Mitochondrial biogenesis
Mitochondrial biogenesis is the process by which cells increase
Background
The ability for a mitochondrion to self-replicate is rooted in its evolutionary history. It is commonly thought that mitochondria descend from cells that formed
Mitochondria are produced from the transcription and translation of genes both in the nuclear genome and in the
Protein import
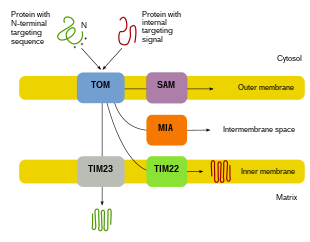
Since the majority of mitochondrial protein comes from the nuclear genome, the proteins need to be properly targeted and transported into the mitochondria to perform their functions.[8][10][11] First, mRNA is translated in the cell's cytosol.[10][11] The resulting unfolded precursor proteins will then be able to reach their respective mitochondrial compartments.[11][10] Precursor proteins will be transported to one of four areas of the mitochondria, which include the outer membrane, inner membrane, intermembrane space, and matrix.[10][11] All proteins will enter the mitochondria by a translocase on the outer mitochondrial membrane (TOM).[11][10][5] Some proteins will have an N-terminal targeting signal, and these proteins will be detected and transported into the matrix, where they will then be cleaved and folded.[12][11][10] Other proteins may have targeting information in their sequences and will not include an N-terminal signal.[11][10] During the past two decades, researchers have discovered over thirty proteins that participate in mitochondrial protein import.[11] As researchers learn more about these proteins and how they reach the respective mitochondrial compartments that utilize them, it becomes evident that there is a multitude of processes that work together in the cell to allow for mitochondrial biogenesis.[11][8]
Fusion and fission
Mitochondria are highly versatile and are able to change their shape through fission and fusion events.[9][8] Definitively, fission is the event of a single entity breaking apart, whereas fusion is the event of two or more entities joining to form a whole.[8] The processes of fission and fusion oppose each other and allow the mitochondrial network to constantly remodel itself.[9][8] If a stimulus induces a change in the balance of fission and fusion in a cell, it could significantly alter the mitochondrial network.[9][13] For example, an increase in mitochondrial fission would create many fragmented mitochondria, which has been shown to be useful for eliminating damaged mitochondria and for creating smaller mitochondria for efficient transporting to energy-demanding areas.[13][14] Therefore, achieving a balance between these mechanisms allows a cell to have the proper organization of its mitochondrial network during biogenesis and may have an important role in muscle adaptation to physiological stress.[13]

In mammals, mitochondrial fusion and fission are both controlled by GTPases of the dynamin family.[8][13] The process of mitochondrial fission is directed by Drp1, a member of the cytosolic dynamin family.[8][9] This protein forms a spiral around mitochondria and constricts to break apart both the outer and inner membranes of the organelle.[14] On the other hand, the process of fusion is directed by different membrane-anchored dynamin proteins at different levels of the mitochondria.[13] Fusion at the level of the outer mitochondrial membrane is mediated by Mfn1 and Mfn2 (Mitofusins 1 and 2),[15] and fusion at the level of the inner mitochondrial membrane is mediated by Opa1.[8][12][13] Multiple research studies have observed correlated increases between mitochondrial respiratory capacity with Mfn1, Mnf2, and Drp1 gene expression after endurance exercises.[14][15] Therefore, it is supported that reorganization of the mitochondrial network in muscle cells plays an important role in response to exercise.[4][13][15]
Regulation
Post-translational modifications
