Homologous recombination
Homologous recombination is a type of genetic recombination in which genetic information is exchanged between two similar or identical molecules of double-stranded or single-stranded nucleic acids (usually DNA as in cellular organisms but may be also RNA in viruses).
Homologous recombination is widely used by cells to accurately repair harmful DNA breaks that occur on both strands of DNA, known as double-strand breaks (DSB), in a process called homologous recombinational repair (HRR).[1]
Homologous recombination also produces new combinations of DNA sequences during
Homologous recombination is also used in horizontal gene transfer to exchange genetic material between different strains and species of bacteria and viruses. Horizontal gene transfer is the primary mechanism for the spread of antibiotic resistance in bacteria.
Although homologous recombination varies widely among different organisms and cell types, for double-stranded DNA (
Homologous recombination is
History and discovery
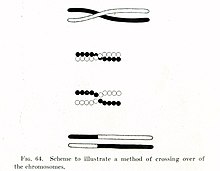
In the early 1900s,
In 1947, the microbiologist
Much of the later work identifying proteins involved in the process and determining their mechanisms has been performed by a number of individuals including James Haber, Patrick Sung, Stephen Kowalczykowski, and others.
In eukaryotes
Homologous recombination (HR) is essential to cell division in eukaryotes like plants, animals, fungi and protists. Homologous recombination repairs double-strand breaks in DNA caused by ionizing radiation or DNA-damaging chemicals.[19] Left unrepaired, these double-strand breaks can cause large-scale rearrangement of chromosomes in somatic cells,[20] which can in turn lead to cancer.[21]
In addition to repairing DNA, homologous recombination also helps produce
Timing within the mitotic cell cycle
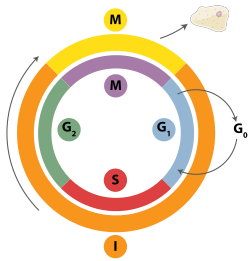
Double-strand breaks can be repaired through homologous recombination, polymerase theta-mediated end joining (TMEJ) or through non-homologous end joining (NHEJ).[28] NHEJ is a DNA repair mechanism which, unlike homologous recombination, does not require a long homologous sequence to guide repair. Whether homologous recombination or NHEJ is used to repair double-strand breaks is largely determined by the phase of cell cycle. Homologous recombination repairs DNA before the cell enters mitosis (M phase). It occurs during and shortly after DNA replication, in the S and G2 phases of the cell cycle, when sister chromatids are more easily available.[29] Compared to homologous chromosomes, which are similar to another chromosome but often have different alleles, sister chromatids are an ideal template for homologous recombination because they are an identical copy of a given chromosome. When no homologous template is available or when the template cannot be accessed due to a defect in homologous recombination, the break is repaired via TMEJ in the S and G2 phases of the cell cycle. In contrast to homologous recombination and TMEJ, NHEJ is predominant in the G1 phase of the cell cycle, when the cell is growing but not yet ready to divide. It occurs less frequently after the G1 phase, but maintains at least some activity throughout the cell cycle. The mechanisms that regulate homologous recombination and NHEJ throughout the cell cycle vary widely between species.[30]
The role of chromatin
The packaging of eukaryotic DNA into chromatin presents a barrier to all DNA-based processes that require recruitment of enzymes to their sites of action. To allow homologous recombination (HR) DNA repair, the chromatin must be remodeled. In eukaryotes, ATP dependent chromatin remodeling complexes and histone-modifying enzymes are two predominant factors employed to accomplish this remodeling process.[34]
Chromatin relaxation occurs rapidly at the site of a DNA damage.
γH2AX, the phosphorylated form of H2AX is also involved in the early steps leading to chromatin decondensation after DNA double-strand breaks. The histone variant H2AX constitutes about 10% of the H2A histones in human chromatin.[38] γH2AX (H2AX phosphorylated on serine 139) can be detected as soon as 20 seconds after irradiation of cells (with DNA double-strand break formation), and half maximum accumulation of γH2AX occurs in one minute.[38] The extent of chromatin with phosphorylated γH2AX is about two million base pairs at the site of a DNA double-strand break.[38] γH2AX does not, itself, cause chromatin decondensation, but within 30 seconds of irradiation, RNF8 protein can be detected in association with γH2AX.[39] RNF8 mediates extensive chromatin decondensation, through its subsequent interaction with CHD4,[40] a component of the nucleosome remodeling and deacetylase complex NuRD.
