been tailored to the specific application, such as segmentation and group analysis. Mapping an image to an atlas usually involves registering the image and the atlas. This deformation can be used to address variability in medical images.
Single template
The simplest approach is to model medical images as deformed versions of a single template image. For example, anatomical MRI brain scans are often mapped to the MNI template [35] as to represent all the brain scans in common coordinates. The main drawback of a single-template approach is that if there are significant differences between the template and a given test image, then there may not be a good way to map one onto the other. For example, an anatomical MRI brain scan of a patient with severe brain abnormalities (i.e., a tumor or surgical procedure), may not easily map to the MNI template.
Multiple templates
Rather than relying on a single template, multiple templates can be used. The idea is to represent an image as a deformed version of one of the templates. For example, there could be one template for a healthy population and one template for a diseased population. However, in many applications, it is not clear how many templates are needed. A simple albeit computationally expensive way to deal with this is to have every image in a training dataset be a template image and thus every new image encountered is compared against every image in the training dataset. A more recent approach automatically finds the number of templates needed.[36]
Statistical analysis
Statistical methods combine the medical imaging field with modern
Group analysis
In the Group Analysis, the objective is to detect and quantize abnormalities induced by a disease by comparing the images of two or more cohorts. Usually one of these cohorts consist of normal (control) subjects, and the other one consists of abnormal patients. Variation caused by the disease can manifest itself as abnormal deformation of anatomy (see
The comparison between groups is usually conducted on the
Classification
Although group analysis can quantify the general effects of a pathology on an anatomy and function, it does not provide subject level measures, and hence cannot be used as biomarkers for diagnosis (see Imaging Biomarkers). Clinicians, on the other hand, are often interested in early diagnosis of the pathology (i.e. classification, The main difficulties are as follows:
- Small sample size (Curse of Dimensionality): a large medical imaging dataset contains hundreds to thousands of images, whereas the number of voxels in a typical volumetric image can easily go beyond millions. A remedy to this problem is to reduce the number of features in an informative sense (see dimensionality reduction). Several unsupervised and semi-/supervised,[44][45][46][47]approaches have been proposed to address this issue.
- Interpretability: A good generalization accuracy is not always the primary objective, as clinicians would like to understand which parts of anatomy are affected by the disease. Therefore, interpretability of the results is very important; methods that ignore the image structure are not favored. Alternative methods based on feature selection have been proposed,.[45][46][47][48]
Clustering
Image-based pattern classification methods typically assume that the neurological effects of a disease are distinct and well defined. This may not always be the case. For a number of medical conditions, the patient populations are highly heterogeneous, and further categorization into sub-conditions has not been established. Additionally, some diseases (e.g.,
Shape analysis
Shape Analysis is the field of Medical Image Computing that studies geometrical properties of structures obtained from different imaging modalities. Shape analysis recently become of increasing interest to the medical community due to its potential to precisely locate morphological changes between different populations of structures, i.e. healthy vs pathological, female vs male, young vs elderly. Shape Analysis includes two main steps: shape correspondence and statistical analysis.
- Shape correspondence is the methodology that computes correspondent locations between geometric shapes represented by triangle meshes, contours, point sets or volumetric images. Obviously definition of correspondence will influence directly the analysis. Among the different options for correspondence frameworks we can find: Anatomical correspondence, manual landmarks, functional correspondence (i.e. in brain morphometry locus responsible for same neuronal functionality), geometry correspondence, (for image volumes) intensity similarity, etc. Some approaches, e.g. spectral shape analysis, do not require correspondence but compare shape descriptors directly.
- Statistical analysis will provide measurements of structural change at correspondent locations.
Longitudinal studies
In longitudinal studies the same person is imaged repeatedly. This information can be incorporated both into the image analysis, as well as into the statistical modeling.
- In longitudinal image processing, segmentation and analysis methods of individual time points are informed and regularized with common information usually from a within-subject template. This regularization is designed to reduce measurement noise and thus helps increase sensitivity and statistical power. At the same time over-regularization needs to be avoided, so that effect sizes remain stable. Intense regularization, for example, can lead to excellent test-retest reliability, but limits the ability to detect any true changes and differences across groups. Often a trade-off needs to be aimed for, that optimizes noise reduction at the cost of limited effect size loss. Another common challenge in longitudinal image processing is the, often unintentional, introduction of processing bias. When, for example, follow-up images get registered and resampled to the baseline image, interpolation artifacts get introduced to only the follow-up images and not the baseline. These artifact can cause spurious effects (usually a bias towards overestimating longitudinal change and thus underestimating required sample size). It is therefore essential that all-time points get treated exactly the same to avoid any processing bias.
