DNA synthesis

DNA synthesis is the natural or artificial creation of
There are several different definitions for DNA synthesis: it can refer to
DNA synthesis occurs in all eukaryotes and prokaryotes, as well as some viruses. The accurate synthesis of DNA is important in order to avoid mutations to DNA. In humans, mutations could lead to diseases such as cancer so DNA synthesis, and the machinery involved in vivo, has been studied extensively throughout the decades. In the future these studies may be used to develop technologies involving DNA synthesis, to be used in data storage.
DNA replication


In nature, DNA molecules are synthesised by all living
Continuously, eukaryotic enzymes encounter DNA damage which can perturb DNA replication. This damage is in the form of DNA lesions that arise spontaneously or due to DNA damaging agents. DNA replication machinery is therefore highly controlled in order to prevent collapse when encountering damage.[2] Control of the DNA replication system ensures that the genome is replicated only once per cycle; over-replication induces DNA damage. Deregulation of DNA replication is a key factor in genomic instability during cancer development.[3]
This highlights the specificity of DNA synthesis machinery in vivo. Various means exist to artificially stimulate the replication of naturally occurring DNA, or to create artificial gene sequences. However, DNA synthesis in vitro can be a very error-prone process.
DNA repair synthesis
Reverse Transcription
Reverse transcription is part of the replication cycle of particular virus families, including retroviruses. It involves copying RNA into double-stranded complementary DNA (cDNA), using reverse transcriptase enzymes. In retroviruses, viral RNA is inserted into a host cell nucleus. There, a viral reverse transcriptase enzyme adds DNA nucleotides onto the RNA sequence, generating cDNA that is inserted into the host cell genome by the enzyme integrase, encoding viral proteins.[5]
Polymerase chain reaction
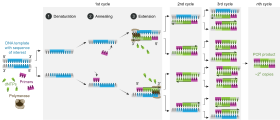
A
DNA synthesis during PCR is very similar to living cells but has very specific reagents and conditions. During PCR, DNA is chemically extracted from host chaperone proteins then heated, causing thermal dissociation of the DNA strands. Two new cDNA strands are built from the original strand, these strands can be split again to act as the template for further PCR products. The original DNA is multiplied through many rounds of PCR.[1] More than a billion copies of the original DNA strand can be made.
Random mutagenesis
For many experiments, such as structural and evolutionary studies, scientists need to produce a large library of variants of a particular DNA sequence. Random mutagenesis takes place in vitro, when mutagenic replication with a low fidelity DNA polymerase is combined with selective PCR amplification to produce many copies of mutant DNA.[6]
RT-PCR
RT-PCR differs from conventional PCR as it synthesizes cDNA from mRNA, rather than template DNA. The technique couples a reverse transcription reaction with PCR-based amplification, as an RNA sequence acts as a template for the enzyme, reverse transcriptase. RT-PCR is often used to test gene expression in particular tissue or cell types at various developmental stages or to test for genetic disorders.[7]
Gene synthesis
Artificial gene synthesis is the process of synthesizing a gene in vitro without the need for initial template DNA samples. In 2010
Oligonucleotide synthesis
Oligonucleotide synthesis is the chemical synthesis of sequences of nucleic acids. The majority of biological research and bioengineering involves synthetic DNA, which can include oligonucleotides, synthetic genes, or even chromosomes. Today, all synthetic DNA is custom-built using the phosphoramidite method by Marvin H. Caruthers. Oligos are synthesized from building blocks which replicate natural bases. The process has been automated since the late 1970s and can be used to form desired genetic sequences as well as for other uses in medicine and molecular biology. However, creating sequences chemically is impractical beyond 200-300 bases, and is an environmentally hazardous process. These oligos, of around 200 bases, can be connected using DNA assembly methods, creating larger DNA molecules.[9]
Some studies have explored the possibility of enzymatic synthesis using terminal deoxynucleotidyl transferase (TdT), a DNA polymerase that requires no template. However, this method is not yet as effective as chemical synthesis, and is not commercially available.[10]
With advances in artificial DNA synthesis, the possibility of
Base pair synthesis
It has been reported that new nucleobase pairs can be synthesized, as well as A-T (adenine - thymine) and G-C (guanine - cytosine). Synthetic nucleotides can be used to expand the genetic alphabet and allow specific modification of DNA sites. Even just a third base pair would expand the number of amino acids that can be encoded by DNA from the existing 20 amino acids to a possible 172.[8] Hachimoji DNA is built from eight nucleotide letters, forming four possible base pairs. It therefore doubles the information density of natural DNA. In studies, RNA has even been produced from hachimoji DNA. This technology could also be used to allow data storage in DNA.[12]
References
- ^ S2CID 215257488.
- PMID 30514768.
- PMID 30700044.
- ^ a b Tiwari V, Wilson DM 3rd. DNA Damage and Associated DNA Repair Defects in Disease and Premature Aging. Am J Hum Genet. 2019 Aug 1;105(2):237-257. doi: 10.1016/j.ajhg.2019.06.005. Review. PMID 31374202
- PMID 26104704.
- PMID 29496818.
- PMID 24034314.
- ^ a b Fikes, Bradley J. (May 8, 2014). "Life engineered with expanded genetic code". San Diego Union Tribune. Archived from the original on 9 May 2014. Retrieved 8 May 2014.
- S2CID 49271982.
- PMID 30804572.
- PMID 32269230.
- PMID 30792304.