After undergoing relaxation subsequent to DNA damage, followed by DNA repair, chromatin recovers to a compaction state close to its pre-damage level after about 20 min.[35]
Homologous recombination during meiosis
In vertebrates the locations at which recombination occurs are determined by the binding locations of PRDM9, a protein which recognizes a specific sequence motif by its zinc finger array.[41] At these sites, another protein, SPO11 catalyses recombination-initiating double strand breaks (DSBs), a subset of which are repaired by recombination with the homologous chromosome. PRDM9 deposits both H3K4me3 and H3K36me3 histone methylation marks at the sites it binds, and this methyltransferase activity is essential for its role in DSB positioning. Following their formation, DSB sites are processed by resection, resulting in single-stranded DNA (ssDNA) that becomes decorated with DMC1. From mid-zygotene to early pachytene, as part of the recombinational repair process, DMC1 dissociates from the ssDNA and counts decrease until all breaks (except those on the XY chromosomes) are repaired at late pachytene. Several other proteins are involved in this process, including ZCWPW1, the [42] first protein directly positioned by PRDM9's dual histone marks. ZCWPW1 is important for homologous DSB repair, not positioning.
Models
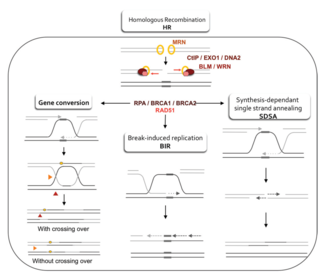
Two primary models for how homologous recombination repairs double-strand breaks in DNA are the double-strand break repair (DSBR) pathway (sometimes called the double Holliday junction model) and the synthesis-dependent strand
The
DSBR pathway

After the stages of resection, strand invasion and DNA synthesis, the DSBR and SDSA pathways become distinct.
Whether recombination in the DSBR pathway results in chromosomal crossover is determined by how the double Holliday junction is cut, or "resolved". Chromosomal crossover will occur if one Holliday junction is cut on the crossing strand and the other Holliday junction is cut on the non-crossing strand (in Figure 5, along the horizontal purple arrowheads at one Holliday junction and along the vertical orange arrowheads at the other). Alternatively, if the two Holliday junctions are cut on the crossing strands (along the horizontal purple arrowheads at both Holliday junctions in Figure 5), then chromosomes without crossover will be produced.[47]
SDSA pathway
Homologous recombination via the SDSA pathway occurs in cells that divide through mitosis and meiosis and results in non-crossover products. In this model, the invading 3' strand is extended along the recipient DNA duplex by a DNA polymerase, and is released as the Holliday junction between the donor and recipient DNA molecules slides in a process called branch migration. The newly synthesized 3' end of the invading strand is then able to
During mitosis, the major homologous recombination pathway for repairing DNA double-strand breaks appears to be the SDSA pathway (rather than the DSBR pathway).[49] The SDSA pathway produces non-crossover recombinants (Figure 5). During meiosis non-crossover recombinants also occur frequently and these appear to arise mainly by the SDSA pathway as well.[49][50] Non-crossover recombination events occurring during meiosis likely reflect instances of repair of DNA double-strand damages or other types of DNA damages.
SSA pathway
The single-strand annealing (SSA) pathway of homologous recombination repairs double-strand breaks between two repeat sequences. The SSA pathway is unique in that it does not require a separate similar or identical molecule of DNA, like the DSBR or SDSA pathways of homologous recombination. Instead, the SSA pathway only requires a single DNA duplex, and uses the repeat sequences as the identical sequences that homologous recombination needs for repair. The pathway is relatively simple in concept: after two strands of the same DNA duplex are cut back around the site of the double-strand break, the two resulting 3' overhangs then align and anneal to each other, restoring the DNA as a continuous duplex.[48][51]
As DNA around the double-strand break is cut back, the single-stranded 3' overhangs being produced are coated with the
BIR pathway
During
The BIR pathway can also help to maintain the length of
Maintaining telomere length is critical for
In bacteria

Homologous recombination is a major DNA repair process in bacteria. It is also important for producing genetic diversity in bacterial populations, although the process differs substantially from
RecBCD pathway
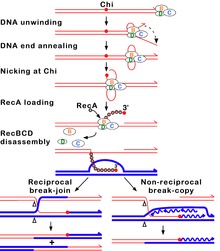
The RecBCD pathway is the main recombination pathway used in many bacteria to repair double-strand breaks in DNA, and the proteins are found in a broad array of bacteria.