SIRT7, a histone H3K18-specific deacetylase, epigenetically controls mitochondria biogenesis, ribosomal biosynthesis, and DNA repair. SIRT7 is methylated at arginine 388 (R388), which inhibits its H3K18 deacetylase activity. Protein arginine methyltransferase 6 (PRMT6) directly interacts with and methylates SIRT7 at R388 in vitro and in vivo R388 methylation suppresses the H3K18 deacetylase activity of SIRT7 without modulating its subcellular localization. PRMT6-induced H3K18 hyperacetylation at SIRT7-target gene promoter epigenetically promotes mitochondria biogenesis and maintains mitochondria respiration. Moreover, high glucose enhances R388 methylation in mouse fibroblasts and liver tissue. PRMT6 signals glucose availability to SIRT7 in an AMPK-dependent manner. AMPK induces R388 hypomethylation by disrupting the association between PRMT6 and SIRT7. Together, PRMT6-induced arginine methylation of SIRT7 coordinates glucose availability with mitochondria biogenesis to maintain energy homeostasis. [16]
Gene transcription
Cell signaling
AMP-activated kinase (AMPK) also regulates mitochondrial biogenesis by phosphorylating and activating PGC-1α upon sensing an energy deficiency in muscle.[5][17] In mice with reduced ATP/AMP ratios that would occur during exercise, the energy depletion has been shown to correlate with AMPK activation.[5][19][17] AMPK activation then continued to activate PGC- 1α and NRFs in these mice, and mitochondrial biogenesis was stimulated.[5][19][17]
Fumarate signaling
Human genome encodes three different malic enzymes. ME1 is NADP-depdendent. ME2 is NAD-depdendent, while ME3 displays dual coenzyme specificity. ME2 is allosterically activated by fumarate, which is distinctive from the other two malic enzymes (ME1 and ME3). The
Aging
The capacity for mitochondrial biogenesis has been shown to decrease with age, and such decreased mitochondrial function has been associated with diabetes and cardiovascular disease.[22][23][24] Aging and disease can induce changes in the expression levels of proteins involved in the fission and fusion mechanisms of mitochondria, thus creating dysfunctional mitochondria.[25][26] One hypothesis for the detrimental results of aging is associated with the loss of telomeres, the end segments of chromosomes that protect genetic information from degradation.[23][26] Telomere loss has also been associated with decreased mitochondrial function.[26][23] Deficiency of telomerase reverse transcriptase (TERT), an enzyme that plays a role in preserving telomeres, has been correlated with activated p53, a protein that suppresses PGC-1α.[26][25][23] Therefore, the loss of telomeres and TERT that comes with aging has been associated with impaired mitochondrial biogenesis.[23][25][26] AMPK expression has also been shown to diminish with age, which may also contribute to suppressing mitochondrial biogenesis.[5][26]
Clinical Applications of Targeting Mitochondrial Biogenesis
Mitochondrial biogenesis can be targeted to prevent cancer proliferation. Specifically, two biogenesis regulators—PGC1α and
References
- ^ PMID 24606795.
- ^ PMID 24606801.
- PMID 23737207.
- ^ S2CID 39267910.
- ^ PMID 20533901.
- PMID 28754286.
- PMID 14736705.
- ^ PMID 26858267.
- ^ S2CID 12627451.
- ^ PMID 22683763.
- ^ PMID 17825565.
- ^ PMID 18430751.
- ^ PMID 22936770.
- ^ S2CID 33936266.
- ^ PMID 15961417.
- PMID 30420520.
- ^ PMID 23602910.
- ^ PMID 15454086.
- ^ PMID 20933024.
- ^ PMID 18628400.
- PMID 33770508.
- PMID 22146081.
- ^ PMID 21407239.
- S2CID 25332524.
- ^ PMID 21307849.
- ^ PMID 22588366.
- PMID 20933024.
- S2CID 256111946.
- PMID 18288422.
- PMID 31501275.
Further reading
- Smith JA, Stallons LJ, Collier JB, Chavin KD, Schnellmann RG (February 2015). "Suppression of mitochondrial biogenesis through toll-like receptor 4-dependent mitogen-activated protein kinase kinase/extracellular signal-regulated kinase signaling in endotoxin-induced acute kidney injury". The Journal of Pharmacology and Experimental Therapeutics. 352 (2): 346–57. PMID 25503387.
- Cameron RB, Beeson CC, Schnellmann RG (December 2016). "Development of Therapeutics That Induce Mitochondrial Biogenesis for the Treatment of Acute and Chronic Degenerative Diseases". Journal of Medicinal Chemistry. 59 (23): 10411–10434. PMID 27560192.
- Whitaker RM, Corum D, Beeson CC, Schnellmann RG (2016). "Mitochondrial Biogenesis as a Pharmacological Target: A New Approach to Acute and Chronic Diseases". Annual Review of Pharmacology and Toxicology. 56: 229–49. PMID 26566156.