- Post-processing and statistical analysis of longitudinal data usually requires dedicated statistical tools such as repeated measure ANOVA or the more powerful linear mixed effects models. Additionally, it is advantageous to consider the spatial distribution of the signal. For example, cortical thickness measurements will show a correlation within-subject across time and also within a neighborhood on the cortical surface - a fact that can be used to increase statistical power. Furthermore, time-to-event (aka survival) analysis is frequently employed to analyze longitudinal data and determine significant predictors.
Image-based physiological modelling
Traditionally, medical image computing has seen to address the quantification and fusion of structural or functional information available at the point and time of image acquisition. In this regard, it can be seen as quantitative sensing of the underlying anatomical, physical or physiological processes. However, over the last few years, there has been a growing interest in the predictive assessment of disease or therapy course. Image-based modelling, be it of biomechanical or physiological nature, can therefore extend the possibilities of image computing from a descriptive to a predictive angle.
According to the STEP research roadmap,[50][51] the Virtual Physiological Human (VPH) is a methodological and technological framework that, once established, will enable the investigation of the human body as a single complex system. Underlying the VPH concept, the International Union for Physiological Sciences (IUPS) has been sponsoring the IUPS Physiome Project for more than a decade,.[52][53] This is a worldwide public domain effort to provide a computational framework for understanding human physiology. It aims at developing integrative models at all levels of biological organization, from genes to the whole organisms via gene regulatory networks, protein pathways, integrative cell functions, and tissue and whole organ structure/function relations. Such an approach aims at transforming current practice in medicine and underpins a new era of computational medicine.[54]
In this context, medical imaging and image computing play an increasingly important role as they provide systems and methods to image, quantify and fuse both structural and functional information about the human being in vivo. These two broad research areas include the transformation of generic computational models to represent specific subjects, thus paving the way for personalized computational models.[55] Individualization of generic computational models through imaging can be realized in three complementary directions:
- definition of the subject-specific computational domain (anatomy) and related subdomains (tissue types);
- definition of boundary and initial conditions from (dynamic and/or functional) imaging; and
- characterization of structural and functional tissue properties.
In addition, imaging also plays a pivotal role in the evaluation and validation of such models both in humans and in animal models, and in the translation of models to the clinical setting with both diagnostic and therapeutic applications. In this specific context, molecular, biological, and pre-clinical imaging render additional data and understanding of basic structure and function in molecules, cells, tissues and animal models that may be transferred to human physiology where appropriate.
The applications of image-based VPH/Physiome models in basic and clinical domains are vast. Broadly speaking, they promise to become new virtual imaging techniques. Effectively more, often non-observable, parameters will be imaged in silico based on the integration of observable but sometimes sparse and inconsistent multimodal images and physiological measurements. Computational models will serve to engender interpretation of the measurements in a way compliant with the underlying biophysical, biochemical or biological laws of the physiological or pathophysiological processes under investigation. Ultimately, such investigative tools and systems will help our understanding of disease processes, the natural history of disease evolution, and the influence on the course of a disease of pharmacological and/or interventional therapeutic procedures.
Cross-fertilization between imaging and modelling goes beyond interpretation of measurements in a way consistent with physiology. Image-based patient-specific modelling, combined with models of medical devices and pharmacological therapies, opens the way to predictive imaging whereby one will be able to understand, plan and optimize such interventions in silico.
Mathematical methods in medical imaging
A number of sophisticated mathematical methods have entered medical imaging, and have already been implemented in various software packages. These include approaches based on
Modality specific computing
Some imaging modalities provide very specialized information. The resulting images cannot be treated as regular scalar images and give rise to new sub-areas of Medical Image Computing. Examples include diffusion MRI, functional MRI and others.