In this pathway, a three-subunit
Upon encountering a Chi site, the activity of the RecBCD enzyme changes drastically.[64][61][67] DNA unwinding pauses for a few seconds and then resumes at roughly half the initial speed. This is likely because the slower RecB helicase unwinds the DNA after Chi, rather than the faster RecD helicase, which unwinds the DNA before Chi.[68][69] Recognition of the Chi site also changes the RecBCD enzyme so that it cuts the DNA strand with Chi and begins loading multiple RecA proteins onto the single-stranded DNA with the newly generated 3' end. The resulting RecA-coated nucleoprotein filament then searches out similar sequences of DNA on a homologous chromosome. The search process induces stretching of the DNA duplex, which enhances homology recognition (a mechanism termed conformational proofreading[70][71][72]). Upon finding such a sequence, the single-stranded nucleoprotein filament moves into the homologous recipient DNA duplex in a process called strand invasion.[73] The invading 3' overhang causes one of the strands of the recipient DNA duplex to be displaced, to form a D-loop. If the D-loop is cut, another swapping of strands forms a cross-shaped structure called a Holliday junction.[65] Resolution of the Holliday junction by some combination of RuvABC or RecG can produce two recombinant DNA molecules with reciprocal genetic types, if the two interacting DNA molecules differ genetically. Alternatively, the invading 3’ end near Chi can prime DNA synthesis and form a replication fork. This type of resolution produces only one type of recombinant (non-reciprocal).
RecF pathway
Bacteria appear to use the RecF pathway of homologous recombination to repair single-strand gaps in DNA. When the RecBCD pathway is inactivated by mutations and additional mutations inactivate the SbcCD and ExoI nucleases, the RecF pathway can also repair DNA double-strand breaks.
Although the proteins and specific mechanisms involved in their initial phases differ, the two pathways are similar in that they both require single-stranded DNA with a 3' end and the RecA protein for strand invasion. The pathways are also similar in their phases of branch migration, in which the Holliday junction slides in one direction, and resolution, in which the Holliday junctions are cleaved apart by enzymes.[75][76] The alternative, non-reciprocal type of resolution may also occur by either pathway.
Branch migration
Immediately after strand invasion, the Holliday junction moves along the linked DNA during the branch migration process. It is in this movement of the Holliday junction that base pairs between the two homologous DNA duplexes are exchanged. To catalyze branch migration, the RuvA protein first recognizes and binds to the Holliday junction and recruits the RuvB protein to form the RuvAB complex. Two sets of the RuvB protein, which each form a ring-shaped ATPase, are loaded onto opposite sides of the Holliday junction, where they act as twin pumps that provide the force for branch migration. Between those two rings of RuvB, two sets of the RuvA protein assemble in the center of the Holliday junction such that the DNA at the junction is sandwiched between each set of RuvA. The strands of both DNA duplexes—the "donor" and the "recipient" duplexes—are unwound on the surface of RuvA as they are guided by the protein from one duplex to the other.[77][78]
Resolution
In the resolution phase of recombination, any Holliday junctions formed by the strand invasion process are cut, thereby restoring two separate DNA molecules. This cleavage is done by RuvAB complex interacting with RuvC, which together form the
Facilitating genetic transfer
Homologous recombination is an important method of integrating donor DNA into a recipient organism's genome in
In bacterial conjugation, where DNA is transferred between bacteria through direct cell-to-cell contact, homologous recombination helps integrate foreign DNA into the host genome via the RecBCD pathway. The RecBCD enzyme promotes recombination after DNA is converted from single-strand DNA–in which form it originally enters the bacterium–to double-strand DNA during replication. The RecBCD pathway is also essential for the final phase of transduction, a type of horizontal gene transfer in which DNA is transferred from one bacterium to another by a virus. Foreign, bacterial DNA is sometimes misincorporated in the capsid head of bacteriophage virus particles as DNA is packaged into new bacteriophages during viral replication. When these new bacteriophages infect other bacteria, DNA from the previous host bacterium is injected into the new bacterial host as double-strand DNA. The RecBCD enzyme then incorporates this double-strand DNA into the genome of the new bacterial host.[65]
Bacterial transformation
Natural bacterial transformation involves the transfer of DNA from a donor bacterium to a recipient bacterium, where both donor and recipient are ordinarily of the same species. Transformation, unlike bacterial conjugation and transduction, depends on numerous bacterial gene products that specifically interact to perform this process.[84] Thus transformation is clearly a bacterial adaptation for DNA transfer. In order for a bacterium to bind, take up and integrate donor DNA into its resident chromosome by homologous recombination, it must first enter a special physiological state termed competence. The RecA/Rad51/DMC1 gene family plays a central role in homologous recombination during bacterial transformation as it does during eukaryotic meiosis and mitosis. For instance, the RecA protein is essential for transformation in Bacillus subtilis and Streptococcus pneumoniae,[85] and expression of the RecA gene is induced during the development of competence for transformation in these organisms.