Diffusion MRI
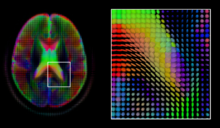
Diffusion MRI is a structural magnetic resonance imaging modality that allows measurement of the diffusion process of molecules. Diffusion is measured by applying a gradient pulse to a magnetic field along a particular direction. In a typical acquisition, a set of uniformly distributed gradient directions is used to create a set of diffusion weighted volumes. In addition, an unweighted volume is acquired under the same magnetic field without application of a gradient pulse. As each acquisition is associated with multiple volumes, diffusion MRI has created a variety of unique challenges in medical image computing.
In medicine, there are two major computational goals in diffusion MRI:
- Estimation of local tissue properties, such as diffusivity;
- Estimation of local directions and global pathways of diffusion.
The
Given the principal direction of diffusion at each location in the volume, it is possible to estimate the global pathways of diffusion through a process known as
Functional MRI
- Task related fMRI is acquired as the subject is performing a sequence of timed experimental conditions. In block-design experiments, the conditions are present for short periods of time (e.g., 10 seconds) and are alternated with periods of rest. Event-related experiments rely on a random sequence of stimuli and use a single time point to denote each condition. The standard approach to analyze task related fMRI is the general linear model (GLM) [76]
- and the linking of network characteristics to behavioral parameters.
There is a rich set of methodology used to analyze functional neuroimaging data, and there is often no consensus regarding the best method. Instead, researchers approach each problem independently and select a suitable model/algorithm. In this context there is a relatively active exchange among neuroscience, computational biology, statistics, and machine learning communities. Prominent approaches include
- Massive univariate approaches that probe individual voxels in the imaging data for a relationship to the experiment condition. The prime approach is the general linear model (GLM) [76]
- Multivariate- and classifier based approaches, often referred to as multi voxel pattern analysis or multi-variate pattern analysis probe the data for global and potentially distributed responses to an experimental condition. Early approaches used support vector machines (SVM) to study responses to visual stimuli.[79] Recently, alternative pattern recognition algorithms have been explored, such as random forest based gini contrast [80] or sparse regression and dictionary learning [81]
- Functional connectivity analysis studies the intrinsic network structure of the brain, including the interactions between regions. The majority of such studies focus on resting state data to parcelate the brain [78] or to find correlates to behavioral measures.[82] Task specific data can be used to study causal relationships among brain regions (e.g., dynamic causal mapping (DCM) [83]).
When working with large cohorts of subjects, the normalization (registration) of individual subjects into a common reference frame is crucial. A body of work and tools exist to perform normalization based on anatomy (FSL, FreeSurfer, SPM). Alignment taking spatial variability across subjects into account is a more recent line of work. Examples are the alignment of the cortex based on fMRI signal correlation,[84] the alignment based on the global functional connectivity structure both in task-, or resting state data,[85] and the alignment based on stimulus specific activation profiles of individual voxels.[86]
Software
Software for medical image computing is a complex combination of systems providing IO, visualization and interaction, user interface, data management and computation. Typically system architectures are layered to serve algorithm developers, application developers, and users. The bottom layers are often libraries and/or toolkits which provide base computational capabilities; while the top layers are specialized applications which address specific medical problems, diseases, or body systems.
Additional notes
Medical Image Computing is also related to the field of
See also
- Brain Connectivity Estimators
- List of Functional Connectivity Software
- Resting State fMRI
- Imaging informatics
- Neuroimaging software
References
- S2CID 237459138.
- .
- ^
J Gee; M Reivich; R Bajcsy (1993). "Elastically Deforming a Three-Dimensional Atlas to Match Anatomical Brain Images". Journal of Computer Assisted Tomography. 17 (1): 225–236. S2CID 25781937.
- ^
MR Sabuncu; BT Yeo; K Van Leemput; B Fischl; PMID 20562040.
- ^
Cootes TF, Taylor CJ, Cooper DH, Graham J (1995). "Active shape models-their training and application". Computer Vision and Image Understanding. 61 (1): 38–59. S2CID 15242659.
- ^ Cootes, T.F.; Edwards, G.J.; Taylor, C.J. (2001). "Active appearance models". IEEE Transactions on Pattern Analysis and Machine Intelligence. 23 (6): 681–685. .
- ISBN 9780128104941.
- PMID 18255491.)