As part of the transformation process, the RecA protein interacts with entering single-stranded DNA (ssDNA) to form RecA/ssDNA nucleofilaments that scan the resident chromosome for regions of homology and bring the entering ssDNA to the corresponding region, where strand exchange and homologous recombination occur.[86] Thus the process of homologous recombination during bacterial transformation has fundamental similarities to homologous recombination during meiosis.
In viruses
Homologous recombination occurs in several
In RNA viruses, homologous recombination can be either precise or imprecise. In the precise type of RNA-RNA recombination, there is no difference between the two parental RNA sequences and the resulting crossover RNA region. Because of this, it is often difficult to determine the location of crossover events between two recombining RNA sequences. In imprecise RNA homologous recombination, the crossover region has some difference with the parental RNA sequences – caused by either addition, deletion, or other modification of nucleotides. The level of precision in crossover is controlled by the sequence context of the two recombining strands of RNA: sequences rich in adenine and uracil decrease crossover precision.[88][90]
Homologous recombination is important in facilitating viral evolution.[88][91] For example, if the genomes of two viruses with different disadvantageous mutations undergo recombination, then they may be able to regenerate a fully functional genome. Alternatively, if two similar viruses have infected the same host cell, homologous recombination can allow those two viruses to swap genes and thereby evolve more potent variations of themselves.[91]
Homologous recombination is the proposed mechanism whereby the DNA virus
When two or more viruses, each containing lethal genomic damage, infect the same host cell, the virus genomes can often pair with each other and undergo homologous recombinational repair to produce viable progeny. This process, known as multiplicity reactivation, has been studied in several
Coronaviruses are capable of genetic recombination when at least two viral genomes are present in the same infected cell. RNA recombination appears to be a major driving force in determining (1) genetic variability within a CoV species, (2) the capability of a CoV species to jump from one host to another, and (3) infrequently, the emergence of novel CoVs.[97] The mechanism of recombination in CoVs likely involves template switching during genome replication.[97] Recombination in RNA viruses appears to be an adaptation for coping with genome damage.[98]
The pandemic SARS-CoV-2's entire receptor binding motif appears to have been introduced through recombination from coronaviruses of pangolins.[99] Such a recombination event may have been a critical step in the evolution of SARS-CoV-2's capability to infect humans.[99] Recombination events are likely key steps in the evolutionary process that leads to the emergence of new human coronaviruses.[100]
During COVID-19 pandemic in 2020, many genomic sequences of Australian SARS‐CoV‐2 isolates have deletions or mutations (29742G>A or 29742G>U; "G19A" or "G19U")in the Coronavirus 3′ stem-loop II-like motif (s2m), an RNA motif in 3' untranslated region of viral genome, suggesting that RNA recombination events may have occurred in s2m of SARS-CoV-2. Based on computational analysis of 1319 Australia SARS‐CoV‐2 sequences using Recco algorithm (https://recco.bioinf.mpi-inf.mpg.de/), 29742G("G19"), 29744G("G21"), and 29751G("G28") were predicted as recombination hotspots.[101]
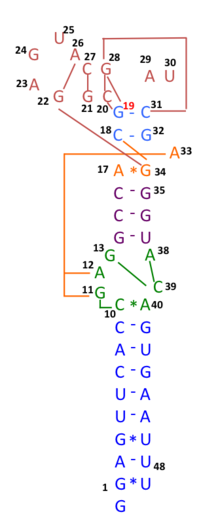
The SARS-CoV-2 outbreak in Diamond Princess cruise most likely originated from either a single person infected with a virus variant identical to the Wuhan WIV04 isolates, or simultaneously with another primary case infected with a virus containing the 11083G > T mutation. Linkage disequilibrium analysis confirmed that RNA recombination with the 11083G > T mutation also contributed to the increase of mutations among the viral progeny. The findings indicate that the 11083G > T mutation of SARS-CoV-2 spread during shipboard quarantine and arose through de novo RNA recombination under positive selection pressure. In addition, in three patients in this cruise, two mutations 29736G > T and 29751G > T ("G13" and "G28") were also located in Coronavirus 3′ stem-loop II-like motif (s2m), as "G28" was predicted as recombination hotspots in Australian SARS-CoV-2 mutants. Although s2m is considered an RNA motif highly conserved among many coronavirus species, this result also suggests that s2m of SARS-CoV-2 is rather a RNA recombination/mutation hotspot.[102]