{{cite journal}}: CS1 maint: multiple names: authors list (link - ^
Karasev, P.; Kolesov I.; Chudy, K.; Vela, P.; Tannenbaum, A. (2011). "Interactive MRI segmentation with controlled active vision". IEEE Conference on Decision and Control and European Control Conference. pp. 2293–2298. PMID 24584213.
- ^ K. Mikula, N. Peyriéras, M. Remešíková, A.Sarti: 3D embryogenesis image segmentation by the generalized subjective surface method using the finite volume technique. Proceedings of FVCA5 – 5th International Symposium on Finite Volumes for Complex Applications, Hermes Publ., Paris 2008.
- ^ A. Sarti, G. Citti: Subjective surfaces and Riemannian mean curvature flow graphs. Acta Math. Univ. Comenian. (N.S.) 70 (2000), 85–103.
- ^ A. Sarti, R. Malladi, J.A. Sethian: Subjective Surfaces: A Method for Completing Missing Boundaries. Proc. Natl. Acad. Sci. mi 12, No. 97 (2000), 6258–6263.
- ^ A. Sarti, R. Malladi, J.A. Sethian: Subjective Surfaces: A Geometric Model for Boundary Completion, International Journal of Computer Vision, mi 46, No. 3 (2002), 201–221.
- arXiv:1511.00561 [cs.CV].
- arXiv:1505.04597 [cs.CV].
- S2CID 206594692.
- S2CID 257489603.
- arXiv:1706.03762 [cs.CL].
- S2CID 211072078.
- S2CID 62571134.
- ^
Lisa Gottesfeld Brown (1992). "A survey of image registration techniques". ACM Computing Surveys. 24 (4): 325–376. S2CID 14576088.
- ^
J. Maintz; M. Viergever (1998). "A survey of medical image registration". Medical Image Analysis. 2 (1): 1–36. PMID 10638851.
- ^ J. Hajnal; D. Hawkes; D. Hill (2001). Medical Image Registration. Baton Rouge, Florida: CRC Press.
- ^ Barbara Zitová; Jan Flusser (2003). "Image registration methods: a survey". Image Vision Comput. 21 (11): 977–1000. .
- ^
J. P. W. Pluim; J. B. A. Maintz; M. A. Viergever (2003). "Mutual information based registration of medical images: A survey". IEEE Trans. Med. Imaging. 22 (8): 986–1004. S2CID 2605077.
- ^ Grenander, Ulf; Miller, Michael I. (1998). "Computational anatomy: an emerging discipline". Q. Appl. Math. LVI (4): 617–694. .
- ^ P. A. Viola (1995). Alignment by Maximization of Mutual Information (Thesis). Massachusetts Institute of Technology.
- ^
C. Wachinger; T. Klein; N. Navab (2011). "Locally adaptive Nakagami-based ultrasound similarity measures". Ultrasonics. 52 (4): 547–554. PMID 22197152.
- ^
C. Wachinger; N. Navab (2012). "Entropy and Laplacian images: structural representations for multi-modal registration". Medical Image Analysis. 16 (1): 1–17. PMID 21632274.
- ISSN 0262-8856.
- S2CID 1698371.
- ^ Pielawski, N., Wetzer, E., Ofverstedt, J., Lu, J., Wählby, C., Lindblad, J., & Sladoje, N. (2020). CoMIR: Contrastive Multimodal Image Representation for Registration. In Advances in Neural Information Processing Systems (pp. 18433–18444). Curran Associates, Inc.
- ^
M. De Craene; A. B. d Aische; B. Macq; S. K. Warfield (2004). "Multi-subject Registration for Unbiased Statistical Atlas Construction" (PDF). Medical Image Computing and Computer-Assisted Intervention – MICCAI 2004. Lecture Notes in Computer Science. Vol. 3216. pp. 655–662. ISBN 978-3-540-22976-6.
- ^
C. J. Twining; T. Cootes; S. Marsland; V. Petrovic; R. Schestowitz; C. Taylor (2005). "A Unified Information-Theoretic Approach to Groupwise Non-rigid Registration and Model Building". Information Processing in Medical Imaging. Lecture Notes in Computer Science. Vol. 19. pp. 1–14. PMID 17354680.
- ^ "The MNI brain and the Talairach atlas".
- ^
M. Sabuncu; S. K. Balci; M. E. Shenton; PMID 19336293.
- ^
J. Ashburner; K.J. Friston (2000). "Voxel-Based Morphometry – The Methods". NeuroImage. 11 (6): 805–821. S2CID 16777465.
- ^
C. Davatzikos (2004). "Why voxel-based morphometric analysis should be used with great caution when characterizing group differences". NeuroImage. 23 (1): 17–20. S2CID 7452089.
- ^
K.J. Friston; W.D. Penny; C. Phillips; S.J. Kiebel; G. Hinton; J. Ashburner (2002). "Classical and Bayesian Inference in Neuroimaging: Theory". NeuroImage. 16 (2): 465–483. S2CID 14911371.
- ^
Yong Fan; Nematollah Batmanghelich; Chris M. Clark; Christos Davatzikos (2008). "Spatial patterns of brain atrophy in MCI patients, identified via high-dimensional pattern classification, predict subsequent cognitive decline". NeuroImage. 39 (4): 1731–1743. PMID 18053747.
- ^
Rémi Cuingnet; Emilie Gerardin; Jérôme Tessieras; Guillaume Auzias; Stéphane Lehéricy; Marie-Odile Habert; Marie Chupin; Habib Benali; Olivier Colliot (2011). "The Alzheimer's Disease Neuroimaging Initiative, Automatic classification of patients with Alzheimer's disease from structural MRI: A comparison of ten methods using the ADNI database" (PDF). NeuroImage. 56 (2): 766–781. S2CID 628131.
- ^
Y. Wang; Y. Fan; P. Bhatt P; C. Davatzikos (2010). "High-dimensional pattern regression using machine learning: from medical images to continuous clinical variables". NeuroImage. 50 (4): 1519–35. PMID 20056158.
- ^
Benoît Magnin; Lilia Mesrob; Serge Kinkingnéhun; Mélanie Pélégrini-Issac; Olivier Colliot; Marie Sarazin; Bruno Dubois; Stéphane Lehéricy; Habib Benali (2009). "Support vector machine-based classification of Alzheimer's disease from whole-brain anatomical MRI". Neuroradiology. 51 (2): 73–83. S2CID 285128.
- ^ a b
N.K. Batmanghelich; B. Taskar; C. Davatzikos (2012). "Generative-discriminative basis learning for medical imaging". IEEE Trans Med Imaging. 31 (1): 51–69. PMID 21791408.
- ^ a b
Glenn Fung; Jonathan Stoeckel (2007). "SVM feature selection for classification of SPECT images of Alzheimer's disease using spatial information". Knowledge and Information Systems. 11 (2): 243–258. S2CID 9901011.
- ^ a b
R. Chaves; J. Ramírez; J.M. Górriz; M. López; D. Salas-Gonzalez; I. Álvarez; F. Segovia (2009). "SVM-based computer-aided diagnosis of the Alzheimer's disease using t-test NMSE feature selection with feature correlation weighting". Neuroscience Letters. 461 (3): 293–297. S2CID 9981775.
- ^ a b )
- ^
Savio A.; Graña M. (2013). "Deformation based feature selection for Computer Aided Diagnosis of Alzheimer's Disease". Expert Systems with Applications. 40 (5): 1619–1628. ISSN 0957-4174.
- ^
R. Filipovych; S. M. Resnick; C. Davatzikos (2011). "Semi-supervised cluster analysis of imaging data". NeuroImage. 54 (3): 2185–2197. PMID 20933091.
- ^ STEP research roadmap Archived 2008-08-28 at the Wayback Machine. europhysiome.org
- S2CID 1211981.
- PMID 11144666.
- S2CID 25185270.
- PMID 23115356.
- ^ N. Ayache, J.-P. Boissel, S. Brunak, G. Clapworthy, G. Lonsdale, J. Fingberg, A. F. Frangi, G.Deco, P. J. Hunter, P.Nielsen, M.Halstead, D. R. Hose, I. Magnin, F. Martin-Sanchez, P. Sloot, J. Kaandorp, A. Hoekstra, S. Van Sint Jan, and M. Viceconti (2005) "Towards virtual physiological human: Multilevel modelling and simulation of the human anatomy and physiology". Directorate General INFSO & Directorate General JRC, White paper
- PMID 18218487.