Effects of dysfunction
Without proper homologous recombination, chromosomes often incorrectly align for the first phase of cell division in
Deficiencies in homologous recombination have been strongly linked to
Decreased rates of homologous recombination cause inefficient DNA repair,
Tumours with a homologous recombination deficiency (including BRCA defects) are described as HRD-positive.[109]
Evolutionary conservation
While the pathways can mechanistically vary, the ability of organisms to perform homologous recombination is universally conserved across all domains of life.[110] Based on the similarity of their amino acid sequences, homologs of a number of proteins can be found in multiple domains of life indicating that they evolved a long time ago, and have since diverged from common ancestral proteins.[110]
RecA recombinase family members are found in almost all organisms with
Related single stranded binding proteins that are important for homologous recombination, and many other processes, are also found in all domains of life.[112]
Rad54, Mre11, Rad50, and a number of other proteins are also found in both archaea and eukaryotes.[110][111][113]
The RecA recombinase family
The proteins of the RecA recombinase family of proteins are thought to be descended from a common ancestral recombinase.
The proteins generally share a long conserved region known as the RecA/Rad51 domain. Within this protein domain are two sequence motifs, Walker A motif and Walker B motif. The Walker A and B motifs allow members of the RecA/Rad51 protein family to engage in ATP binding and ATP hydrolysis.[110][114]
Meiosis-specific proteins
The discovery of Dmc1 in several species of
Technological applications
Gene targeting
Many methods for introducing DNA sequences into organisms to create
Advances in gene targeting technologies which hijack the homologous recombination mechanics of cells are now leading to the development of a new wave of more accurate, isogenic human disease models. These engineered human cell models are thought to more accurately reflect the genetics of human diseases than their mouse model predecessors. This is largely because mutations of interest are introduced into endogenous genes, just as they occur in the real patients, and because they are based on human genomes rather than rat genomes. Furthermore, certain technologies enable the knock-in of a particular mutation rather than just knock-outs associated with older gene targeting technologies.
Protein engineering
Techniques that rely on homologous recombination have been used to engineer new proteins.
Cancer therapy
Homologous recombination proficient (HRP) cancer cells are able to repair the DNA damage, which is caused by chemotherapy such as cisplatin. Thus, HRP cancers are difficult to treat. Studies suggest that homologous recombination can be targeted via c-Abl inhibition.
See also
References
- PMID 11376695.
- OCLC 145080076.
- PMID 2660260.
- S2CID 30212766.
- PMID 6273866.
- PMID 6308623.
- ^ S2CID 39590123.
- PMID 940351.
- PMID 24097900.
- S2CID 26806110.
- ^ "Reginald Crundall Punnett". NAHSTE, University of Edinburgh. Retrieved 3 July 2010.
- ^ Lobo I, Shaw K (2008). "Thomas Hunt Morgan, genetic recombination, and gene mapping". Nature Education. 1 (1).
- ^ PMID 15867161.
- PMID 16587654.
- ^ Stern, C (1931). "Zytologisch-genetische untersuchungen alsbeweise fur die Morgansche theorie des faktoraustauschs". Biologisches Zentralblatt. 51: 547–587.
- ^ "The development of bacterial genetics". US National Library of Medicine. Retrieved 3 July 2010.
- ^ "The Nobel Prize in Physiology or Medicine 1958". Nobelprize.org. Retrieved 3 July 2010.
- ^ PMID 15065659.