- ^
PMID 23645963.
- ^
P Basser; J Mattiello; D LeBihan (January 1994). "MR diffusion tensor spectroscopy, imaging". Biophysical Journal. 66 (1): 259–267. PMID 8130344.
- ^
P Fillard; X Pennec; V Arsigny; N Ayache (2007). "Clinical DT-MRI estimation, smoothing,, fiber tracking with log-Euclidean metrics". IEEE Transactions on Medical Imaging. 26 (11): 1472–1482. PMID 18041263.
- ^
S-K Song; S-W Sun; M Ramsbottom; C Cheng; J Russell; A Cross (November 2002). "Dysmyelination Revealed through MRI as Increased Radial (but Unchanged Axial) Diffusion of Water". NeuroImage. 13 (3): 1429–1436. S2CID 43229972.
- ^
P Barzo; A Marmarou; P Fatouros; K Hayasaki; F Corwin (December 1997). "Contribution of vasogenic and cellular edema to traumatic brain swelling measured by diffusion-weighted imaging". Journal of Neurosurgery. 87 (6): 900–907. PMID 9384402.
- ^
D Alexander; C Pierpaoli; P Basser (January 2001). "Spatial transformation of diffusion tensor magnetic resonance images" (PDF). IEEE Transactions on Medical Imaging. 20 (11): 1131–1139. S2CID 6559551.
- ^
Y Cao; M Miller; S Mori; R Winslow; L Younes (June 2006). "Diffeomorphic Matching of Diffusion Tensor Images". Proceedings of IEEE Computer Society Conference on Computer Vision, Pattern Recognition (CVPR), Workshop on Mathematical Methods in Biomedical Image Analysis (MMBIA 2006). New York. p. 67. PMC 2920614.
- ^
Z Wang; B Vemuri (October 2005). "DTI segmentation using an information theoretic tensor dissimilarity measure". IEEE Transactions on Medical Imaging. 24 (10): 1267–1277. S2CID 32724414.
- ^
Melonakos, J.; Pichon, E.; PMID 18195436.
- ^
S Mori; B Crain; V Chacko; P van Zijl (February 1999). "Three-dimensional tracking of axonal projections in the brain by magnetic resonance imaging". Annals of Neurology. 45 (2): 265–269. S2CID 334903.
- ^
D Tuch; T Reese; M Wiegell; N Makris; J Belliveau; V Wedeen (October 2002). "High angular resolution diffusion imaging reveals intravoxel white matter fiber heterogeneity". Magnetic Resonance in Medicine. 48 (4): 577–582. PMID 12353272.
- ^
D Tuch (December 2004). "Q-ball imaging". Magnetic Resonance in Medicine. 52 (6): 1358–1372. PMID 15562495.
- ^
V Wedeen; P Hagmann; W-Y Tseng; T Reese (December 2005). "Mapping complex tissue architecture with diffusion spectrum magnetic resonance imaging". Magnetic Resonance in Medicine. 54 (6): 1377–1386. PMID 16247738.
- ^ K Jansons; D Alexander (July 2003). "Persistent angular structure: new insights from diffusion magnetic resonance imaging data". Proceedings of Information Processing in Medical Imaging (IPMI) 2003, LNCS 2732. pp. 672–683. .
- ^
J-D Tournier; F Calamante; D Gadian; A Connelly (2007). "Direct estimation of the fiber orientation density function from diffusion-weighted MRI data using spherical deconvolution". NeuroImage. 23 (3): 1176–1185. S2CID 24169627.
- ^
X Geng; T Ross; W Zhan; H Gu; Y-P Chao; C-P Lin; G Christensen; N Schuff; Y Yang (July 2009). "Diffusion MRI Registration Using Orientation Distribution Functions". Proceedings of Information Processing in Medical Imaging (IPMI) 2009, LNCS 5636. Vol. 21. pp. 626–637. PMC 3860746.
- ^
P-T Yap; Y Chen; H An; Y Yang; J Gilmore; W Lin; D Shen (2011). "SPHERE: SPherical Harmonic Elastic REgistration of HARDI data". NeuroImage. 55 (2): 545–556. PMID 21147231.
- ^ P Zhang; M Niethammer; D Shen; P-T Yap (2012). "Large Deformation Diffeomorphic Registration of Diffusion-Weighted Images" (PDF). Proceedings of Medical Image Computing and Computer-Assisted Intervention (MICCAI). .
- ^ M Descoteaux; R Deriche (September 2007). "Segmentation of Q-Ball Images Using Statistical Surface Evolution". Proceedings of Medical Image Computing and Computer-Assisted Intervention (MICCAI) 2007, LNCS 4792. pp. 769–776. .
- ^ a b
Friston, K.; Holmes, A.; Worsley, K.; Poline, J.; Frith, C.; Frackowiak, R.; et al. (1995). "Statistical parametric maps in functional imaging: a general linear approach". Hum Brain Mapp. 2 (4): 189–210. S2CID 9898609.
- ^
Buckner, R. L.; Andrews-Hanna, J. R.; Schacter, D. L. (2008). "The brain's default network: anatomy, function, and relevance to disease". Annals of the New York Academy of Sciences. 1124 (1): 1–38. S2CID 3167595.
- ^ a b
Yeo, B. T. T.; Krienen, F. M.; Sepulcre, J.; Sabuncu, M. R.; Lashkari, D.; Hollinshead, M.; Roffman, J. L.; Smoller, J. W.; Zöllei, L.; Polimeni, J. R.; Fischl, B.; Liu, H.; Buckner, R. L. (2011). "The organization of the human cerebral cortex estimated by intrinsic functional connectivity". J Neurophysiol. 106 (3): 1125–65. PMID 21653723.
- ^
J. V. Haxby; M. I. Gobbini; M. L. Furey; A. Ishai; J. L. Schouten; P. Pietrini (2001). "Distributed and overlapping representations of faces and objects in ventral temporal cortex". Science. 293 (5539): 2425–30. S2CID 6403660.
- ^
Langs, G.; Menze, B. H.; Lashkari, D.; PMID 20709176.
- ^ Varoquaux, G.; Gramfort, A.; Pedregosa, F.; Michel, V.; Thirion, B. (2011). "Multi-subject dictionary learning to segment an atlas of brain spontaneous activity". Inf Process Med Imaging. Vol. 22. pp. 562–73.
- ^
van den Heuvel, M. P.; Stam, C. J.; Kahn, R. S.; Hulshoff Pol, H. E. (2009). "Efficiency of functional brain networks and intellectual performance". J Neurosci. 29 (23): 7619–24. PMID 19515930.
- ^
Friston, K. (2003). "Dynamic causal modelling". NeuroImage. 19 (4): 1273–1302. S2CID 2176588.
- ^
Sabuncu, M. R.; Singer, B. D.; Conroy, B.; Bryan, R. E.; Ramadge, P. J.; Haxby, J. V. (2010). "Function-based Intersubject Alignment of Human Cortical Anatomy". Cerebral Cortex. 20 (1): 130–140. PMID 19420007.
- ^ Langs, G.; Lashkari, D.; Sweet, A.; Tie, Y.; Rigolo, L.; Golby, A. J.; Golland, P. (2011). "Learning an atlas of a cognitive process in its functional geometry". Inf Process Med Imaging. Vol. 22. pp. 135–46.
- ^
Haxby, J. V.; Guntupalli, J. S.; Connolly, A. C.; Halchenko, Y. O.; Conroy, B. R.; Gobbini, M. I.; Hanke, M.; Ramadge, P. J. (2011). "A common, high-dimensional model of the representational space in human ventral temporal cortex". Neuron. 72 (2): 404–416. PMID 22017997.
- S2CID 31031333.
- .
Journals on medical image computing
- Medical Image Analysis (MedIA) ; also the official journal of The MICCAI Society, which organizes the Annual MICCAI Conference a premier conference for medical image computing
- IEEE Transactions on Medical Imaging (IEEE TMI)
- Medical Physics
- Journal of Digital Imaging (JDI) ; the official journal of the Society of Imaging Informatics
- Computerized Medical Imaging and Graphics
- Journal of Computer Aided Radiology and Surgery
- BMC Medical Imaging
In addition the following journals occasionally publish articles describing methods and specific clinical applications of medical image computing or modality specific medical image computing