- ISBN 978-0-7167-3136-8.
- ISBN 978-0-7167-3118-4.
- S2CID 3012823.
- ^ ISBN 978-0-7167-4339-2.
- S2CID 27658497.
- ISBN 978-0-8153-4105-5.
- ^ S2CID 8294596.
- PMID 19464965.
- PMID 423159.
- S2CID 195892718.
- ISBN 978-0-8153-4105-5.
- ^ PMID 18157161.
- ^ PMID 19375328.
- PMID 28623092.
- PMID 18716619.
- PMID 23633913.
- ^ PMID 27733626.
- ^ PMID 27568560.
- ^ PMID 18025084.
- ^ PMID 9488723.
- S2CID 14232192.
- PMID 22531782.
- PMID 20044539.
- PMID 32744506.
- ^ S2CID 30324005.
- PMID 9242902.
- PMID 17988174.

- PMID 10648605.
- ISBN 978-0-8153-4105-5.
- ^ PMID 17363343.
- ^ PMID 20967781.
- S2CID 1878863.
- ^ Haber lab. "Single-strand annealing". Brandeis University. Archived from the original on 19 January 2015. Retrieved 3 July 2010.
- ^ PMID 19260026.
- PMID 19473888.
- PMID 10357855.
- ^ PMID 16756487.
- PMID 19180191.

- PMID 16244317.
- S2CID 4416531.
- PMID 7968921.
- PMID 16132081.

- ^ PMID 18079176.
- S2CID 2916995. Archived from the original(PDF) on 2004-05-25.
- PMID 19542287.
- ^ PMID 22688812.
- ^ PMID 19052323.
- PMID 17395553.
- S2CID 16662983.
- S2CID 4302346.
- PMID 18022364.
- S2CID 1682936. Archived from the original(PDF) on 2012-10-07. Retrieved 2011-08-31.
- PMID 21070960. Archived from the original(PDF) on 2012-10-07. Retrieved 2011-08-31.
- PMID 22560720.
- ISBN 978-0-8153-4105-5.
- PMID 12769856.
- S2CID 2221866.
- PMID 19451222.
- ^ S2CID 28474965.
- ^ ISBN 978-0-8053-4635-0.
- PMID 18043749.

- S2CID 1231127. Archived from the original(PDF) on 2010-06-01.
- PMID 9275198.
- PMID 9475717.
- PMID 10648528.
- S2CID 205499369.
- PMID 19228200.
- S2CID 17272331.
- ISBN 978-0-9631172-1-2.
- ^ PMID 8523555.
- PMID 20454662.

- PMID 10571050.
- ^ PMID 15012521.
- PMID 21458587.
- PMID 6261109.
- ISBN 978-0896038035
- PMID 8456313.
- ^ PMID 27012512.
- PMID 20335491.
- ^ PMID 32937441.
- PMID 32210130.
- PMID 32506536.
- PMID 34248221.
- PMID 15551222.
- ^ Cold Spring Harbor Laboratory (2007). "Human RecQ Helicases, Homologous Recombination And Genomic Instability". ScienceDaily. Retrieved 3 July 2010.
- PMID 11387040.
- S2CID 21218975.
- ^ ISBN 978-0-8153-4110-9.
- ^ PMID 12947386.
- ^ "Use of homologous recombination deficiency (HRD) score to enrich for niraparib sensitive high grade ovarian tumors". Archived from the original on 2017-04-30. Retrieved 2016-12-30.
- ^ PMID 16798872.
- ^ PMID 19282450.
- PMID 20080104.
- PMID 25880130.

- PMID 8051165.
- S2CID 17013247.
- ^ PMID 17921483.
- ISBN 978-0-7167-3136-8.
- ^ "The Nobel Prize in Physiology or Medicine 2007". The Nobel Foundation. Retrieved December 15, 2008.
- PMID 15809422.
- PMID 15644440.
- ^ PMID 17884462.
- PMID 16594730.

- S2CID 4363255.
- S2CID 43516273.
- PMID 17379142.
- S2CID 6493434.
- PMID 34616682.
- ^ PMID 19553640.
- PMID 19553641.
- S2CID 205212044.
External links
- Animations – homologous recombination: Animations showing several models of homologous recombination
- Homologous recombination: Tempy & Trun: Animation of the bacterial RecBCD pathway of homologous recombination
